Detailed DSSR results for the G-quadruplex: PDB entry 156d
Created and maintained by Xiang-Jun Lu <xiangjun@x3dna.org>
Citation: Please cite the NAR'20 DSSR-PyMOL schematics paper and/or the NAR'15 DSSR method paper.
Summary information
- PDB id
- 156d
- Class
- DNA
- Method
- NMR
- Summary
- Refined solution structure of the dimeric quadruplex formed from the oxytricha telomeric oligonucleotide d(ggggttttgggg)
- Reference
- Schultze P, Smith FW, Feigon J (1994): "Refined solution structure of the dimeric quadruplex formed from the Oxytricha telomeric oligonucleotide d(GGGGTTTTGGGG)." Structure, 2, 221-233. doi: 10.1016/S0969-2126(00)00023-X.
- Abstract
- Background: Telomeres, the structures at the ends of linear eukaryotic chromosomes, are essential for chromosome replication and stability. The telomeres of the unicellular ciliate Oxytricha contain a 3' single strand overhang composed of two repeats of the telomere repeat sequence d(TTTTGGGG). It has been proposed that oligonucleotides containing this repeat can form DNA quadruplexes via hydrogen bonding of the guanines into quartets. Such structures may be relevant to the biological function of the telomere, and in G-rich sequences elsewhere in the genome.
Results: We have previously determined from solution NMR data that the Oxy-1.5 Oxytricha repeat oligonucleotide