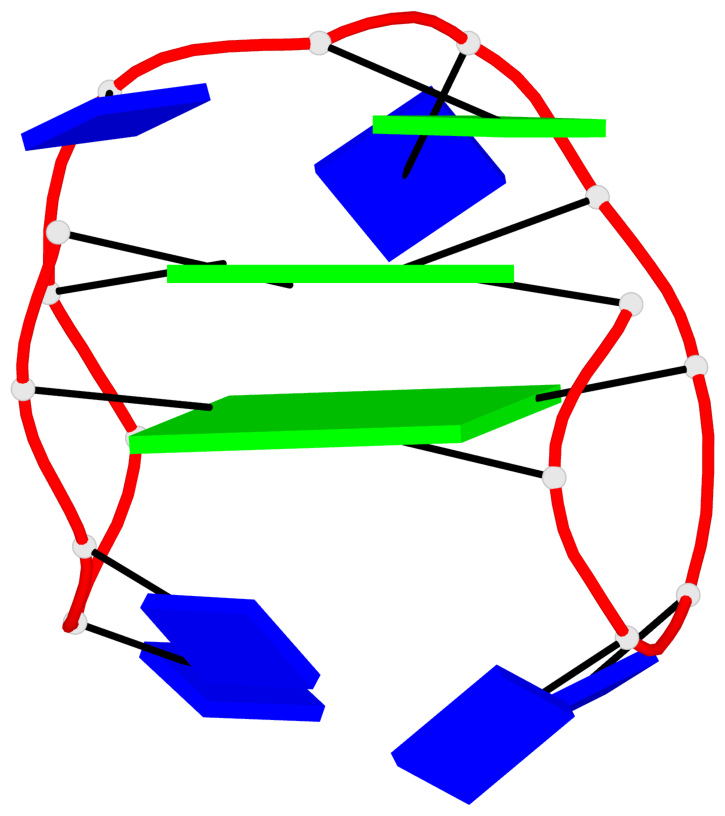
Detailed DSSR results for the G-quadruplex: PDB entry 7v3t
Created and maintained by Xiang-Jun Lu <xiangjun@x3dna.org>
Citation: Please cite the NAR'20 DSSR-PyMOL schematics paper and/or the NAR'15 DSSR method paper.
Summary information
- PDB id
- 7v3t
- Class
- DNA
- Method
- NMR
- Summary
- Solution structure of thrombin binding aptamer g-quadruplex bound a self-adaptive small molecule with rotated ligands
- Reference
- Zhu BC, He J, Xia XY, Jiang J, Liu W, Liu LY, Liang BB, Yao HG, Ke Z, Xia W, Mao ZW (2022): "Solution structure of a thrombin binding aptamer complex with a non-planar platinum(ii) compound." Chem Sci, 13, 8371-8379. doi: 10.1039/d2sc01196d.
- Abstract
- Thrombin Binding Aptamer (TBA) is a monomolecular well-defined two G-tetrad antiparallel G-quadruplex DNA that inhibits the activity of human α-thrombin. In this report, we synthesized a quasi-cross-shaped platinum(ii) compound (L'2LPt) with one cyclometalated and two carbene ligands. We found L'2LPt has selective affinity to bind the TBA G-quadruplex. A fibrinogen clotting assay revealed that L'2LPt can abrogate the inhibitory activity of TBA against thrombin. We solved the 1 : 1 L'2LPt-TBA complex structure by NMR, which revealed a unique self-adaptive property of L'2LPt upon binding to TBA. In the complex, a carbene ligand of L'2LPt rotates to pair with the cyclometalated ligand to form a plane stacking over half of the TBA G-tetrad and covered by lateral TT loops. It is notable that the heavy atom Pt stays out of the G-tetrad. Meanwhile, the other carbene ligand remains relatively perpendicular and forms a hydrogen bond with a guanine to anchor the L'2LPt position. This structure exhibits a quasi-cross-shaped Pt(ii) compound bound to the G-quadruplex with an unusual "wall-mounted" binding mode. Our structures provide insights into the specific recognition of antiparallel G-quadruplex DNA by a self-adaptive Pt(ii) compound and useful information for the design of selective G-quadruplex targeting non-planar molecules.
