Detailed DSSR results for the G-quadruplex: PDB entry 148d
Created and maintained by Xiang-Jun Lu <xiangjun@x3dna.org>
Citation: Please cite the NAR'20 DSSR-PyMOL schematics paper and/or the NAR'15 DSSR method paper.
Summary information
- PDB id
- 148d
- Class
- DNA
- Method
- NMR
- Summary
- Three-dimensional solution structure of the thrombin binding DNA aptamer d(ggttggtgtggttgg)
- Reference
- Schultze P, Macaya RF, Feigon J (1994): "Three-dimensional solution structure of the thrombin-binding DNA aptamer d(GGTTGGTGTGGTTGG)." J.Mol.Biol., 235, 1532-1547. doi: 10.1006/jmbi.1994.1105.
- Abstract
- The DNA oligonucleotide d(GGTTGGTGTGGTTGG) (thrombin aptamer) binds to thrombin and inhibits its enzymatic activity in the chain of reactions that lead to blood clotting. Two-dimensional 1H NMR studies indicate that the oligonucleotide forms a folded structure in solution, composed of two guanine quartets connected by two T-T loops spanning the narrow grooves at one end and a T-G-T loop spanning a wide groove at the other end. We present the assignment strategy used, methods for the structure determination, and the refined three-dimensional structure of the thrombin aptamer. The initial structures were generated by metric matrix distance geometry using distance and dihedral bond angle constraints from NOE and coupling