d(GGGGTTTTGGGG) dimerizes to form an intermolecular quadruplex composed of four guanine quartets and with the thymines in loops across the diagonal at opposite ends of the quadruplex. We report here the refined solution structure of Oxy-1.5. This structure is compared with the previously published crystal structure of the same oligonucleotide.
Conclusions: Oxy-1.5 forms a well-defined, symmetrical structure with ordered thymine loops. Both the solution and crystal structures of Oxy-1.5 are quadruplexes with alternating syn and anti glycosyl conformation of guanines along each strand of the helix and have thymine loops at opposite ends. However, the topology of the two structures is fundamentally different, leading to significant structural differences. A topological pathway for the formation and interconversion of the two structures is proposed. - G4 notes
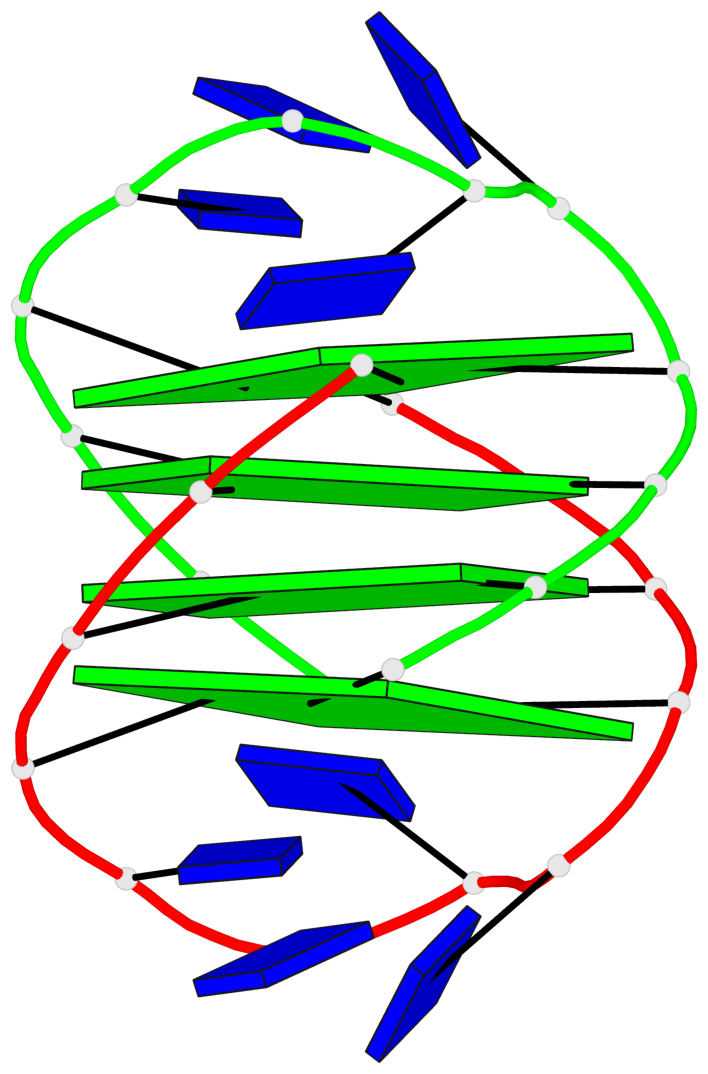
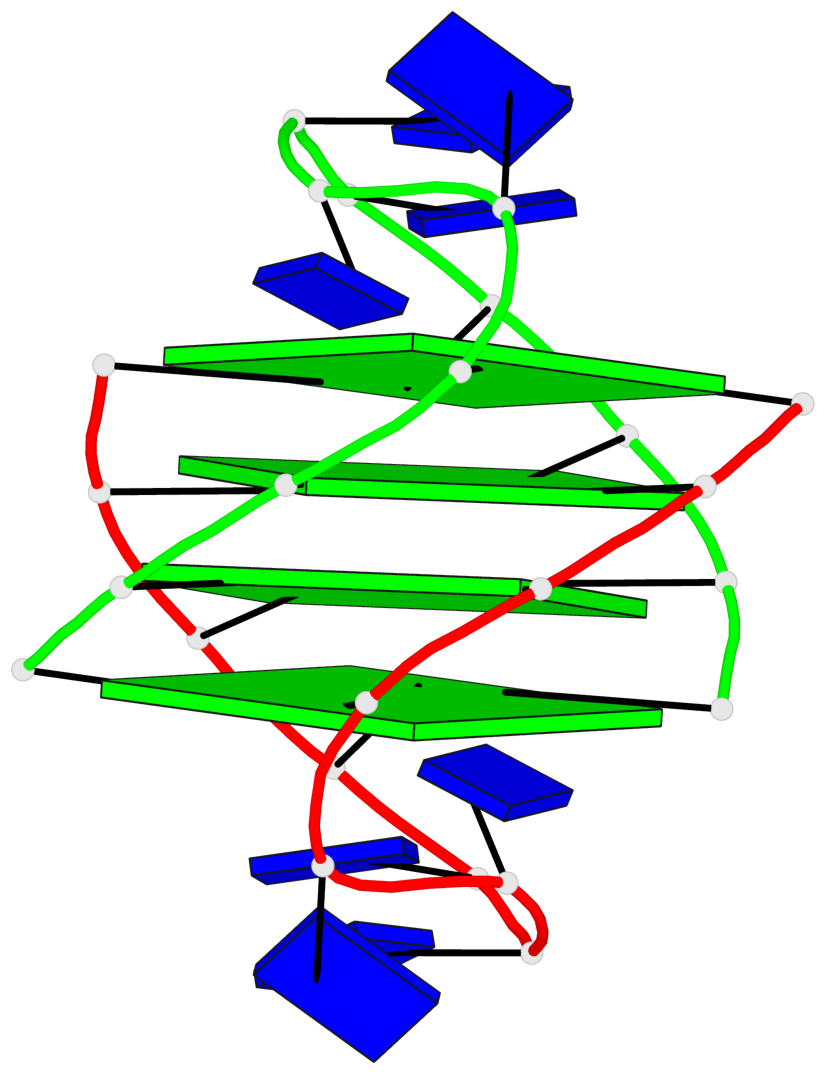
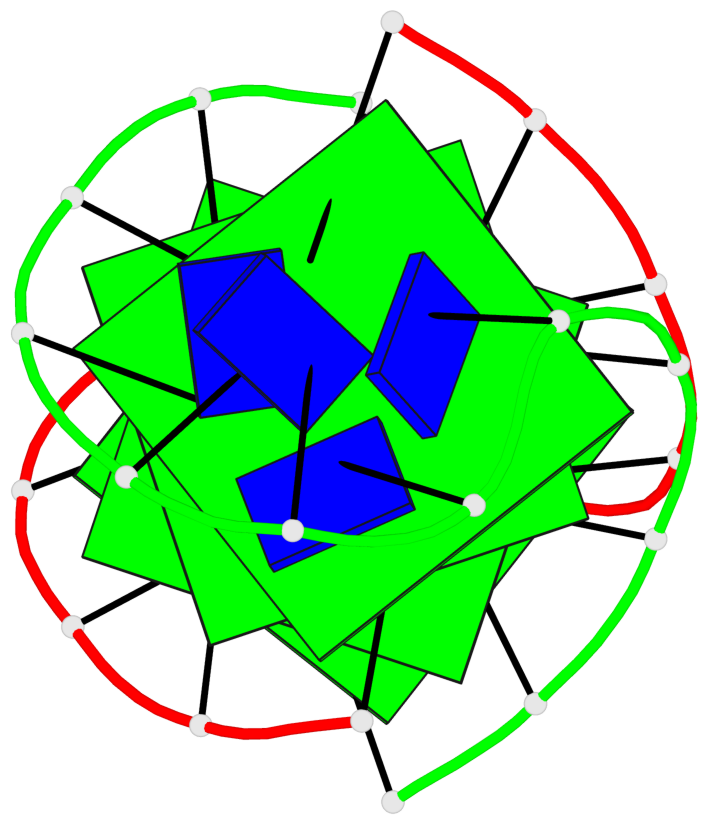
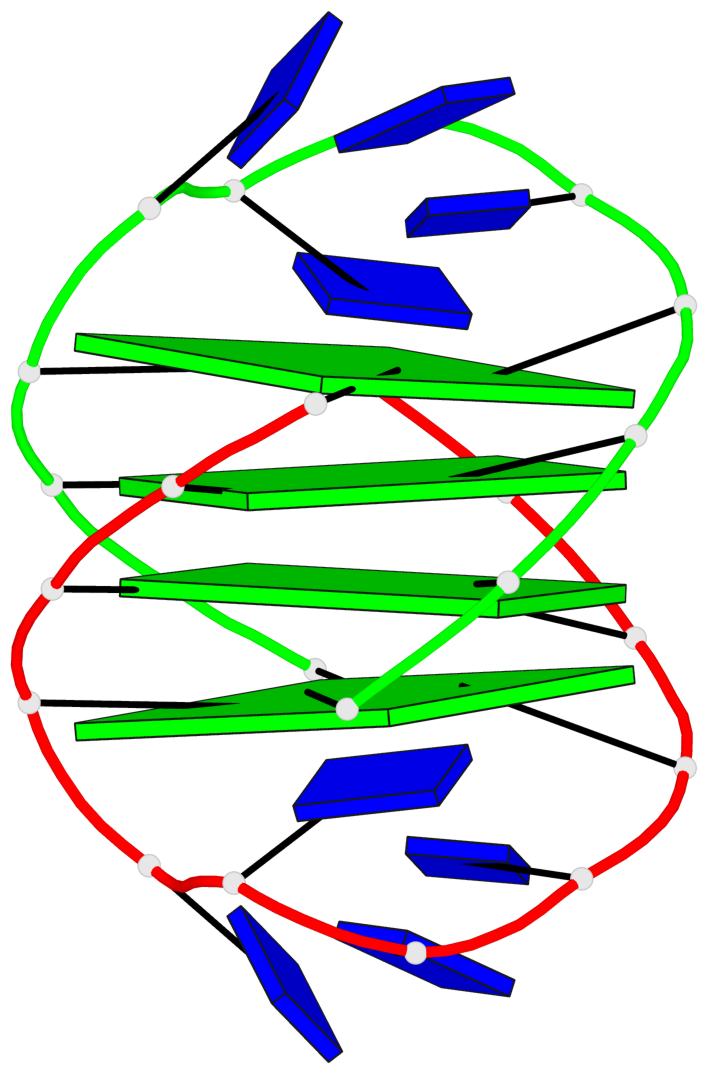
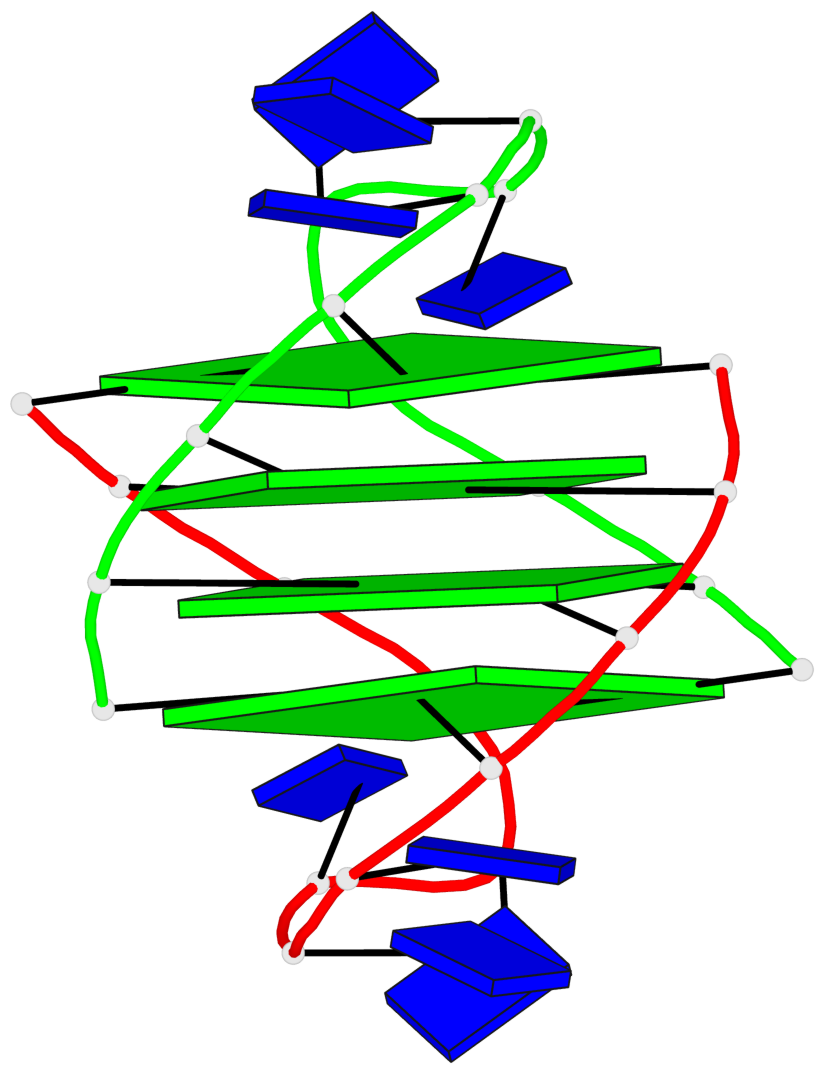
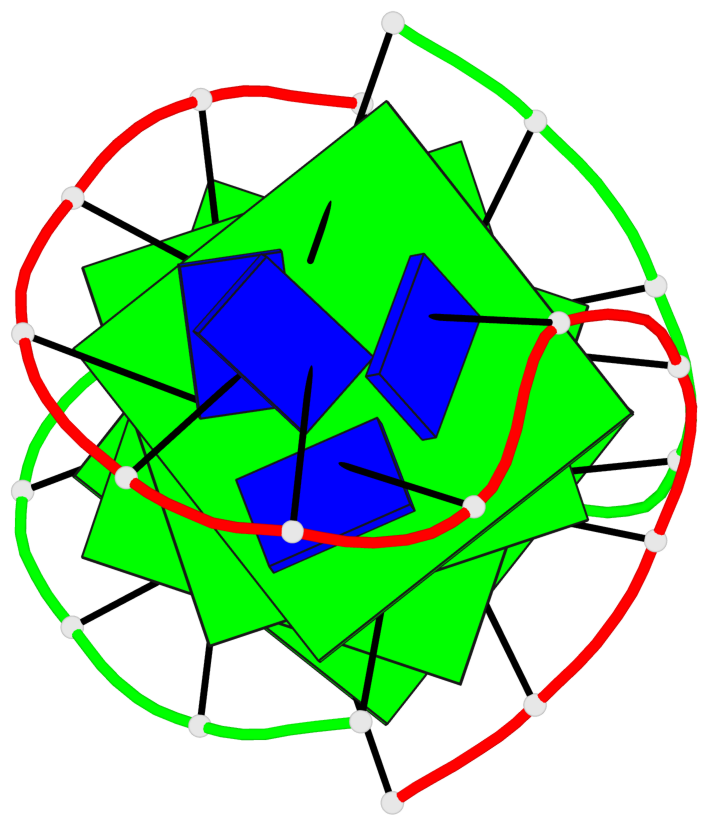
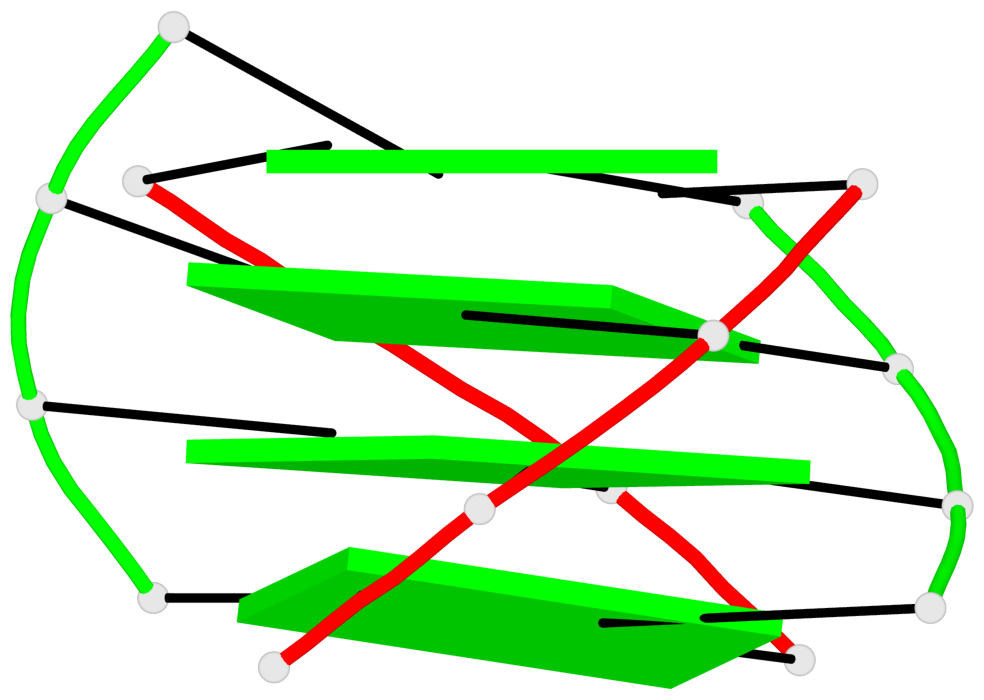
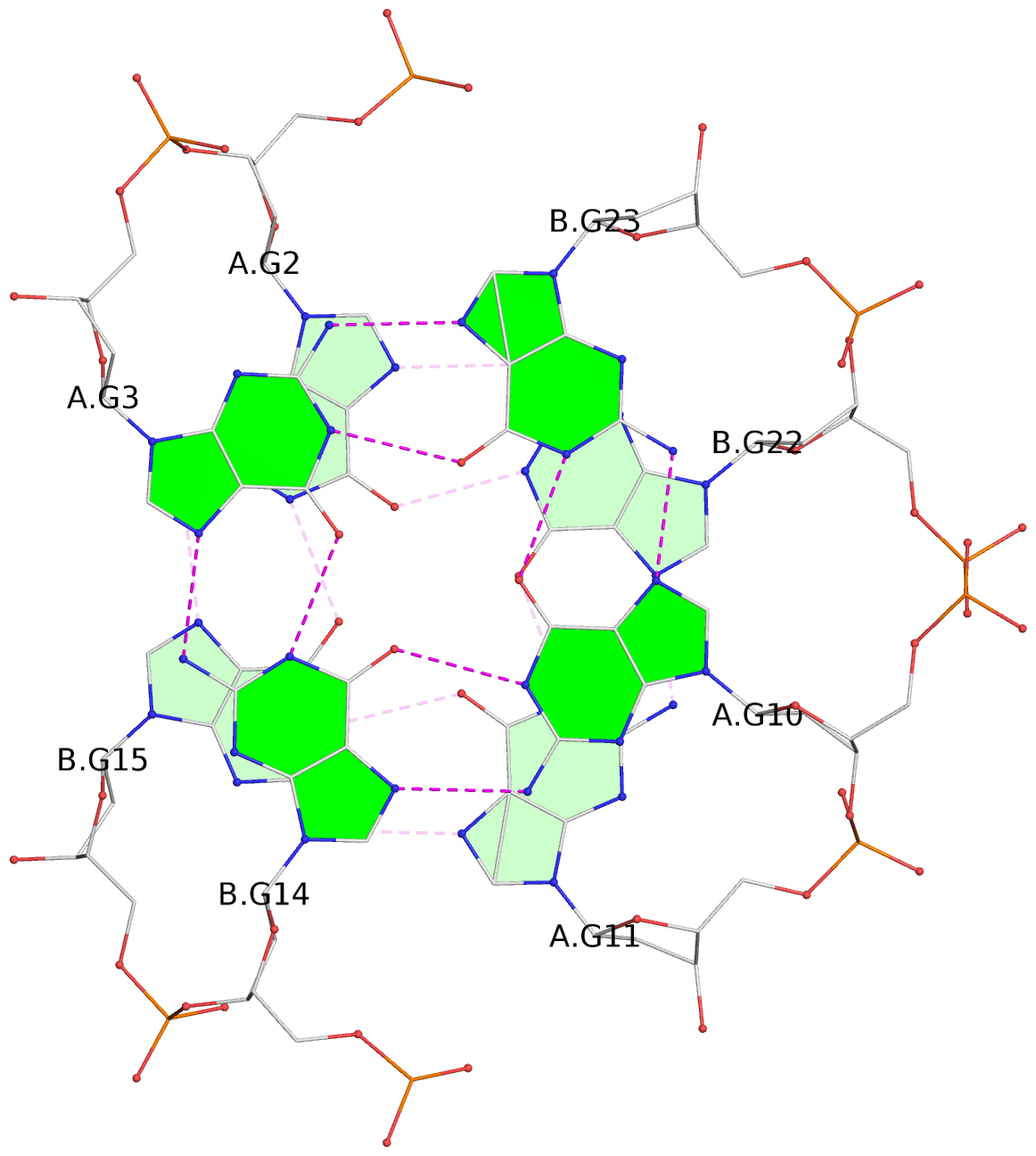
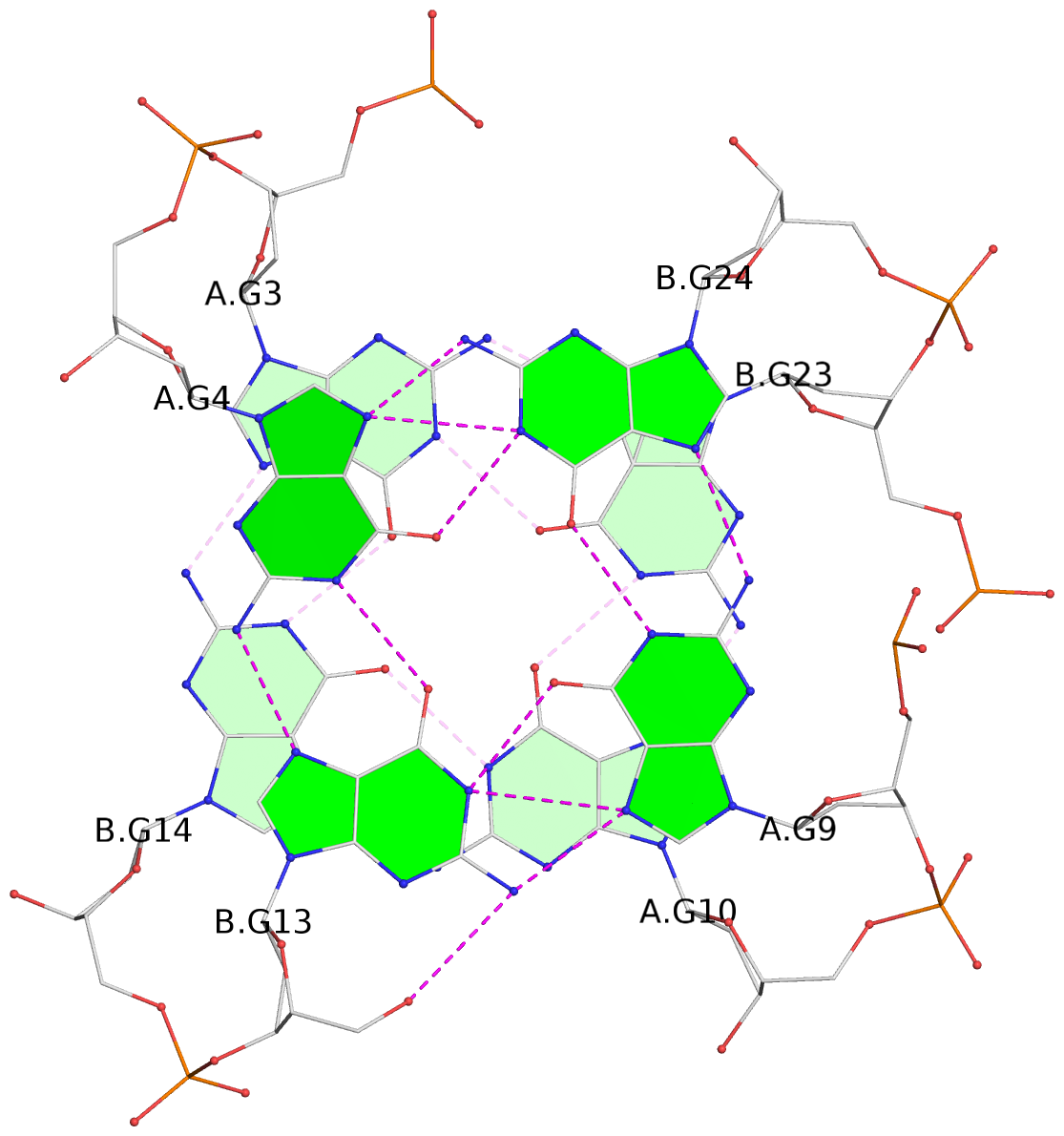
- 4 G-tetrads, 1 G4 helix, 1 G4 stem, (2+2), UDDU
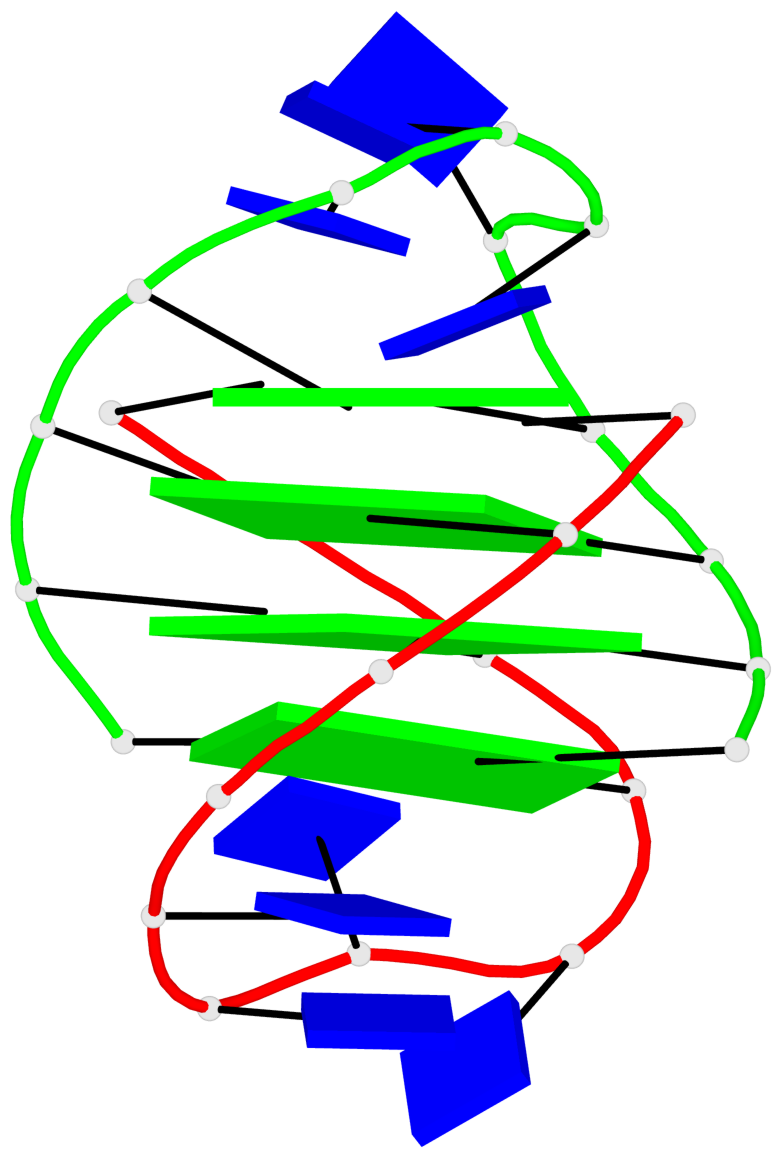
