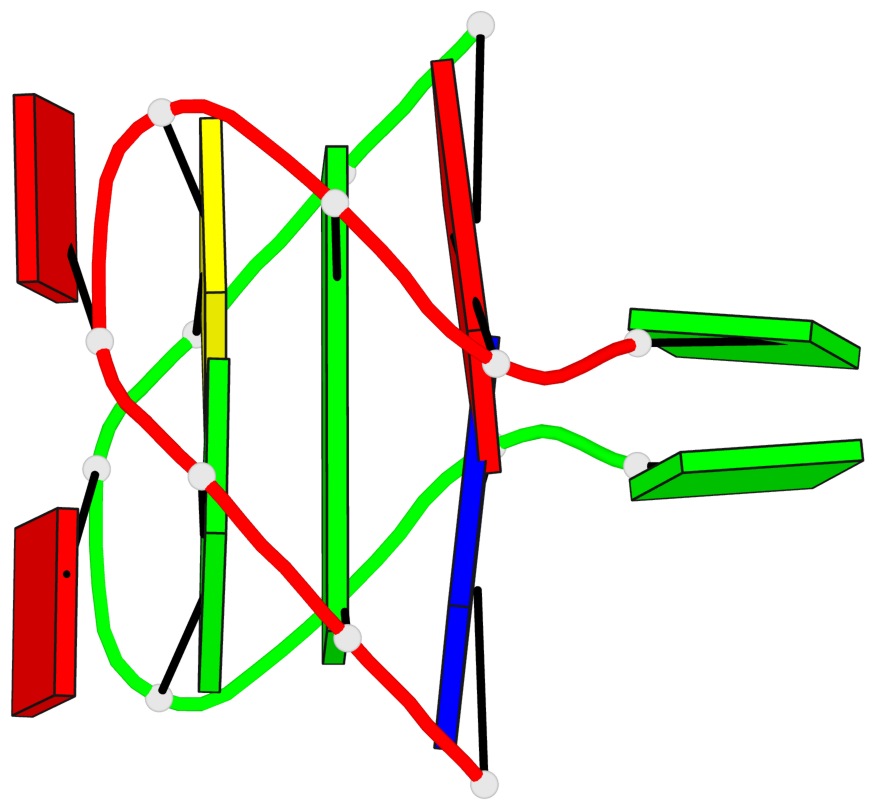
Detailed DSSR results for the G-quadruplex: PDB entry 1jvc
Created and maintained by Xiang-Jun Lu <xiangjun@x3dna.org>
Citation: Please cite the NAR'20 DSSR-PyMOL schematics paper and/or the NAR'15 DSSR method paper.
Summary information
- PDB id
- 1jvc
- Class
- DNA
- Method
- NMR
- Summary
- Dimeric DNA quadruplex containing major groove-aligned a.t.a.t and g.c.g.c tetrads stabilized by inter-subunit watson-crick a:t and g:c pairs
- Reference
- Zhang N, Gorin A, Majumdar A, Kettani A, Chernichenko N, Skripkin E, Patel DJ (2001): "Dimeric DNA quadruplex containing major groove-aligned A-T-A-T and G-C-G-C tetrads stabilized by inter-subunit Watson-Crick A-T and G-C pairs." J.Mol.Biol., 312, 1073-1088. doi: 10.1006/jmbi.2001.5002.
- Abstract
- We report on an NMR study of unlabeled and uniformly 13C,15N-labeled d(GAGCAGGT) sequence in 1 M NaCl solution, conditions under which it forms a head-to-head dimeric quadruplex containing sequentially stacked G-C-G-C, G-G-G-G and A-T-A-T tetrads. We have identified, for the first time, a slipped A-T-A-T tetrad alignment, involving recognition of Watson-Crick A-T pairs along the major groove edges of opposing adenine residues. Strikingly, both Watson-Crick G-C and A-T pairings within the direct G-C-G-C and slipped A-T-A-T tetrads, respectively, occur between rather than within hairpin subunits of the dimeric d(GAGCAGGT) quadruplex. The hairpin turns in the head-to-head dimeric quadruplex involve single adenine residues and adds to our knowledge of chain reversal involving edgewise loops in DNA quadruplexes. Our structural studies, together with those from other laboratories, definitively establish that DNA quadruplex formation is not restricted to G(n) repeat sequences, with their characteristic stacked uniform G-G-G-G tetrad architectures. Rather, the quadruplex fold is a more versatile and robust architecture, accessible to a range of mixed sequences, with the potential to facilitate G-C-G-C and A-T-A-T tetrad through major and minor groove alignment, in addition to G-G-G-G tetrad formation. The definitive experimental identification of such major groove-aligned mixed A-T-A-T and G-C-G-C tetrads within a quadruplex scaffold, has important implications for the potential alignment of duplex segments during homologous recombination.
