Detailed DSSR results for the G-quadruplex: PDB entry 1jjp
Created and maintained by Xiang-Jun Lu <xiangjun@x3dna.org>
Citation: Please cite the NAR'20 DSSR-PyMOL schematics paper and/or the NAR'15 DSSR method paper.
Summary information
- PDB id
- 1jjp
- Class
- DNA
- Method
- NMR
- Summary
- A(gggg) pentad-containing dimeric DNA quadruplex involving stacked g(anti)g(anti)g(anti)g(syn) tetrads
- Reference
- Zhang N, Gorin A, Majumdar A, Kettani A, Chernichenko N, Skripkin E, Patel DJ (2001): "V-shaped scaffold: a new architectural motif identified in an A x (G x G x G x G) pentad-containing dimeric DNA quadruplex involving stacked G(anti) x G(anti) x G(anti) x G(syn) tetrads." J.Mol.Biol., 311, 1063-1079. doi: 10.1006/jmbi.2001.4916.
- Abstract
- We report the results of an NMR study of unlabeled and uniformly (13)C,(15)N-labeled d(G(3)AG(2)T(3)G(3)AT) in 100 mM NaCl, conditions under which it forms a dimeric quadruplex containing several new topological features. The DNA oligomer chain in each symmetry-related monomer subunit undergoes three sharp turns to form a compact domain, with all the purine bases involved in pairing alignments. The first turn is of the double chain reversal type, the