Detailed DSSR results for the G-quadruplex: PDB entry 2awe
Created and maintained by Xiang-Jun Lu <xiangjun@x3dna.org>
Citation: Please cite the NAR'20 DSSR-PyMOL schematics paper and/or the NAR'15 DSSR method paper.
Summary information
- PDB id
- 2awe
- Class
- RNA
- Method
- X-ray (2.1 Å)
- Summary
- Base-tetrad swapping results in dimerization of RNA quadruplexes: implications for formation of i-motif RNA octaplex
- Reference
- Pan B, Shi K, Sundaralingam M (2006): "Base-tetrad swapping results in dimerization of RNA quadruplexes: implications for formation of the i-motif RNA octaplex." Proc.Natl.Acad.Sci.Usa, 103, 3130-3134. doi: 10.1073/pnas.0507730103.
- Abstract
- Nucleic acids adopt different multistranded helical architectures to perform various biological functions. Here, we report a crystal structure of an RNA quadruplex containing "base-tetrad swapping" and bulged nucleotide at 2.1-Angstroms resolution. The base-tetrad swapping results in a dimer of quadruplexes with an intercalated octaplex fragment at the 5' end junction. The intercalated base tetrads provide the basic repeat unit for constructing a model of intercalated RNA octaplex. The model we obtained shows fundamentally different characteristics from duplex, triplex, and quadruplex. We also observed two different orientations of bulged uridine residues that are related to the interaction with surroundings. This structural evidence reflects the conformational flexibility of bulged nucleotides in RNA quadruplexes and implies the potential roles of bulged nucleotides as recognition and interaction sites in RNA-protein and RNA-RNA interactions.