- G4 notes
- 2 G-tetrads, 1 G4 helix, 1 G4 stem, 2(+Ln+Lw+Ln), chair(2+2), UDUD
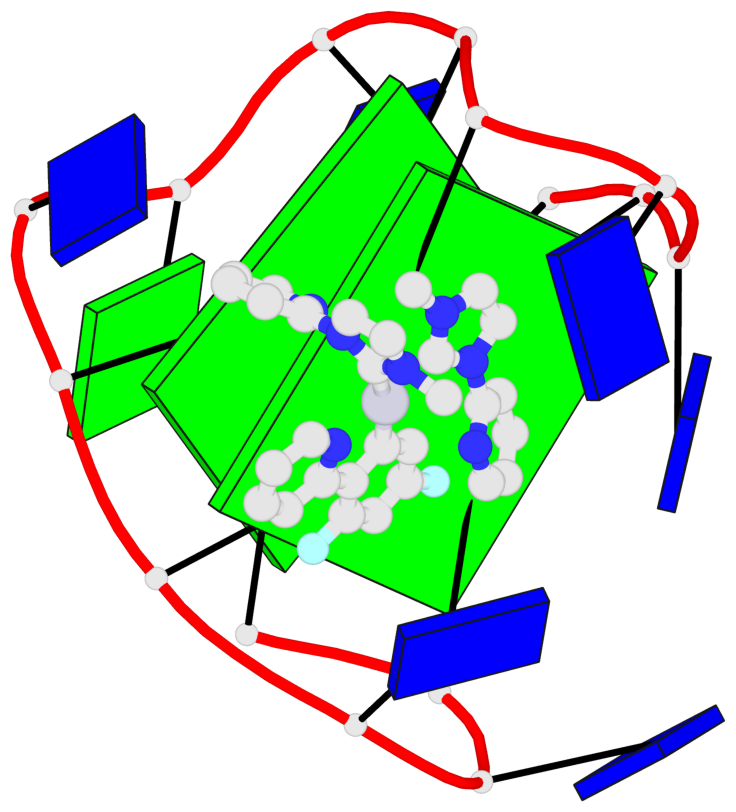
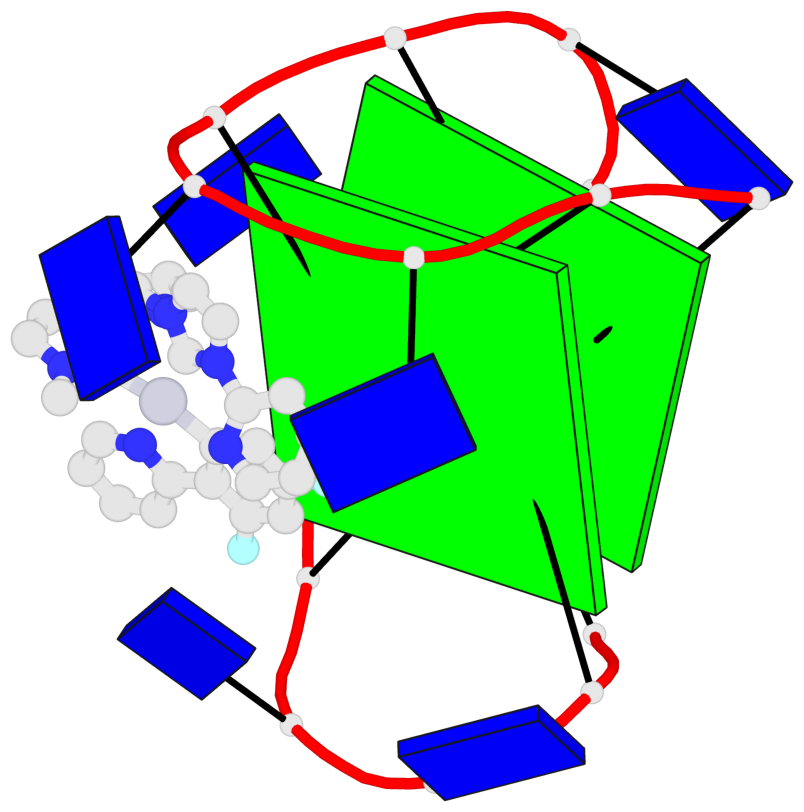
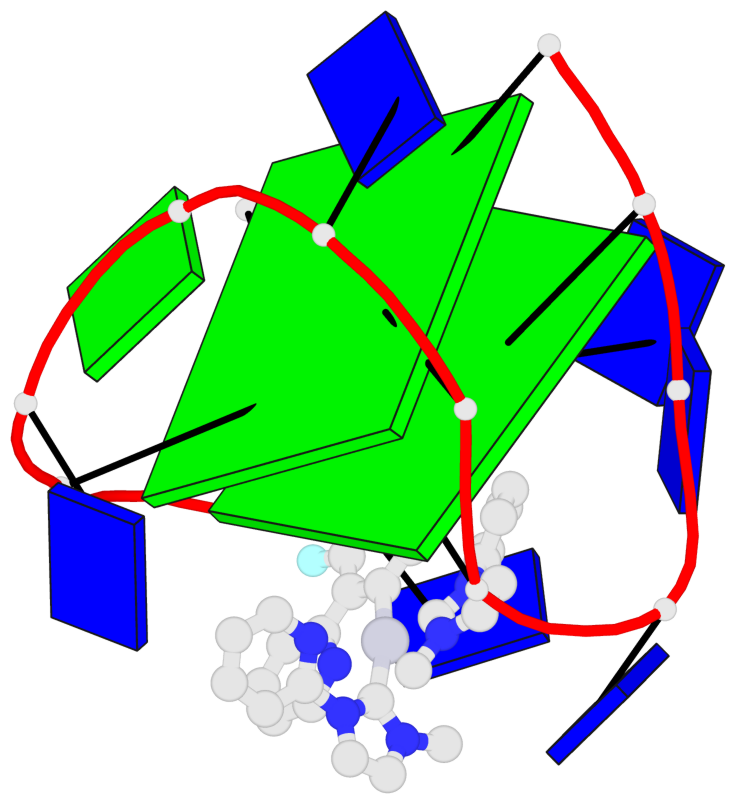
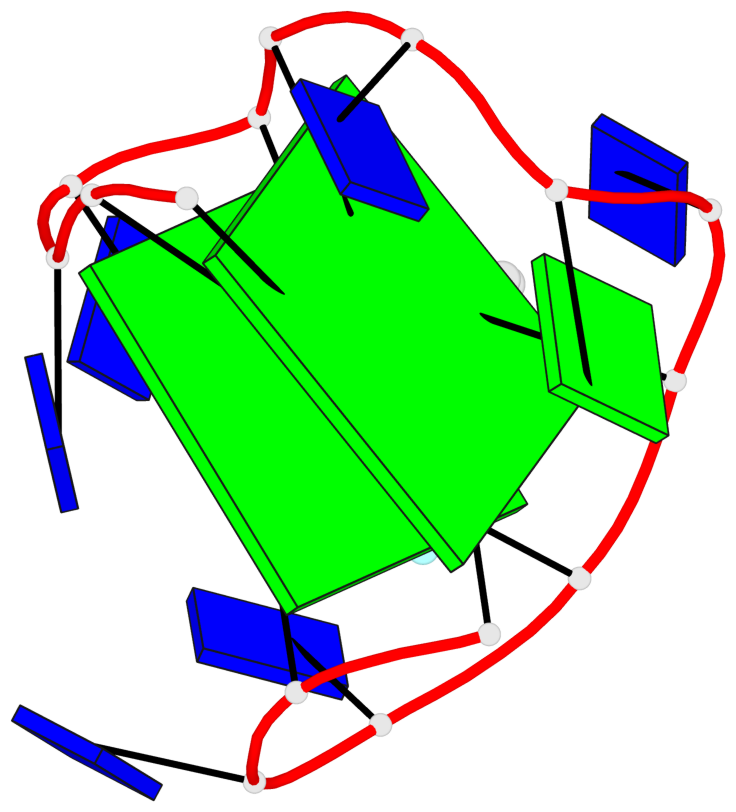
