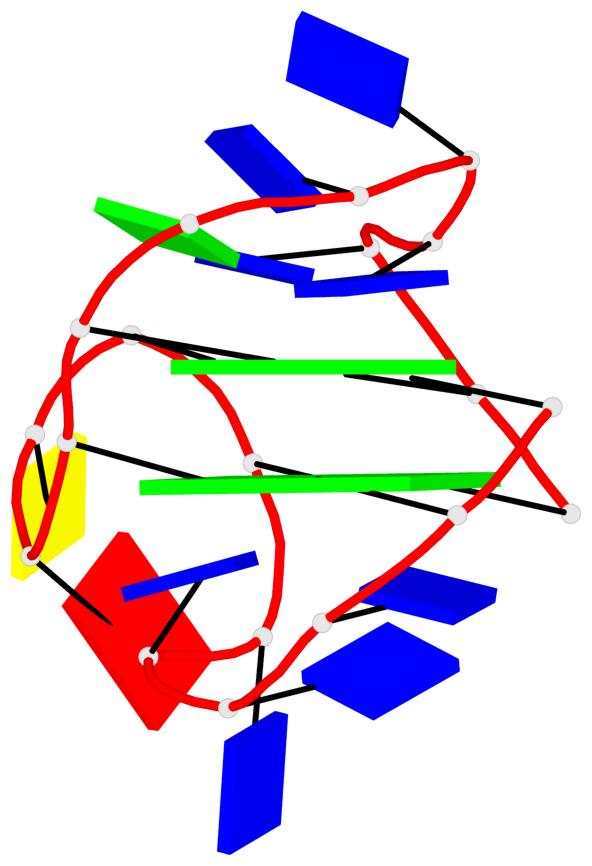
Detailed DSSR results for the G-quadruplex: PDB entry 1i34
Created and maintained by Xiang-Jun Lu <xiangjun@x3dna.org>
Citation: Please cite the NAR'20 DSSR-PyMOL schematics paper and/or the NAR'15 DSSR method paper.
Summary information
- PDB id
- 1i34
- Class
- DNA
- Method
- NMR
- Summary
- Solution DNA quadruplex with double chain reversal loop and two diagonal loops connecting gggg tetrads flanked by g-(t-t) triad and t-t-t triple
- Reference
- Kuryavyi V, Majumdar A, Shallop A, Chernichenko N, Skripkin E, Jones R, Patel DJ (2001): "A double chain reversal loop and two diagonal loops define the architecture of a unimolecular DNA quadruplex containing a pair of stacked G(syn)-G(syn)-G(anti)-G(anti) tetrads flanked by a G-(T-T) Triad and a T-T-T triple." J.Mol.Biol., 310, 181-194. doi: 10.1006/jmbi.2001.4759.