Base-block schematics in six views
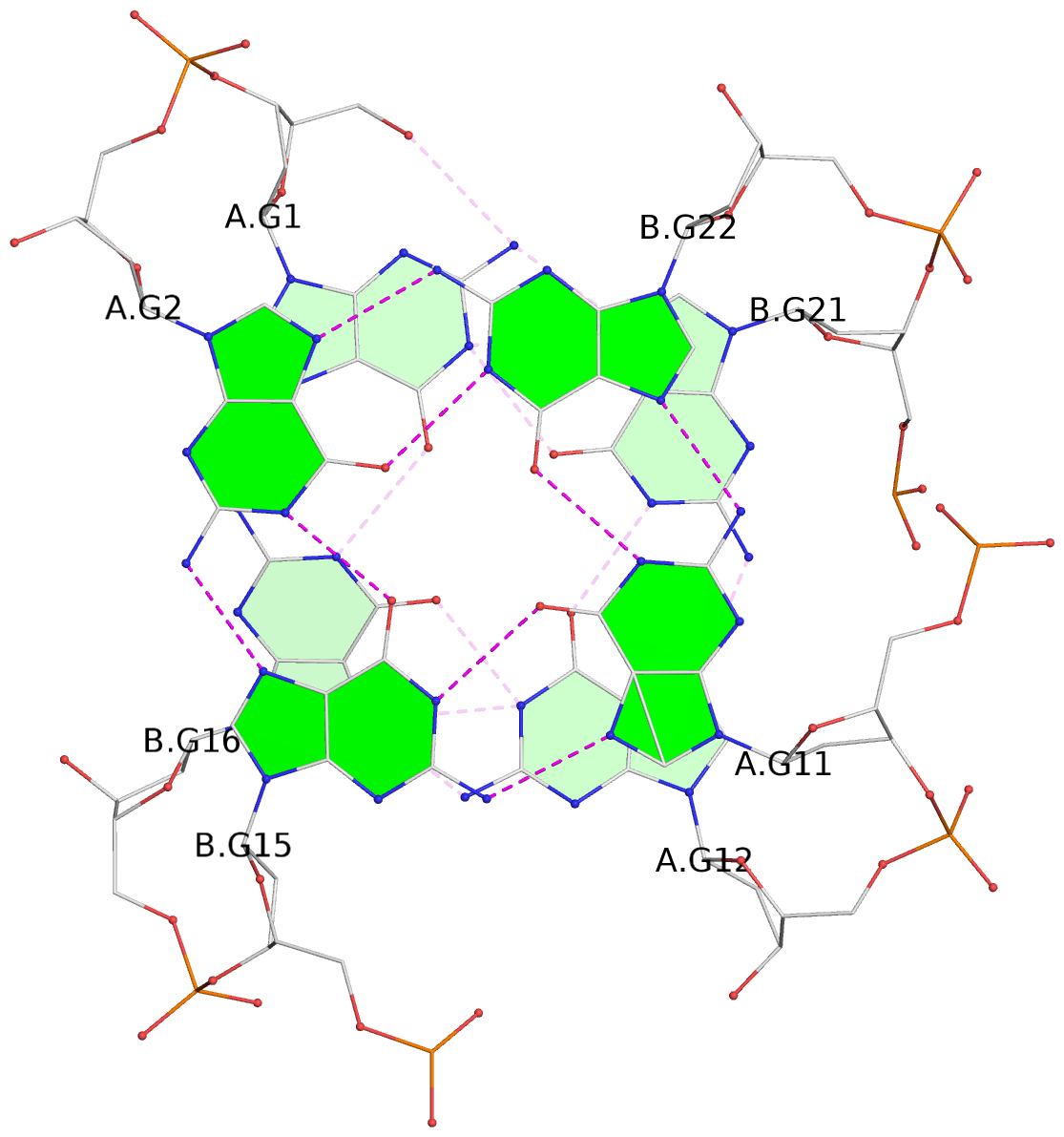
List of 4 G-tetrads
1 glyco-bond=s--s sugar=---- groove=w-n- planarity=0.458 type=other nts=4 GGGG A.DG1,B.DG16,A.DG12,B.DG21 2 glyco-bond=-ss- sugar=---- groove=w-n- planarity=0.303 type=other nts=4 GGGG A.DG2,B.DG15,A.DG11,B.DG22 3 glyco-bond=s--s sugar=---- groove=w-n- planarity=0.304 type=other nts=4 GGGG A.DG3,B.DG14,A.DG10,B.DG23 4 glyco-bond=-ss- sugar=---- groove=w-n- planarity=0.459 type=other nts=4 GGGG A.DG4,B.DG13,A.DG9,B.DG24
List of 1 G4-helix
In DSSR, a G4-helix is defined by stacking interactions of G-tetrads, regardless of backbone connectivity, and may contain more than one G4-stem.
Helix#1, 4 G-tetrad layers, inter-molecular, with 1 stem
List of 1 G4-stem
In DSSR, a G4-stem is defined as a G4-helix with backbone connectivity. Bulges are also allowed along each of the four strands.