- G4 notes
- 1 G-tetrad
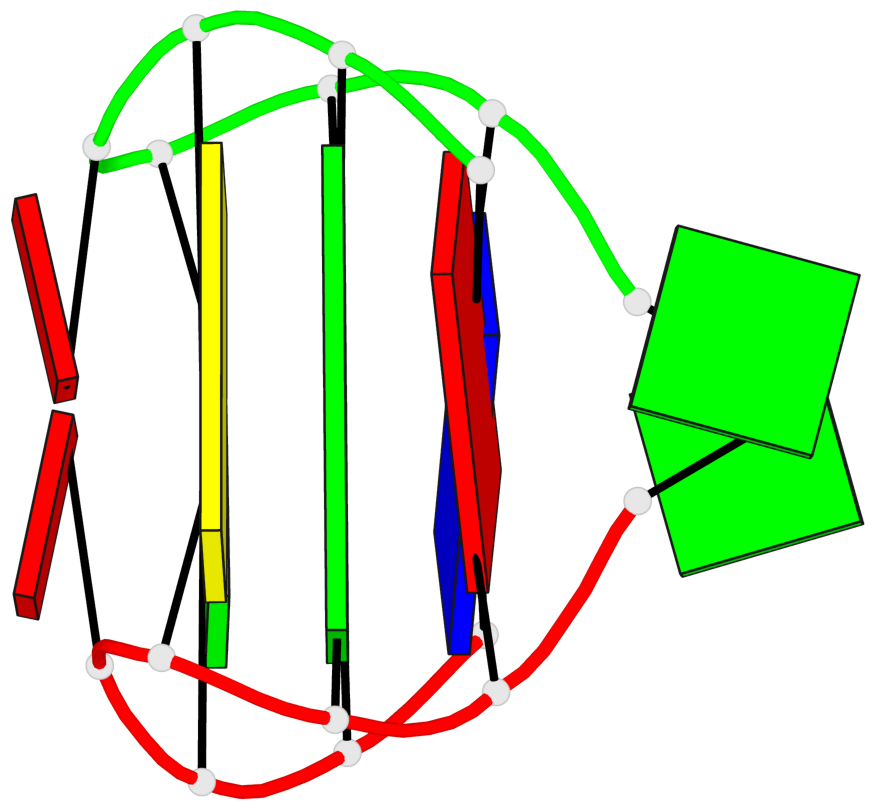
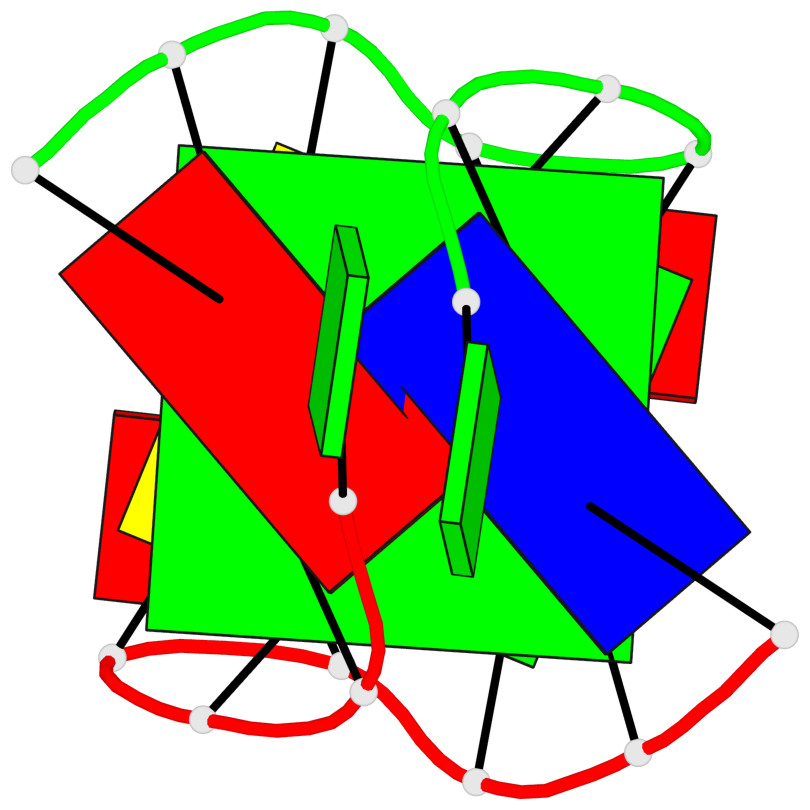
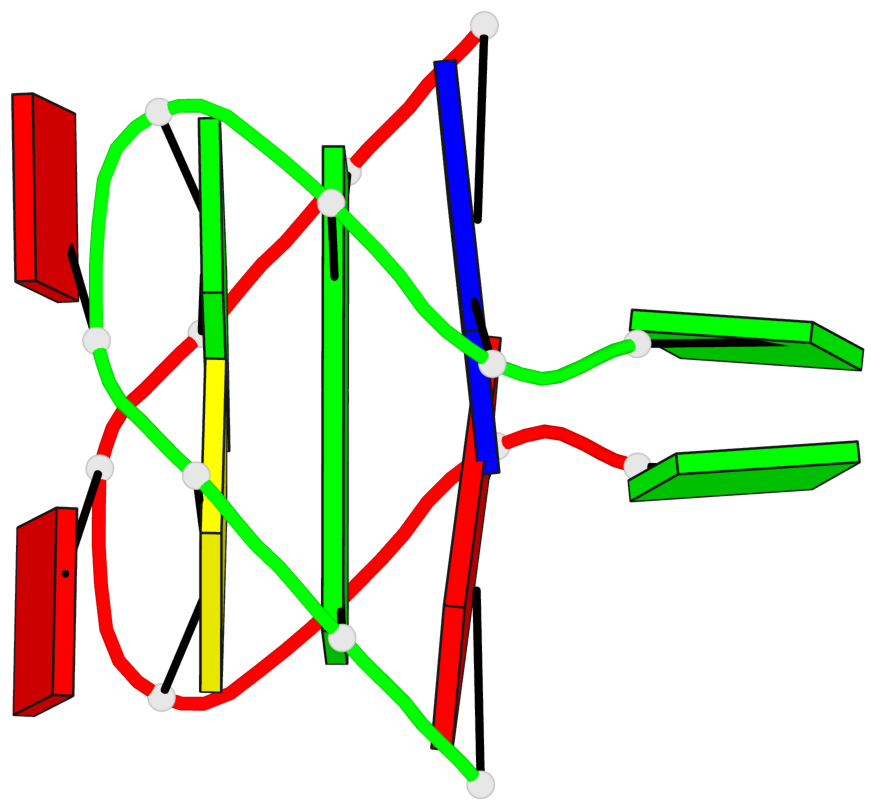
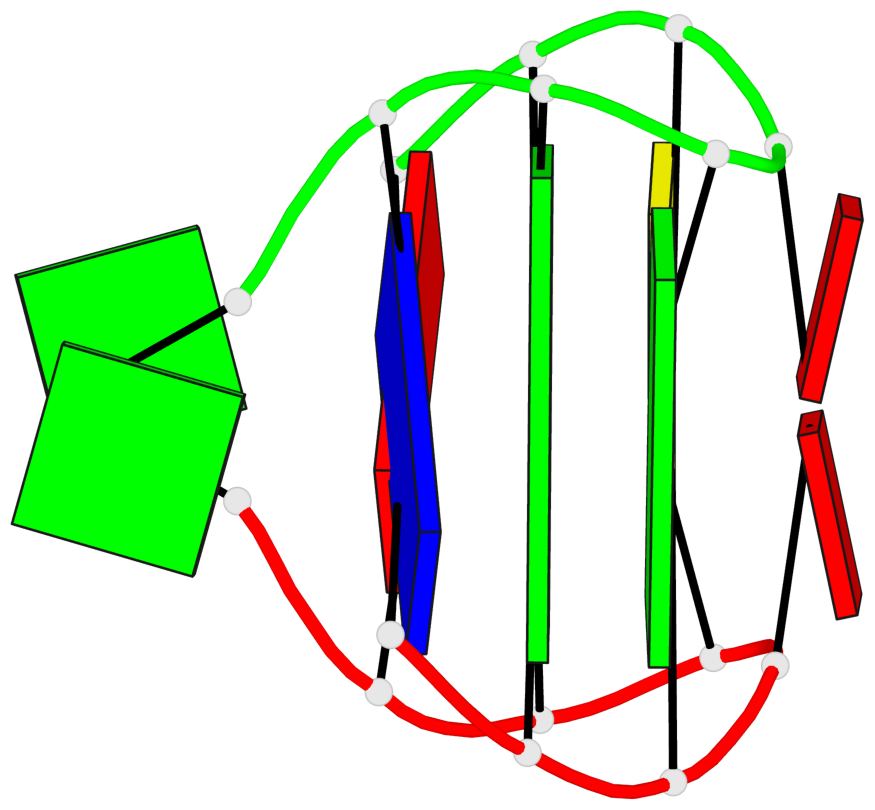
Base-block schematics in six views
List of 1 G-tetrad
1 glyco-bond=s-s- sugar=.-.- groove=wnwn planarity=0.157 type=planar nts=4 GGGG A.DG3,B.DG7,B.DG3,A.DG7
List of 2 non-stem G4-loops (including the two closing Gs)
1 type=lateral helix=#-1 nts=5 GCAGG A.DG3,A.DC4,A.DA5,A.DG6,A.DG7 2 type=lateral helix=#-1 nts=5 GCAGG B.DG3,B.DC4,B.DA5,B.DG6,B.DG7