- Abstract
- The architecture of G-G-G-G tetrad-aligned DNA quadruplexes in monovalent cation solution is dependent on the directionality of the four strands, which in turn are defined by loop connectivities and the guanine syn/anti distribution along individual strands and within individual G-G-G-G tetrads. The smallest unimolecular G-quadruplex belongs to the d(G2NnG2NnG2NnG2) family, which has the potential to form two stacked G-tetrads linked by Nn loop connectivities. Previous studies have focused on the thrombin-binding DNA aptamer d(G2T2G2TGTG2T2G2), where Nn was T2 for the first and third connecting loops and TGT for the middle connecting loop. This DNA aptamer in K(+) cation solution forms a unimolecular G-quadruplex stabilized by two stacked G(syn)-G(anti)-G(syn)-G(anti) tetrads, adjacent strands which are antiparallel to each other and edge-wise connecting T2, TGT and T2 loops. We now report on the NMR-based solution structure of the d(G2T4G2CAG2GT4G2T) sequence, which differs from the thrombin-binding DNA aptamer sequence in having longer first (T4) and third (GT4) loops and a shorter (CA) middle loop. This d(G2T4G2CAG2GT4G2T) sequence in Na(+) cation solution forms a unimolecular G-quadruplex stabilized by two stacked G(syn)-G(syn)-G(anti)-G(anti) tetrads, adjacent strands which have one parallel and one antiparallel neighbors and distinct non-edge-wise loop