Detailed DSSR results for the G-quadruplex: PDB entry 1rau
Created and maintained by Xiang-Jun Lu <xiangjun@x3dna.org>
Citation: Please cite the NAR'20 DSSR-PyMOL schematics paper and/or the NAR'15 DSSR method paper.
Summary information
- PDB id
- 1rau
- Class
- RNA
- Method
- NMR
- Summary
- Solution structure of an unusually stable RNA tetraplex containing g-and u-quartet structures
- Reference
- Cheong C, Moore PB (1992): "Solution structure of an unusually stable RNA tetraplex containing G- and U-quartet structures." Biochemistry, 31, 8406-8414. doi: 10.1021/bi00151a003.
- Abstract
- A model for the solution structure of an RNA tetraplex, (rUGGGGU)4, has been obtained by two-dimensional NMR spectroscopy and molecular dynamics. The molecule is parallel stranded and Hoogsteen base-paired in 50 mM KCl, and it is so stable that three of its six imino protons have exchange half-lives measured in days at 40 degrees C. The tetraplex is stabilized by base stacking and by the hydrogen bonds in four G quartets and at least one U quartet. This is the first indication of the existence of U-quartet structures of which we are aware.
- G4 notes
- 4 G-tetrads, 1 G4 helix, 1 G4 stem, parallel(4+0), UUUU
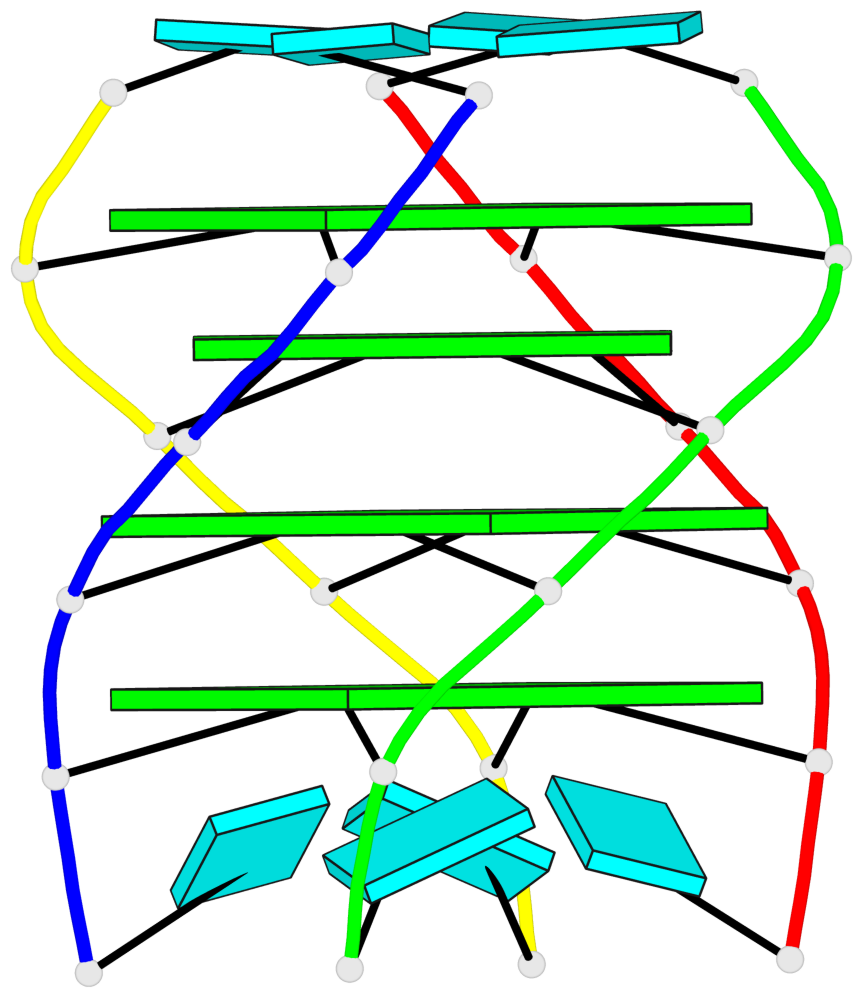
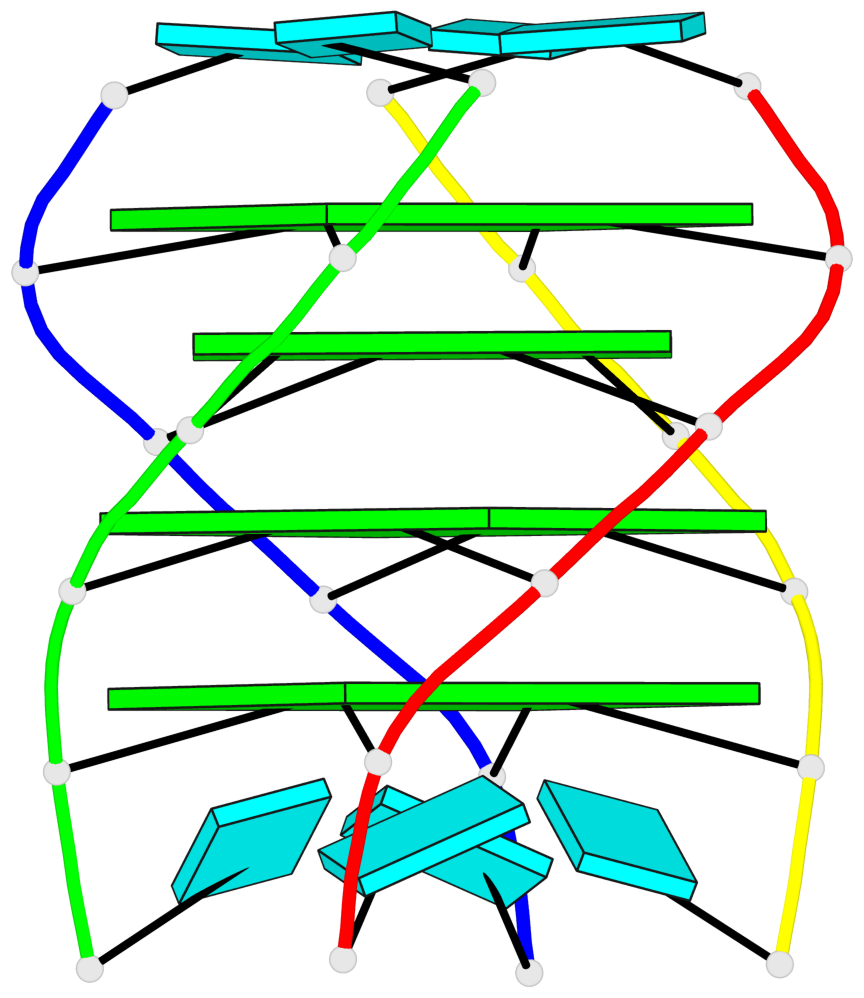
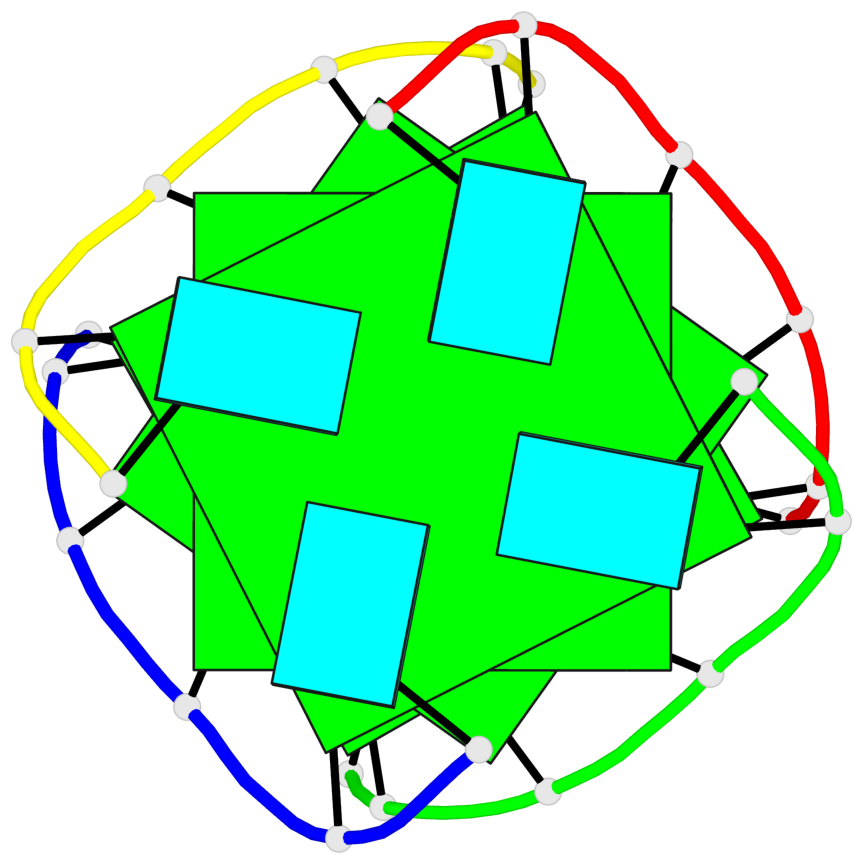
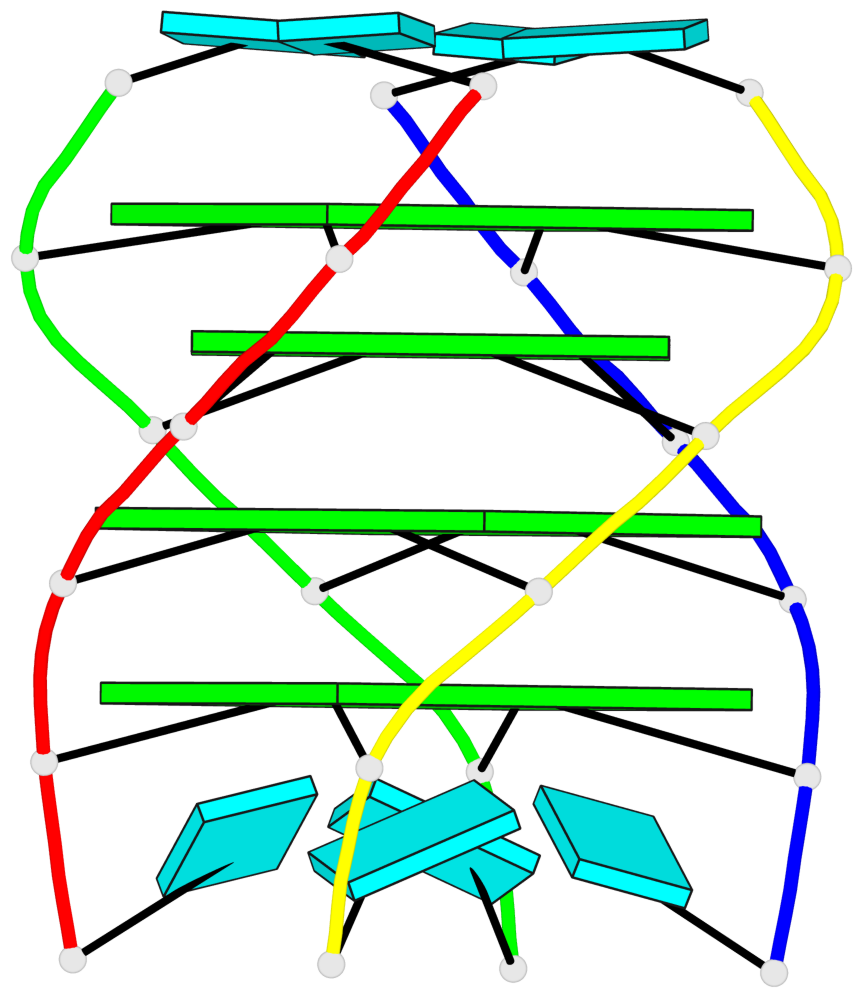