second is of the edgewise type, and the third represents a new alignment, the V-shaped type. Each monomer of the dimeric quadruplex contains two stacked G(anti) x G(anti) x G(anti) x G(syn) tetrads, one of which forms a newly identified A x (G x G x G x G) pentad, through sheared G.A mismatch formation. There is a break in one of the four G-G columns that link adjacent G x G x G x G tetrads within each monomer. This architectural interruption is compensated by a new topological feature of quadruplex architecture, the V-shaped scaffold. The missing G-G column results in an opening that could facilitate insertion of planar ligands into the quadruplex. The dimeric interface contains stacked A.(G.G.G.G) pentads, with each pentad containing four bases from one monomer and a syn G1 from the partner monomer. Several potential ligand-binding pockets, positioned towards either end of the folded architecture, were identifiable in a surface view of the solution structure of the dimeric d(G(3)AG(2)T(3)G(3)AT) quadruplex.
- G4 notes
- 4 G-tetrads, 1 G4 helix
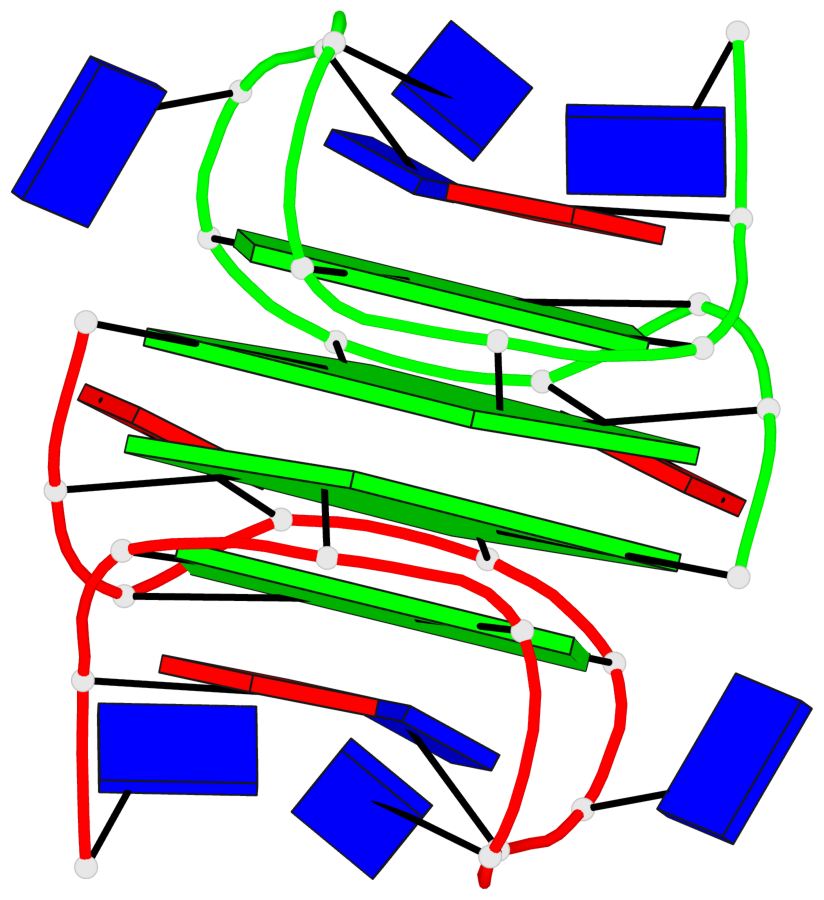
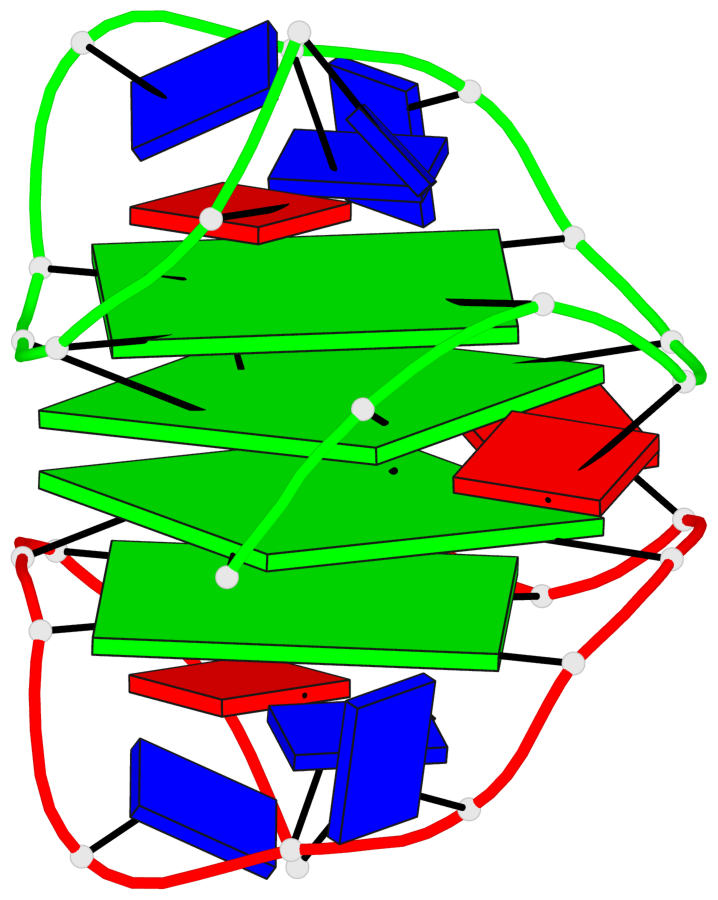
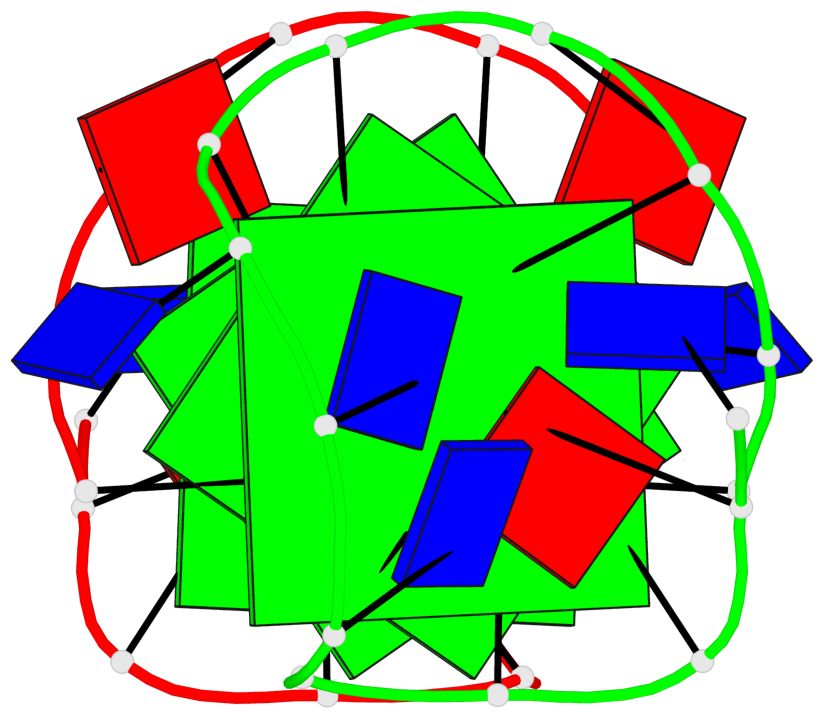
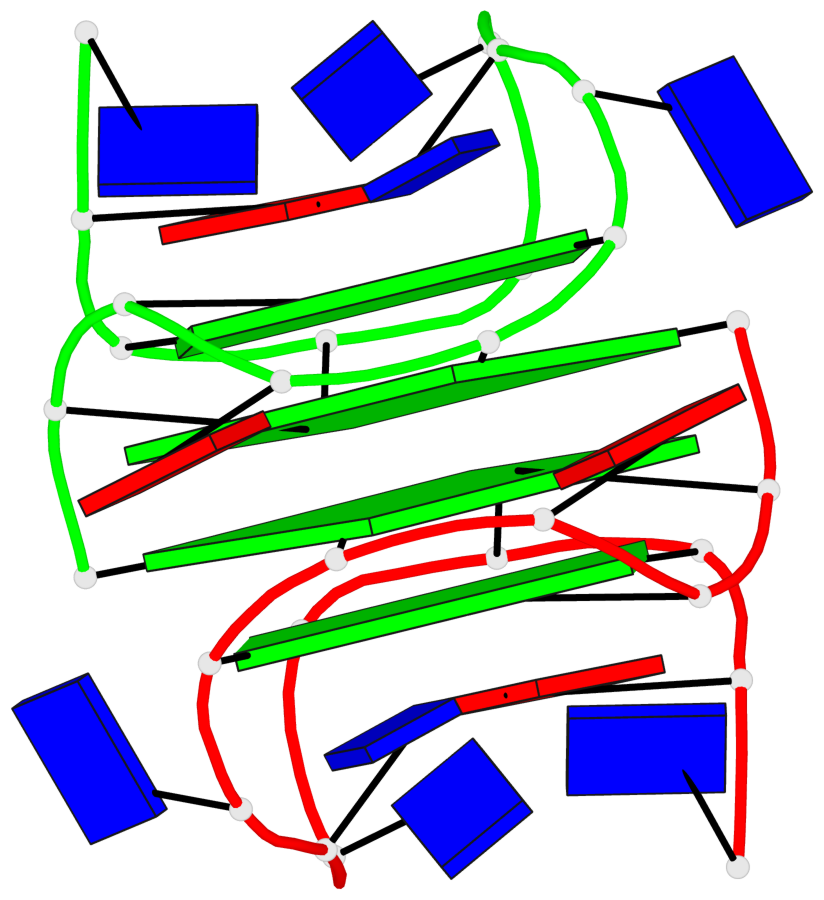