constants, respectively, and refined by restrained molecular dynamics and direct NOE refinement. Knowledge of the three-dimensional structure of this thrombin aptamer may be relevant for the design of improved thrombin-inhibiting anti-coagulants with similar structural motifs.
- G4 notes
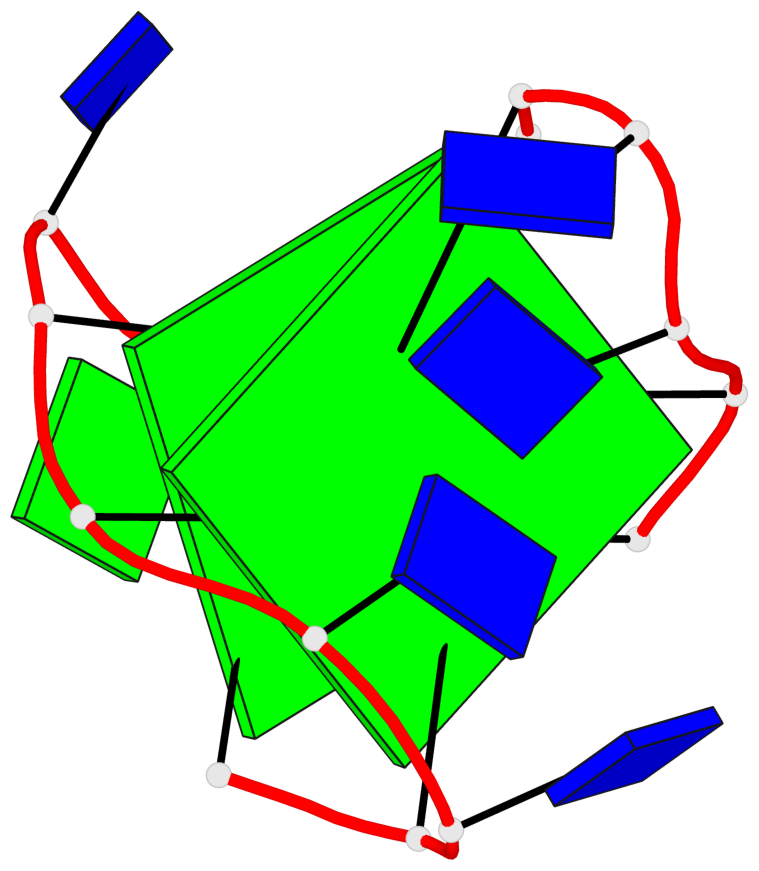
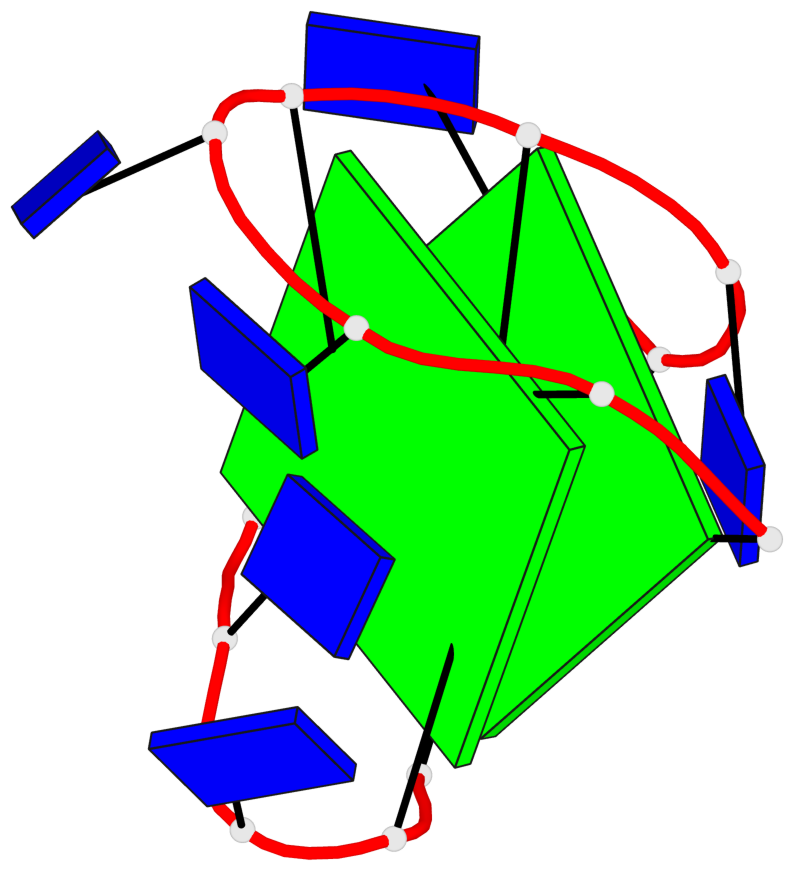
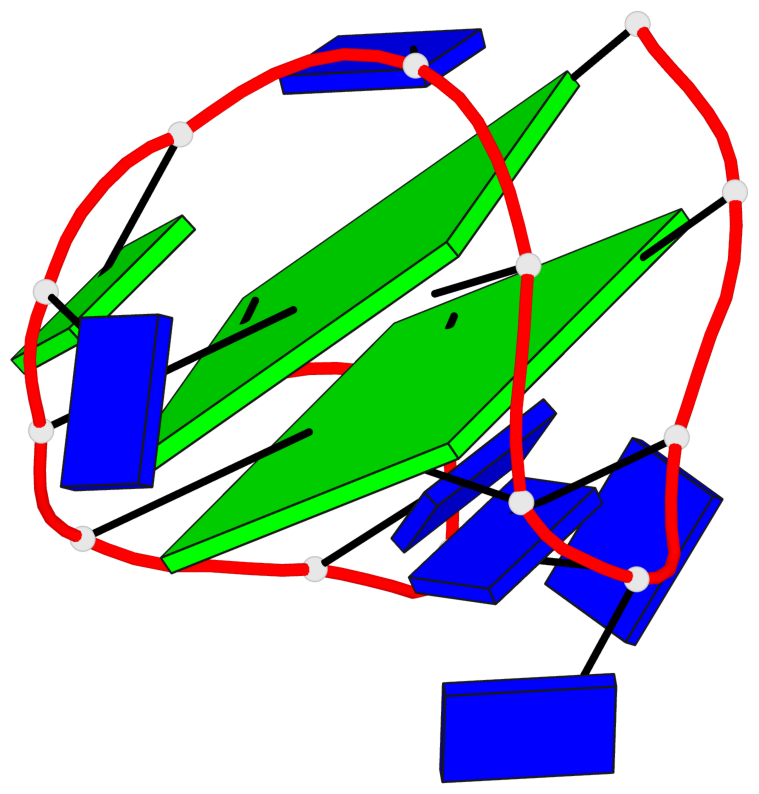
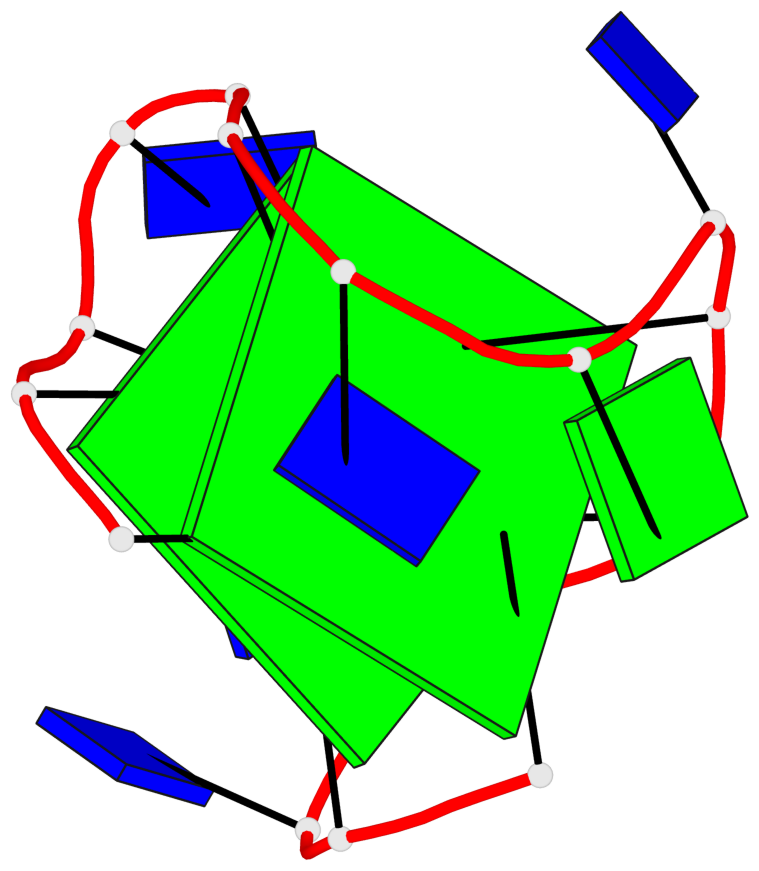