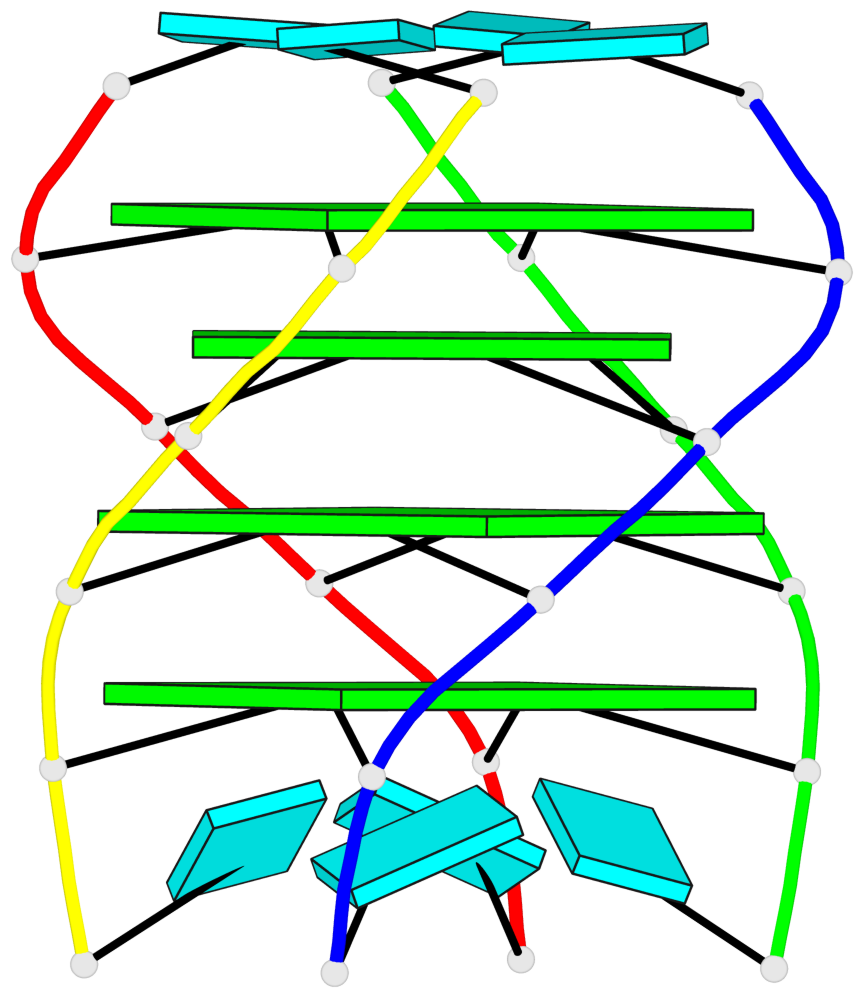
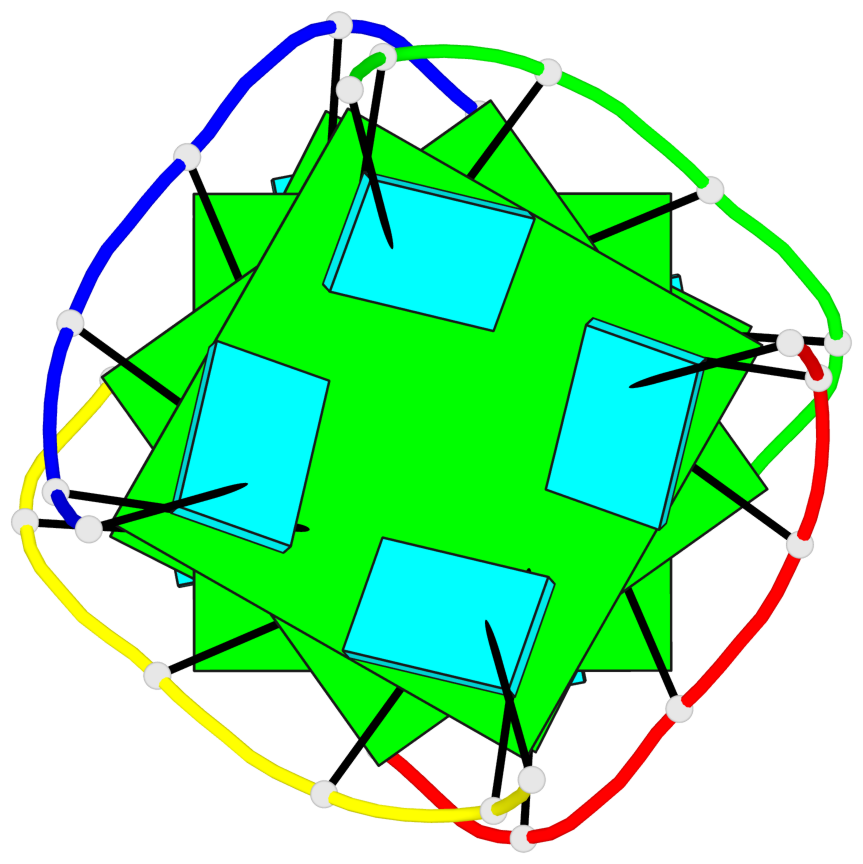
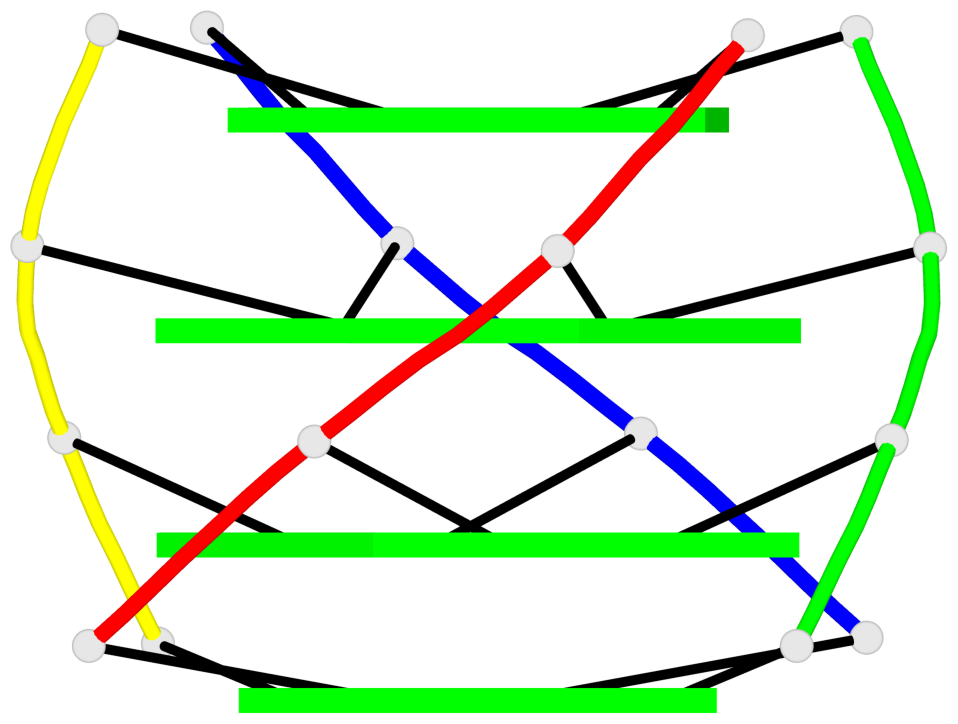
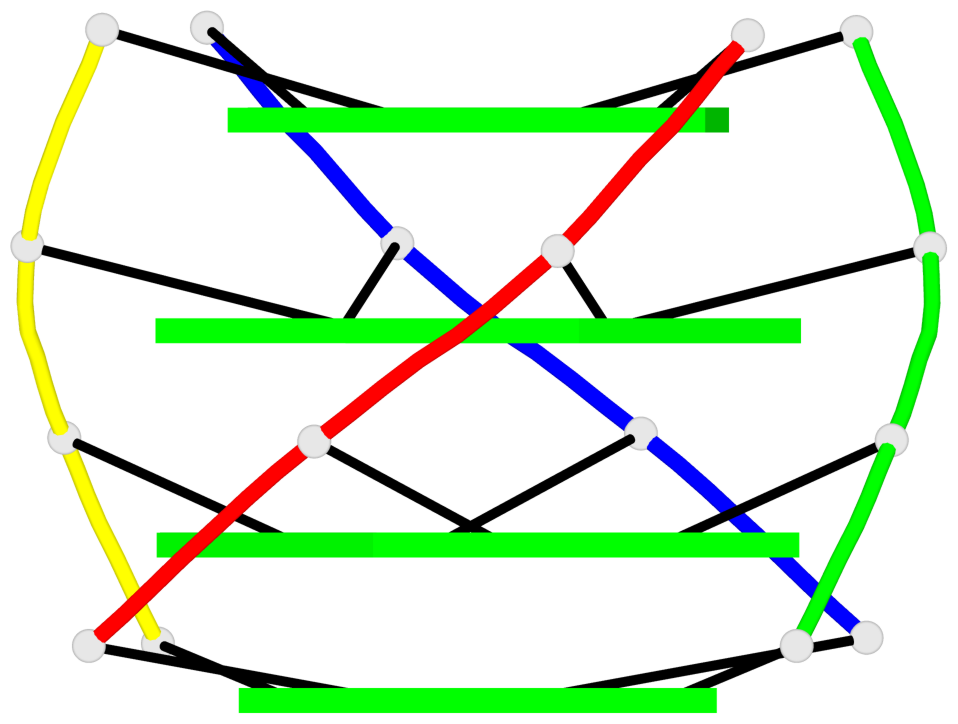
Base-block schematics in six views
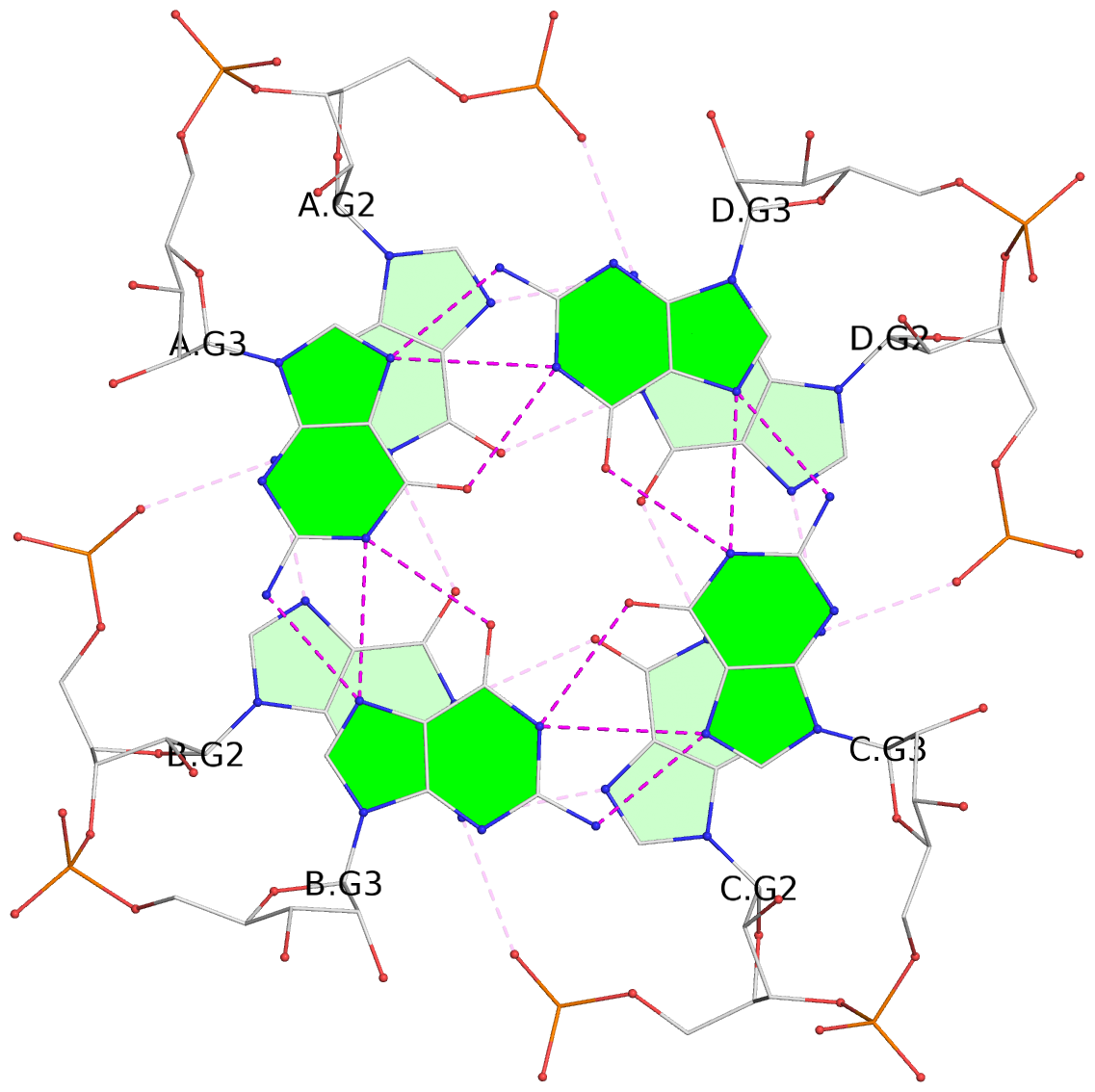
List of 4 G-tetrads
1 glyco-bond=---- sugar=---- groove=---- planarity=0.105 type=planar nts=4 GGGG A.G2,B.G2,C.G2,D.G2 2 glyco-bond=---- sugar=3333 groove=---- planarity=0.056 type=planar nts=4 GGGG A.G3,B.G3,C.G3,D.G3 3 glyco-bond=---- sugar=3333 groove=---- planarity=0.063 type=planar nts=4 GGGG A.G4,B.G4,C.G4,D.G4 4 glyco-bond=---- sugar=3333 groove=---- planarity=0.078 type=planar nts=4 GGGG A.G5,B.G5,C.G5,D.G5
List of 1 G4-helix
In DSSR, a G4-helix is defined by stacking interactions of G-tetrads, regardless of backbone connectivity, and may contain more than one G4-stem.
Helix#1, 4 G-tetrad layers, inter-molecular, with 1 stem
List of 1 G4-stem
In DSSR, a G4-stem is defined as a G4-helix with backbone connectivity. Bulges are also allowed along each of the four strands.