connectivities. Specifically, the longer first (T4) and third (GT4) loops are of the diagonal type while the shorter middle loop is of the double chain reversal type. In addition, the pair of stacked G-G-G-G tetrads are flanked on one side by a G-(T-T) triad and on the other side by a T-T-T triple. The distinct differences in strand directionalities, loop connectivities and syn/anti distribution within G-G-G-G tetrads between the thrombin-binding DNA aptamer d(G2T2G2TGTG2T2G2) quadruplex reported previously, and the d(G2T4G2CAG2GT4G2T) quadruplex reported here, reinforces the polymorphic nature of higher-order DNA architectures. Further, these two small unimolecular G-quadruplexes, which are distinct from each other and from parallel-stranded G-quadruplexes, provide novel targets for ligand recognition. Our results demonstrate that the double chain reversal loop connectivity identified previously by our laboratory within the Tetrahymena telomere d(T2G4)4 quadruplex, is a robust folding topology, since it has now also been observed within the d(G2T4G2CAG2GT4G2T) quadruplex. The identification of a G-(T-T) triad and a T-T-T triple, expands on the available recognition alignments for base triads and triples.
- G4 notes
- 2 G-tetrads, 1 G4 helix, 1 G4 stem, 2(D+PD), (2+2), UDDU
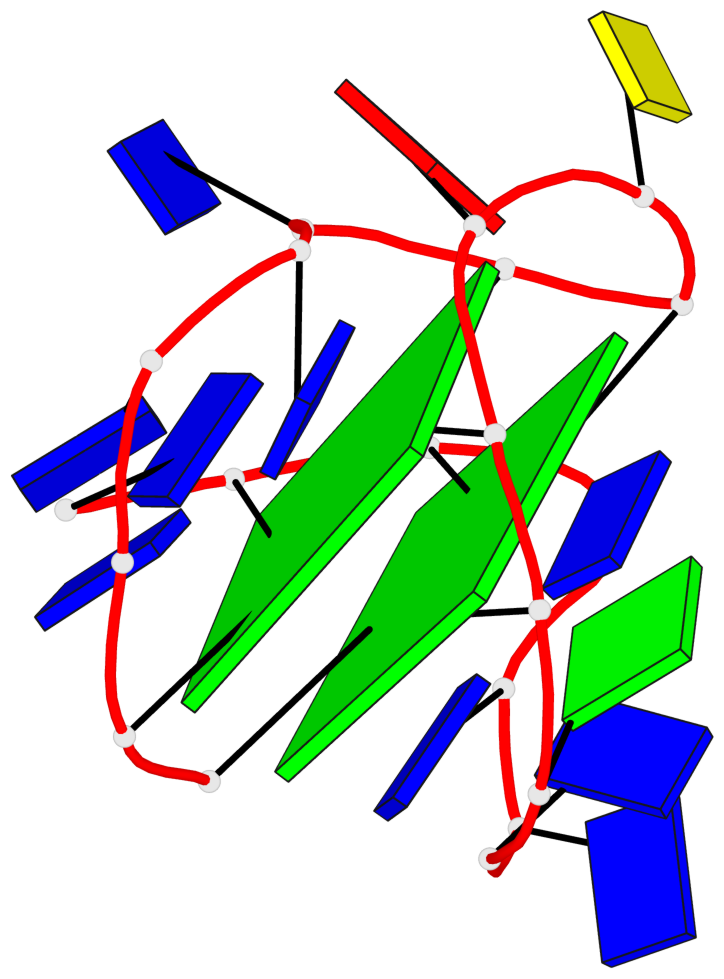
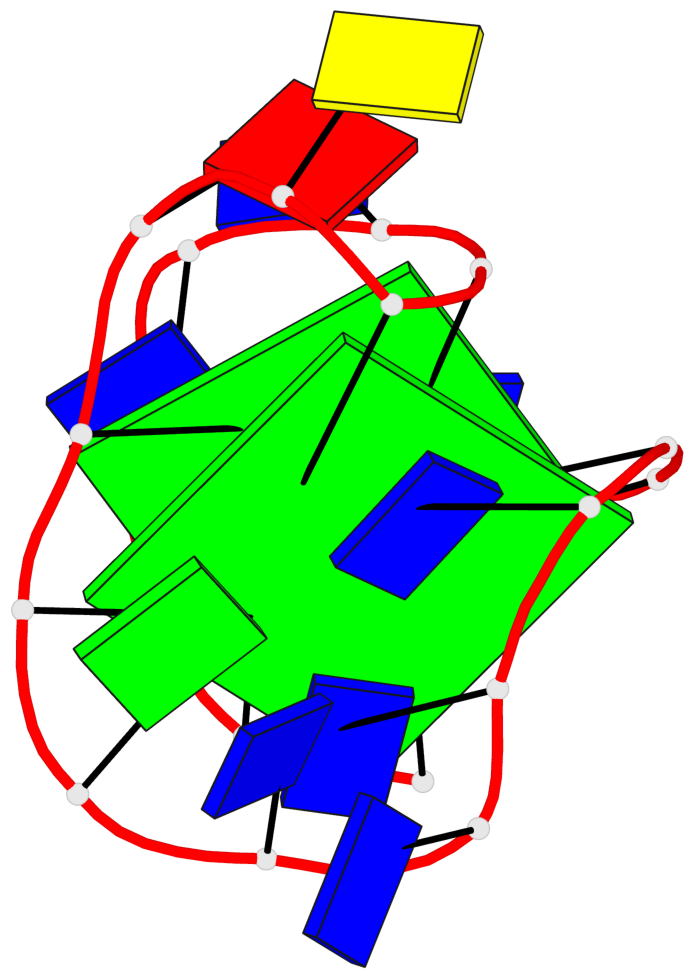
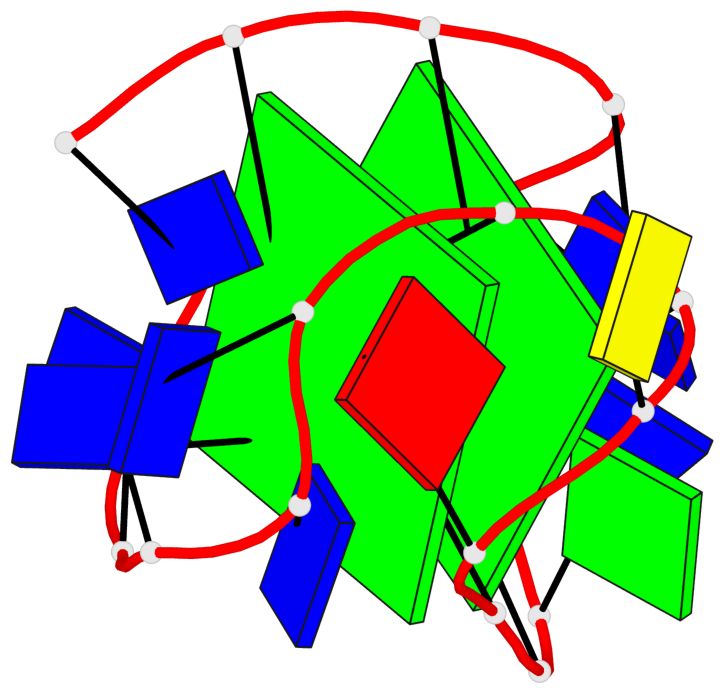
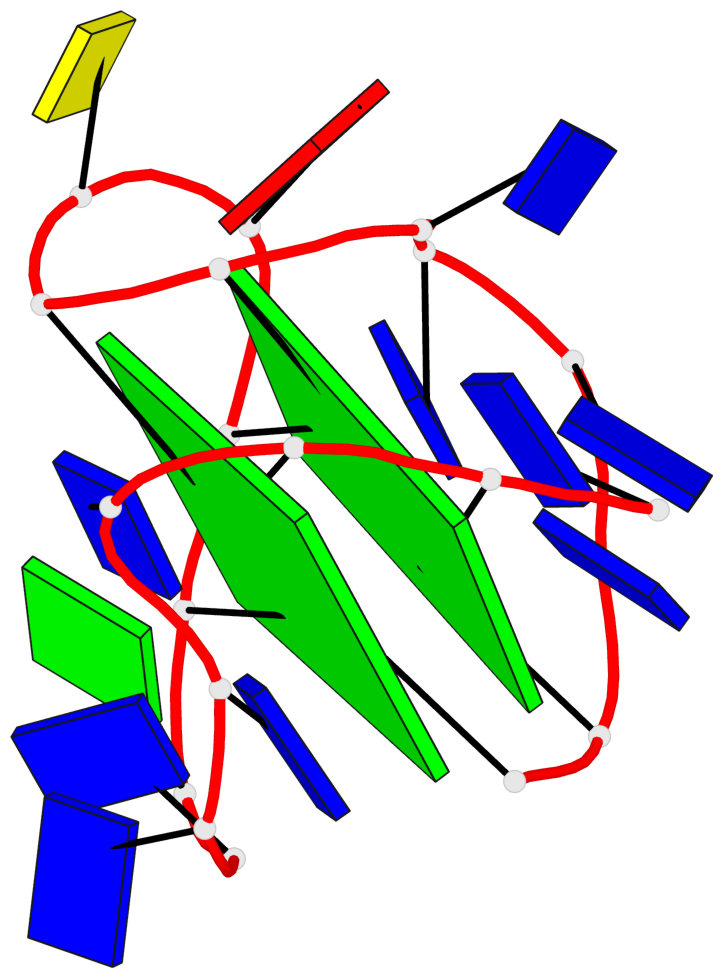
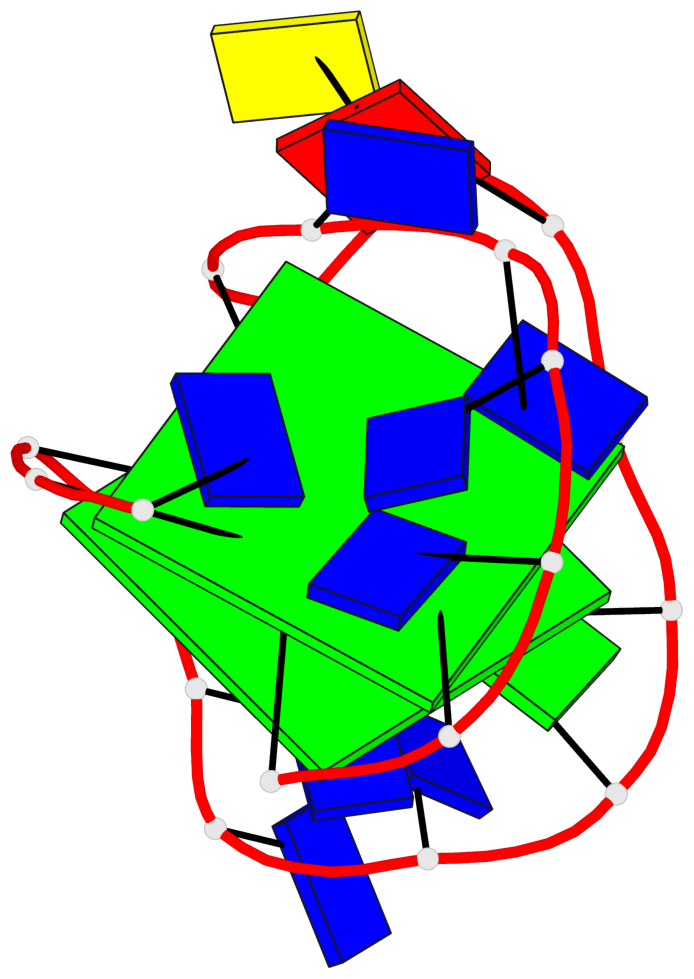