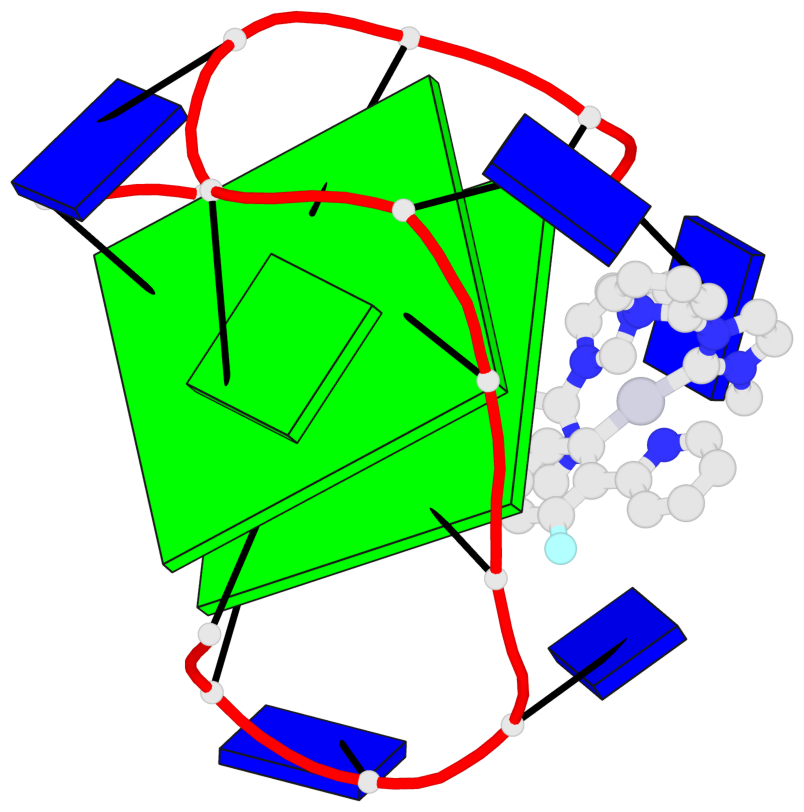
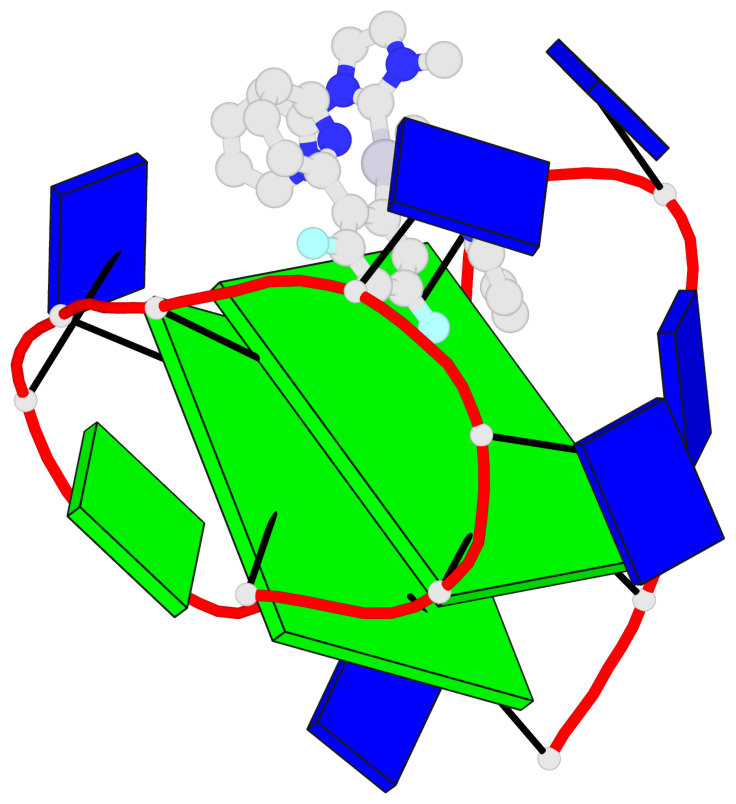
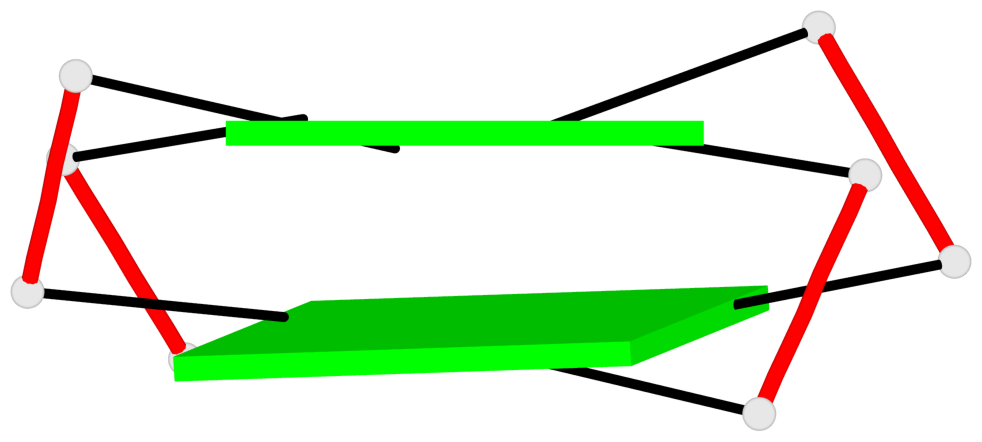
Base-block schematics in six views
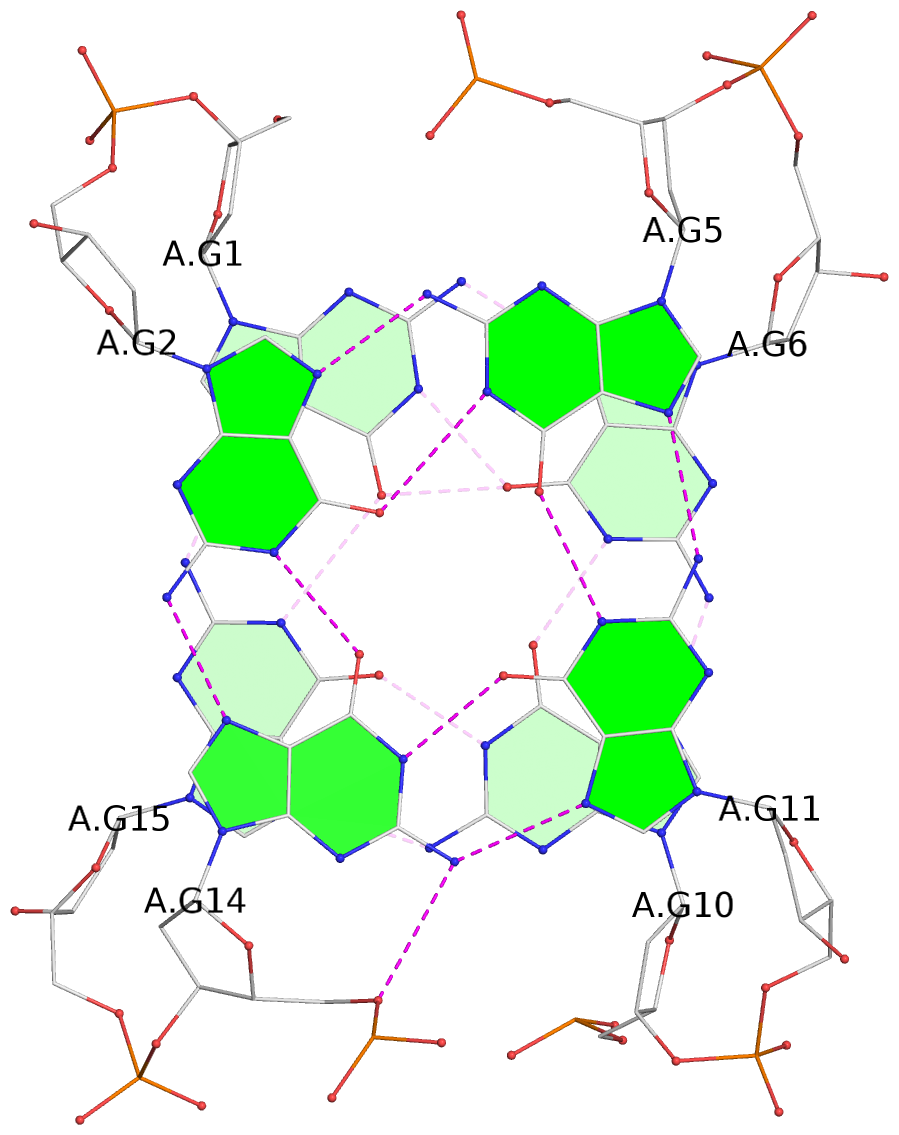
List of 2 G-tetrads
1 glyco-bond=s-s- sugar=.... groove=wnwn planarity=0.207 type=other nts=4 GGGG A.DG1,A.DG15,A.DG10,A.DG6 2 glyco-bond=-s-s sugar=..3- groove=wnwn planarity=0.232 type=other nts=4 GGGG A.DG2,A.DG14,A.DG11,A.DG5
List of 1 G4-helix
In DSSR, a G4-helix is defined by stacking interactions of G-tetrads, regardless of backbone connectivity, and may contain more than one G4-stem.
Helix#1, 2 G-tetrad layers, INTRA-molecular, with 1 stem
List of 1 G4-stem
In DSSR, a G4-stem is defined as a G4-helix with backbone connectivity. Bulges are also allowed along each of the four strands.