- 2 G-tetrads, 1 G4 helix, 1 G4 stem, 2(+Ln+Lw+Ln), chair(2+2), UDUD
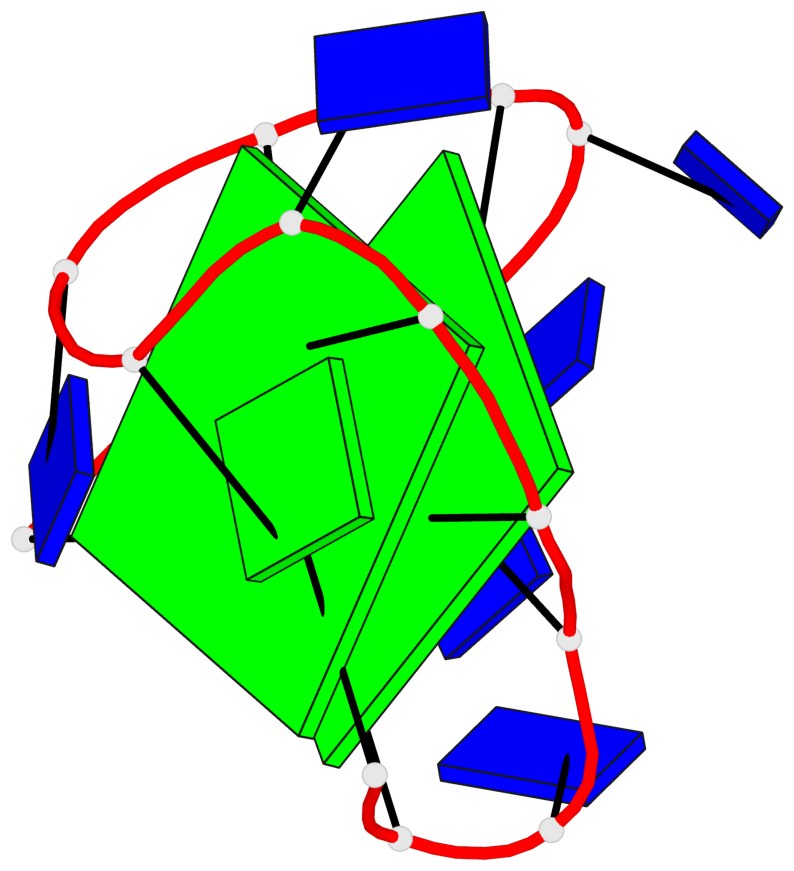
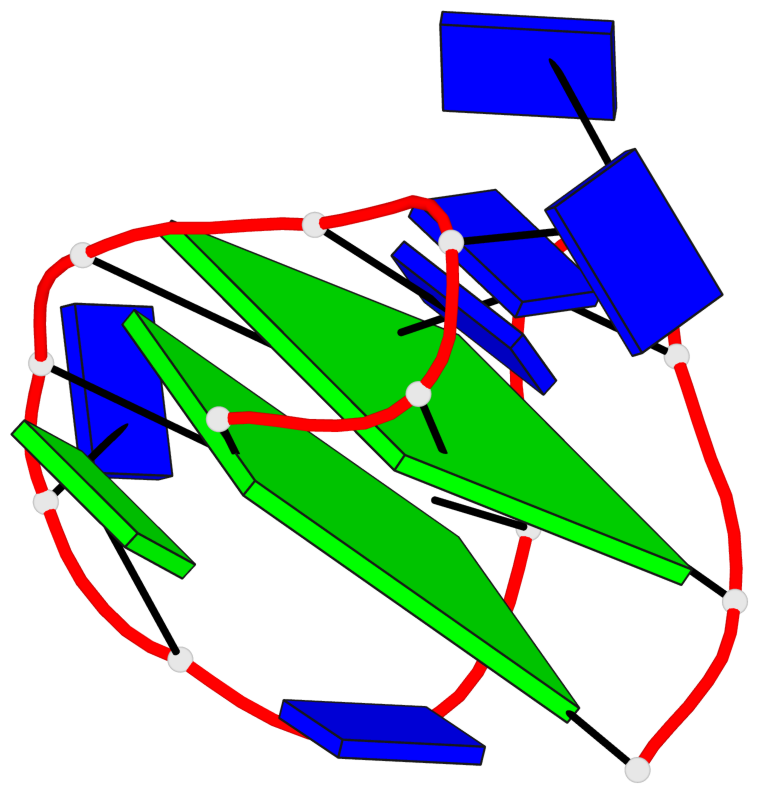
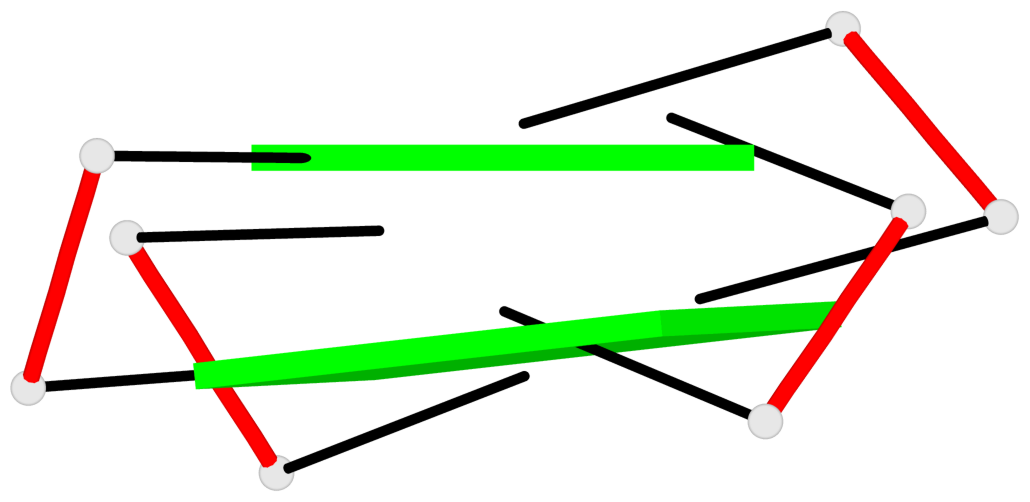
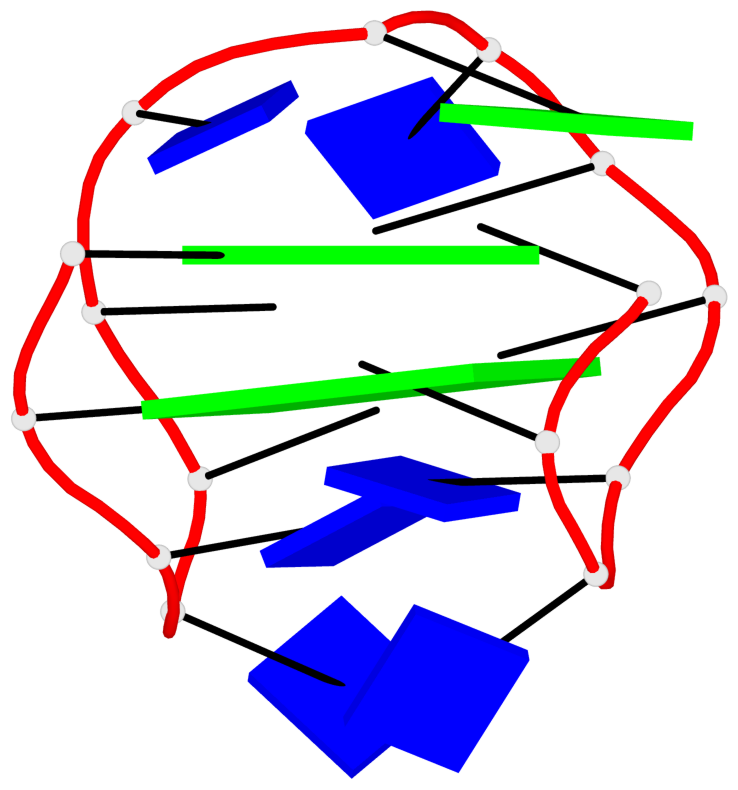
Base-block schematics in six views
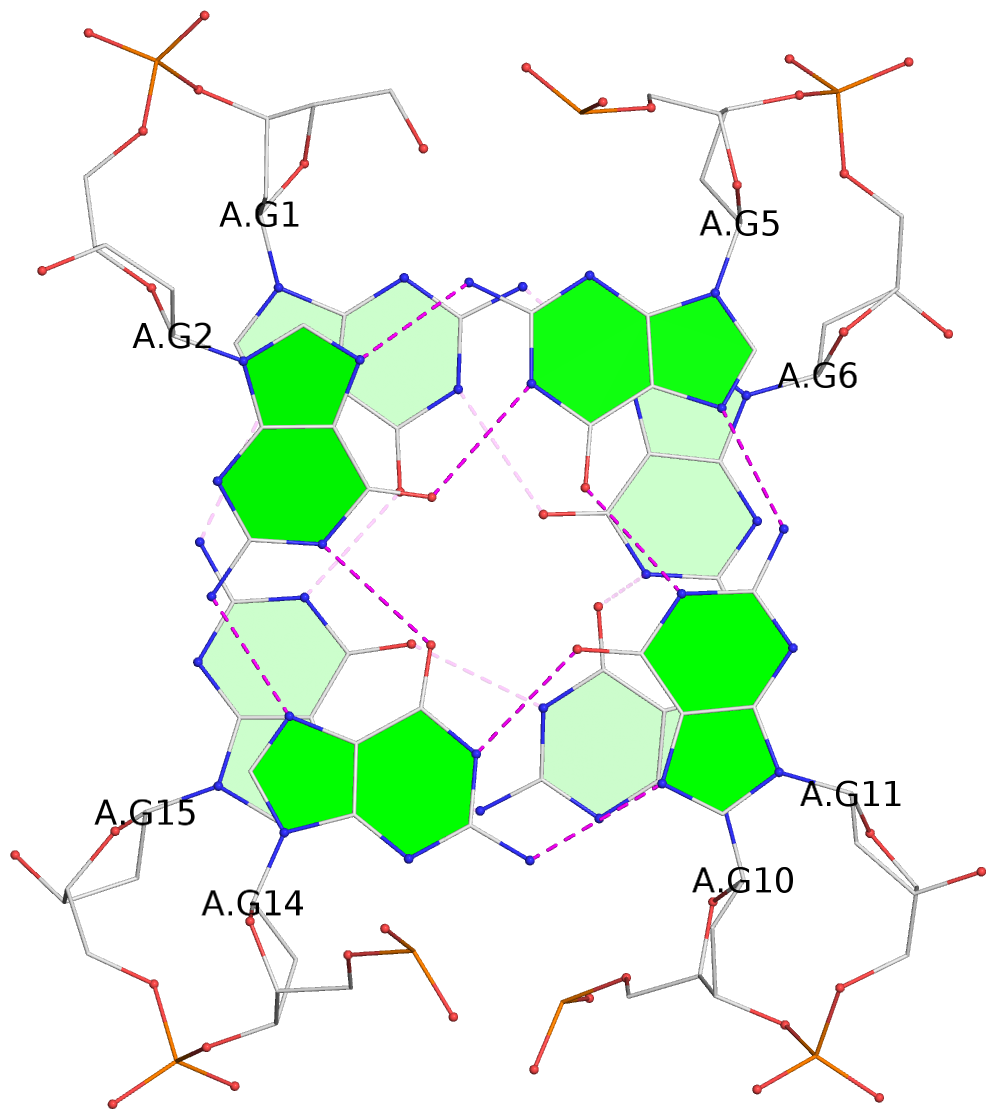
List of 2 G-tetrads
1 glyco-bond=s-s- sugar=---- groove=wnwn planarity=0.583 type=other nts=4 GGGG A.DG1,A.DG15,A.DG10,A.DG6 2 glyco-bond=-s-s sugar=---- groove=wnwn planarity=0.526 type=other nts=4 GGGG A.DG2,A.DG14,A.DG11,A.DG5
List of 1 G4-helix
In DSSR, a G4-helix is defined by stacking interactions of G-tetrads, regardless of backbone connectivity, and may contain more than one G4-stem.
Helix#1, 2 G-tetrad layers, INTRA-molecular, with 1 stem
List of 1 G4-stem
In DSSR, a G4-stem is defined as a G4-helix with backbone connectivity. Bulges are also allowed along each of the four strands.