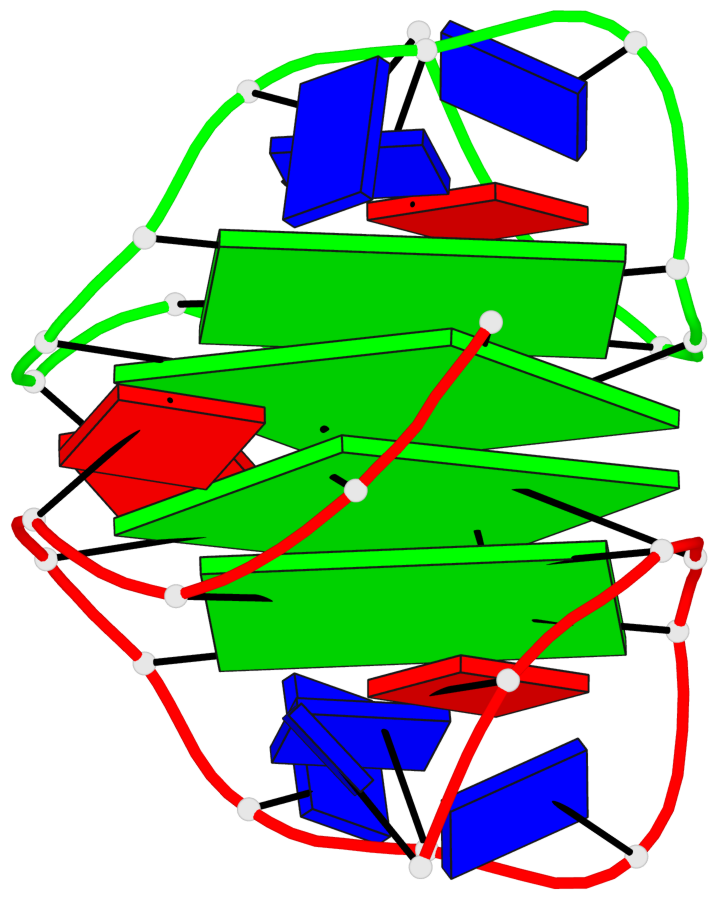
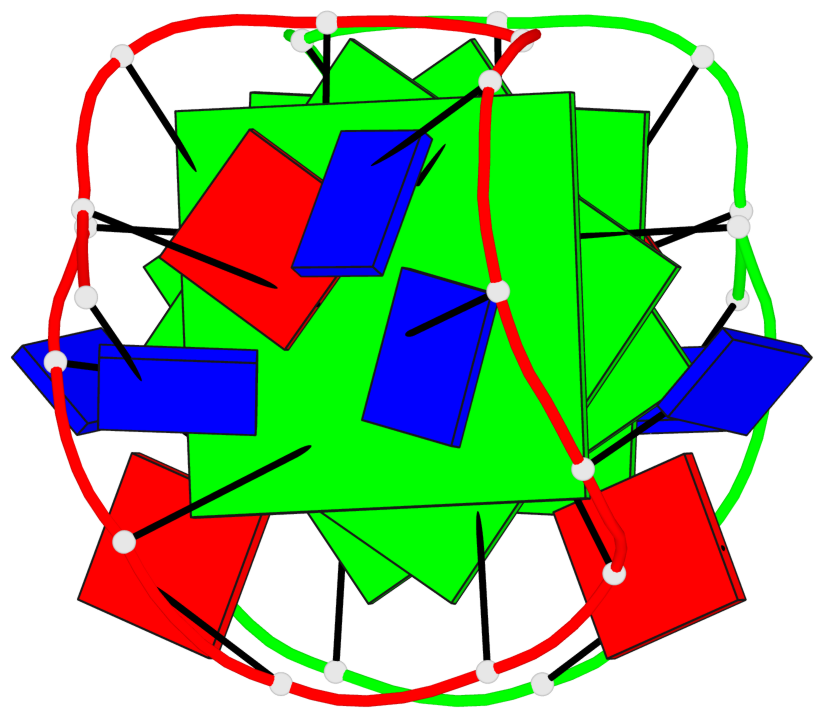
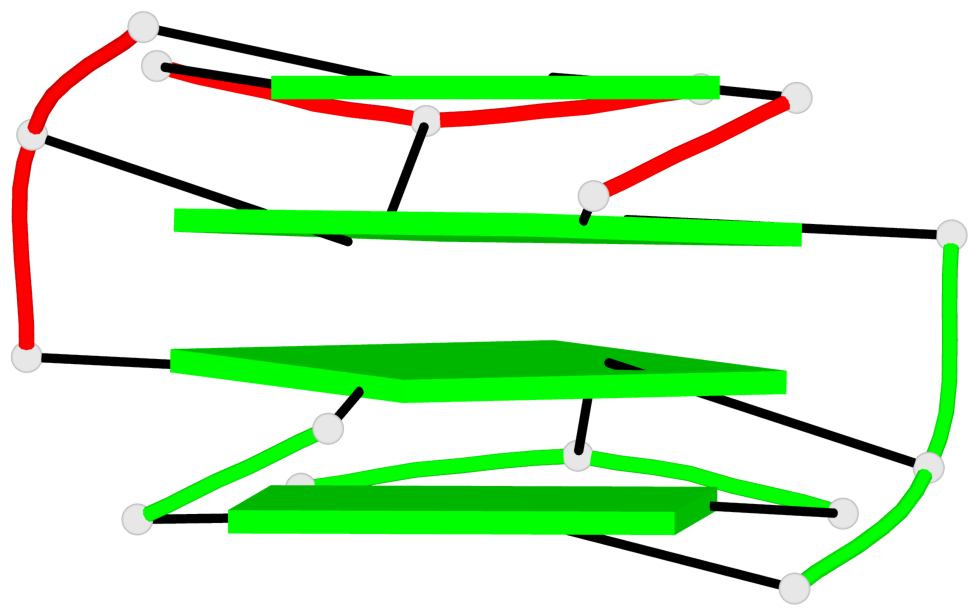
Base-block schematics in six views
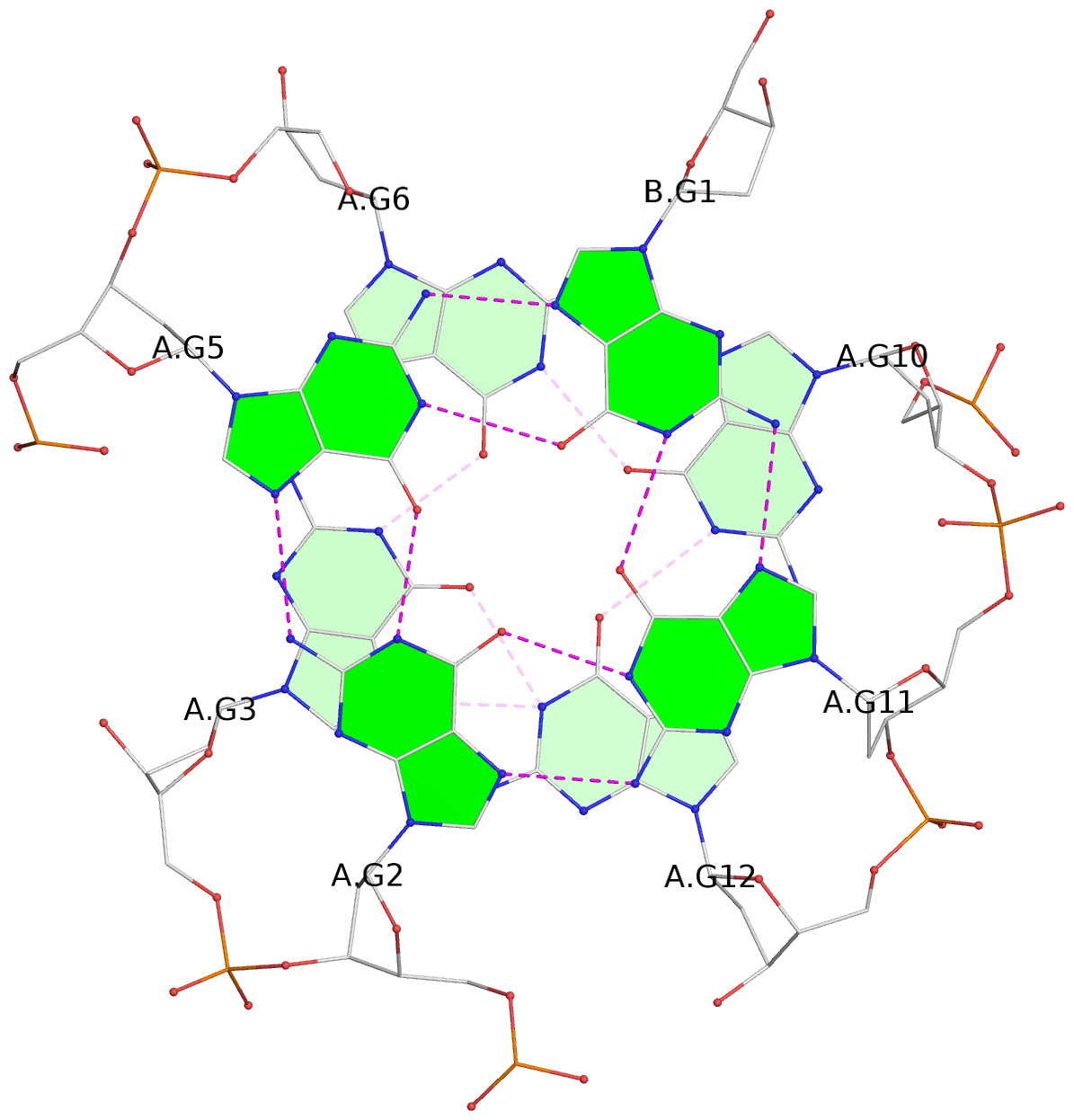
List of 4 G-tetrads
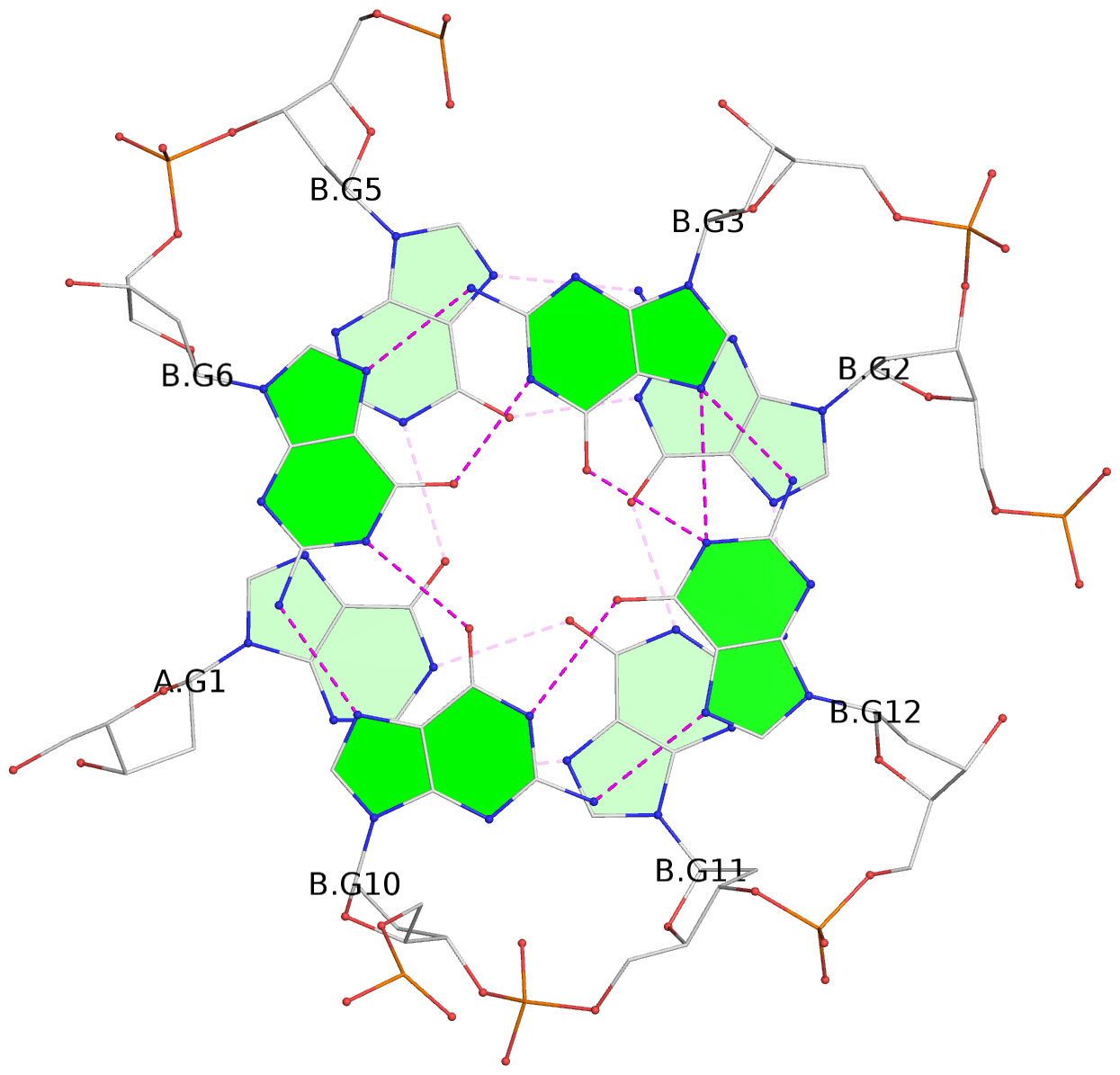
1 glyco-bond=---- sugar=.3-- groove=---- planarity=0.151 type=planar nts=4 GGGG A.DG1,B.DG11,B.DG2,B.DG5 2 glyco-bond=---- sugar=--.3 groove=---- planarity=0.150 type=planar nts=4 GGGG A.DG2,A.DG5,B.DG1,A.DG11 3 glyco-bond=--s- sugar=-.3- groove=-wn- planarity=0.191 type=other nts=4 GGGG A.DG3,A.DG6,A.DG10,A.DG12 4 glyco-bond=--s- sugar=-.3- groove=-wn- planarity=0.191 type=other nts=4 GGGG B.DG3,B.DG6,B.DG10,B.DG12
List of 1 G4-helix
In DSSR, a G4-helix is defined by stacking interactions of G-tetrads, regardless of backbone connectivity, and may contain more than one G4-stem.
Helix#1, 4 G-tetrad layers, inter-molecular
List of 4 non-stem G4-loops (including the two closing Gs)
1 type=lateral helix=#1 nts=5 GTTTG A.DG6,A.DT7,A.DT8,A.DT9,A.DG10 2 type=V-shaped helix=#1 nts=3 GGG A.DG10,A.DG11,A.DG12 3 type=lateral helix=#1 nts=5 GTTTG B.DG6,B.DT7,B.DT8,B.DT9,B.DG10 4 type=V-shaped helix=#1 nts=3 GGG B.DG10,B.DG11,B.DG12