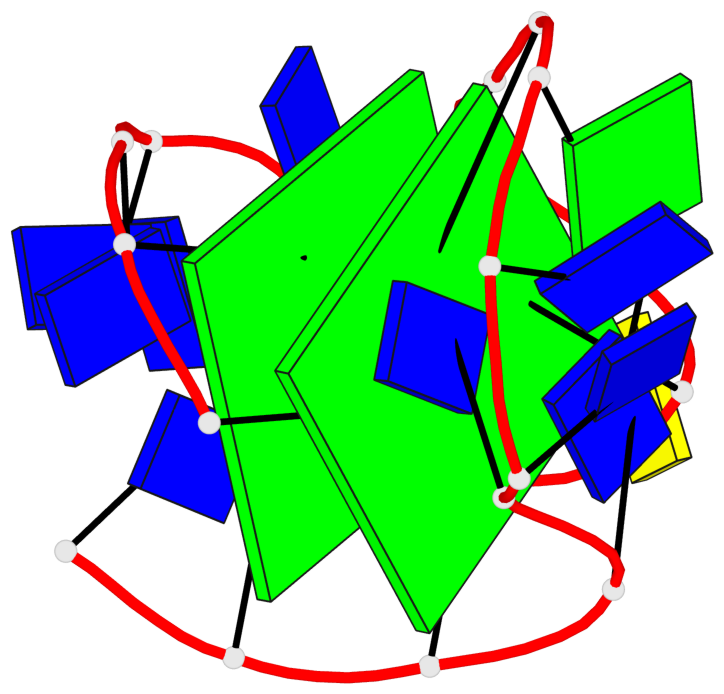
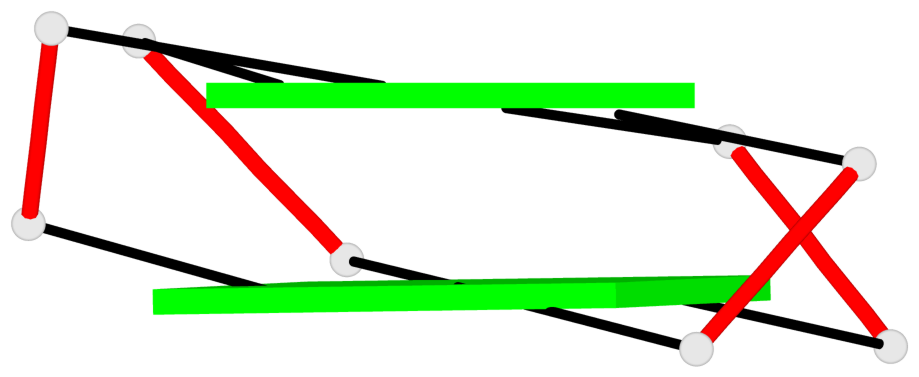
Base-block schematics in six views
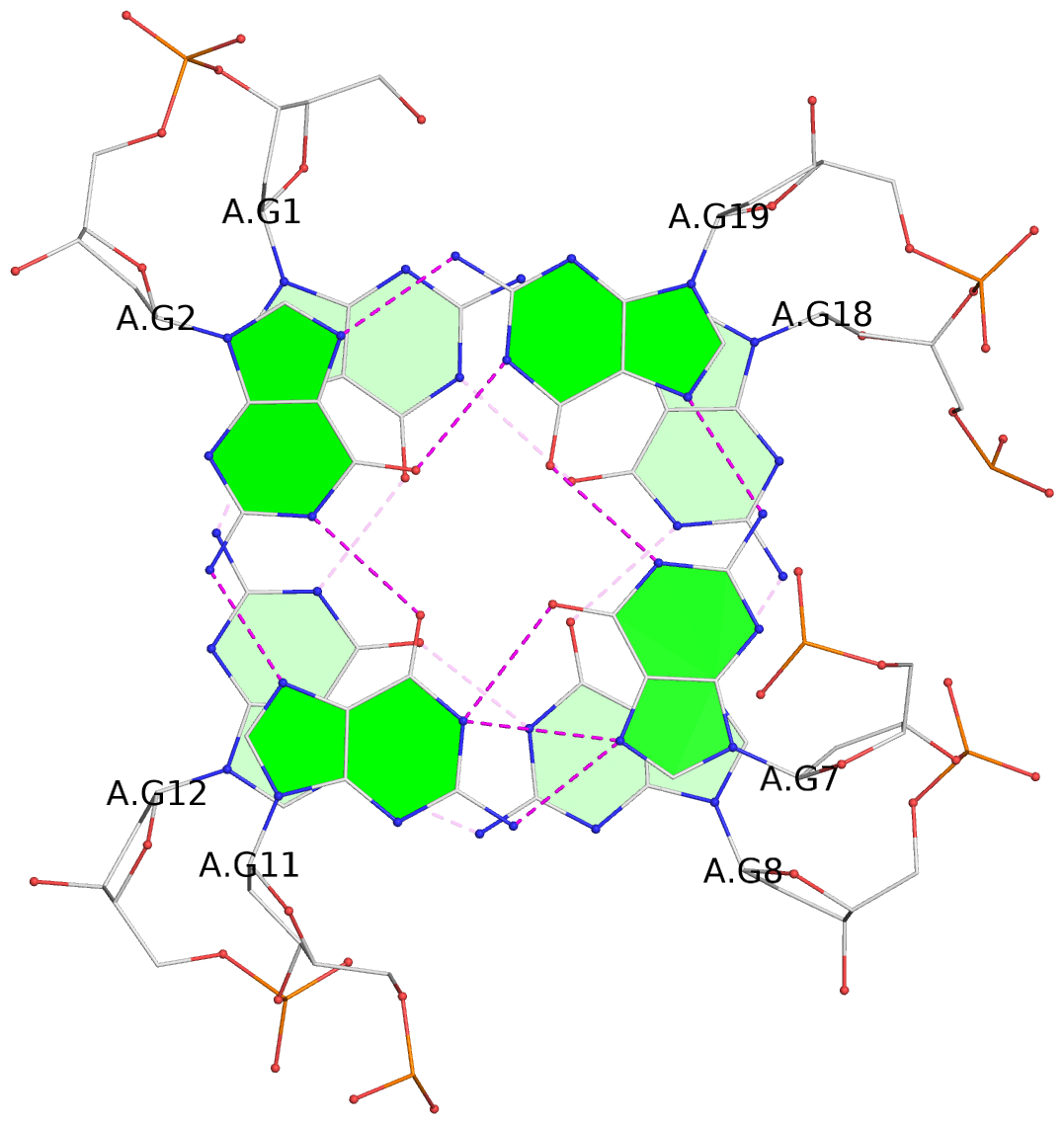
List of 2 G-tetrads
1 glyco-bond=s--s sugar=-... groove=w-n- planarity=0.130 type=planar nts=4 GGGG A.DG1,A.DG12,A.DG8,A.DG18 2 glyco-bond=-ss- sugar=-3.. groove=w-n- planarity=0.163 type=other nts=4 GGGG A.DG2,A.DG11,A.DG7,A.DG19
List of 1 G4-helix
In DSSR, a G4-helix is defined by stacking interactions of G-tetrads, regardless of backbone connectivity, and may contain more than one G4-stem.
Helix#1, 2 G-tetrad layers, INTRA-molecular, with 1 stem
List of 1 G4-stem
In DSSR, a G4-stem is defined as a G4-helix with backbone connectivity. Bulges are also allowed along each of the four strands.