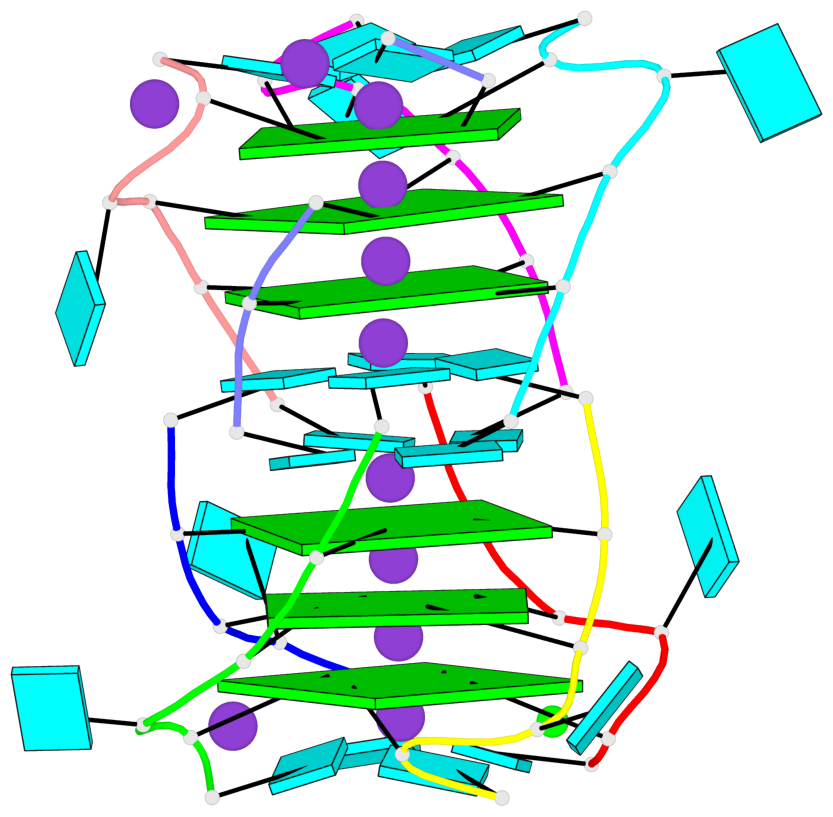
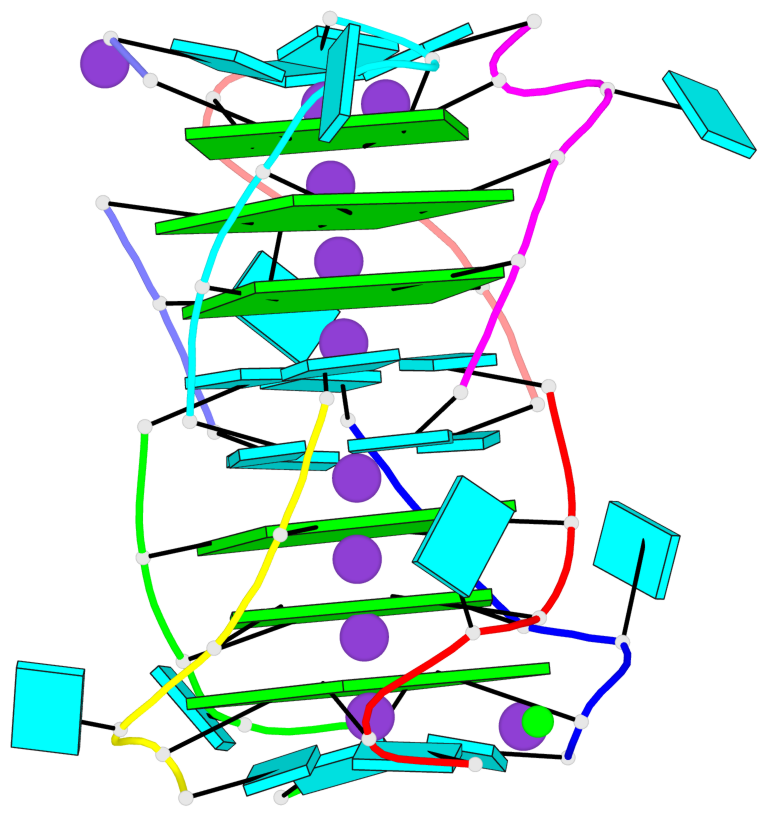
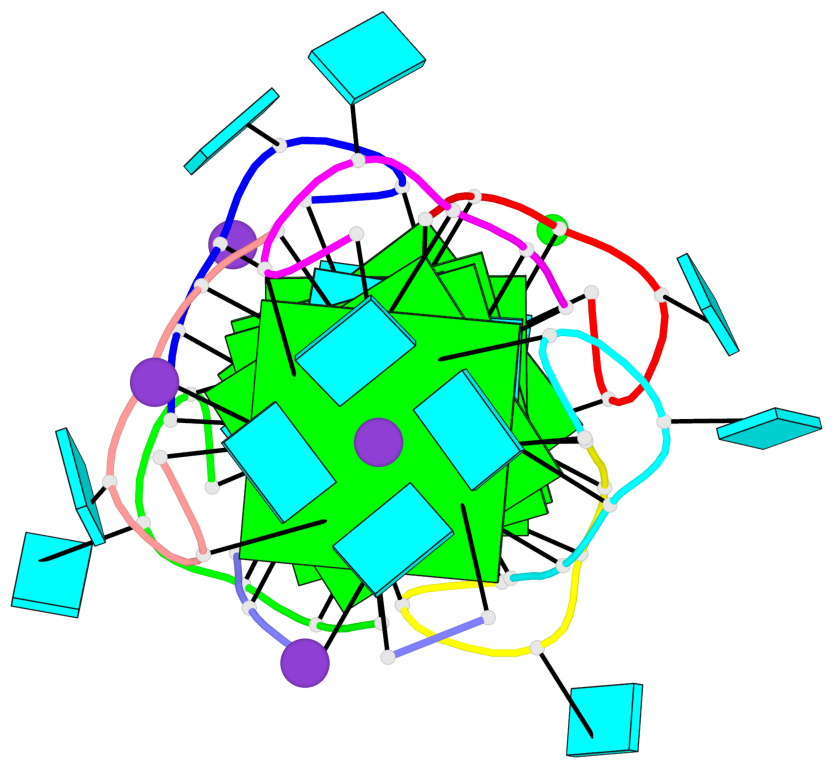
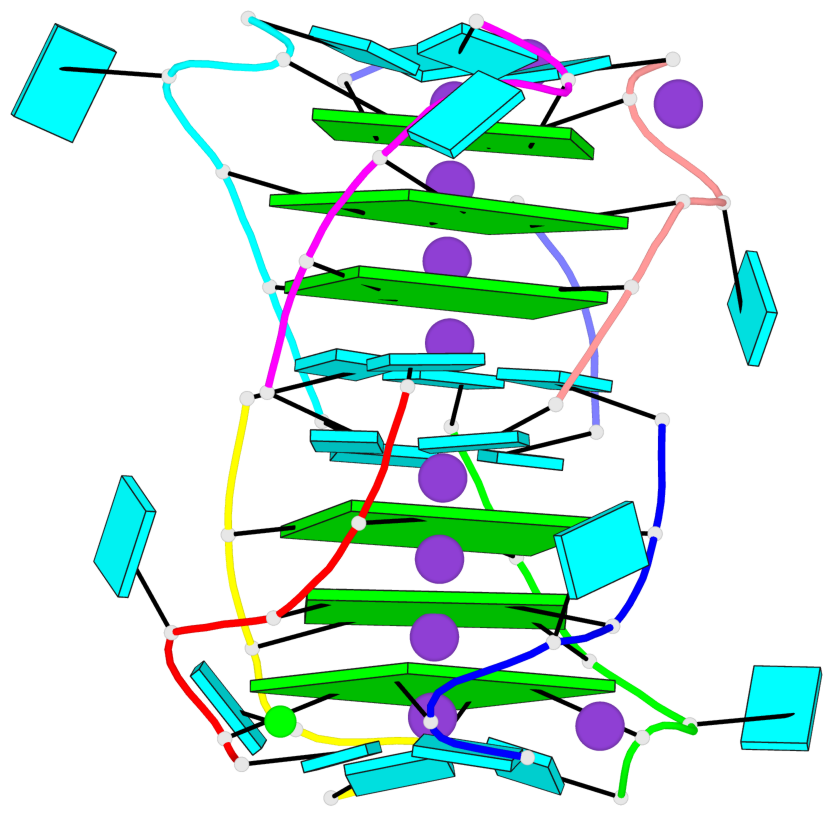
- G4 notes
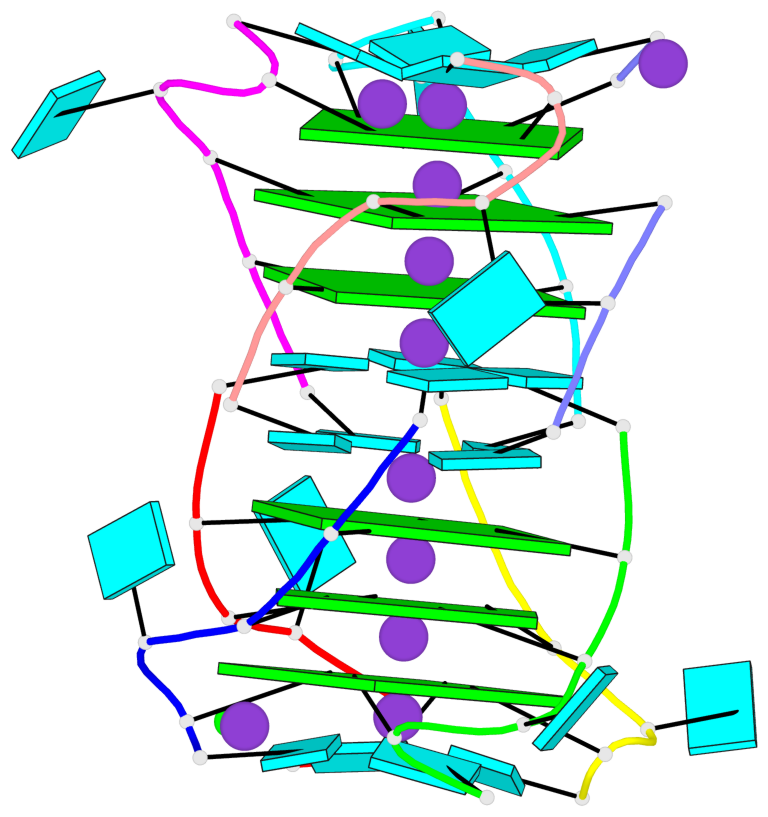
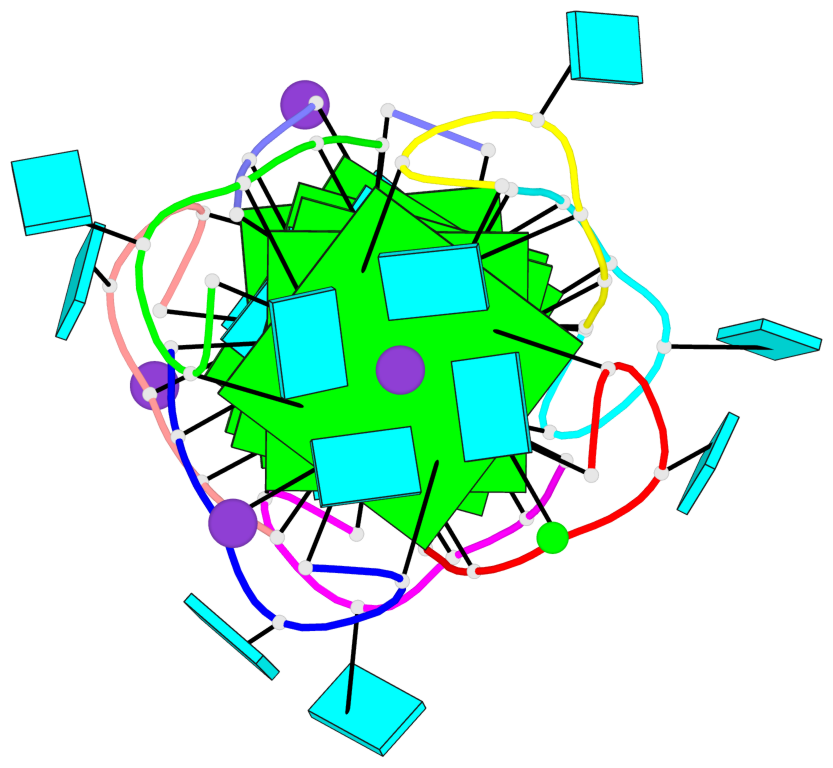
- 6 G-tetrads, 2 G4 helices, 2 G4 stems, parallel(4+0), UUUU
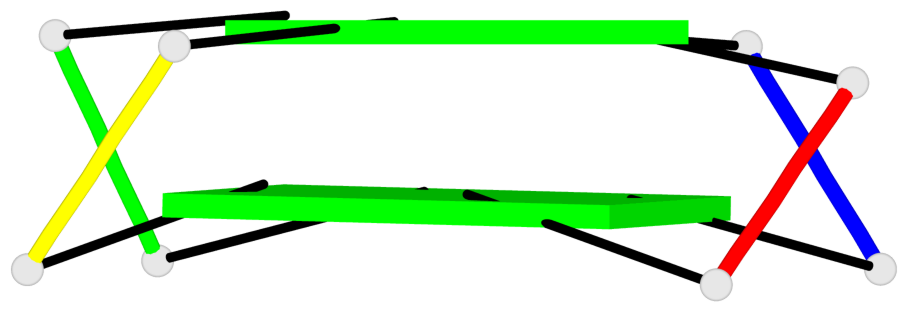
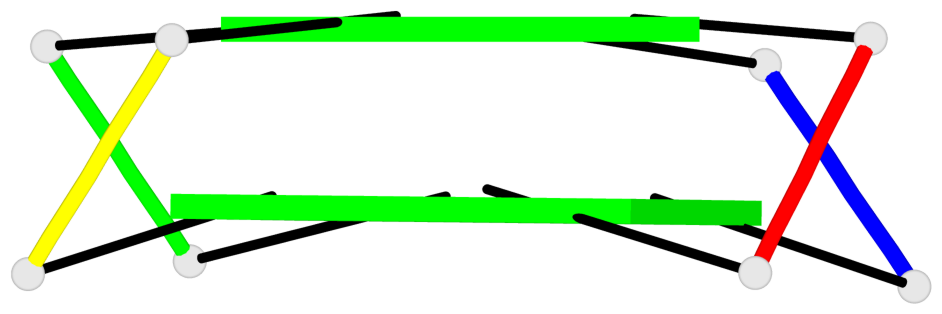
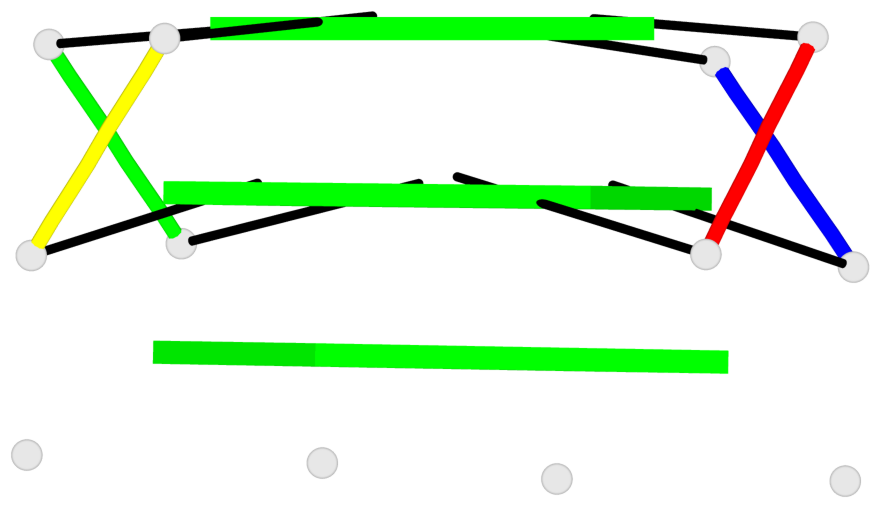
Base-block schematics in six views
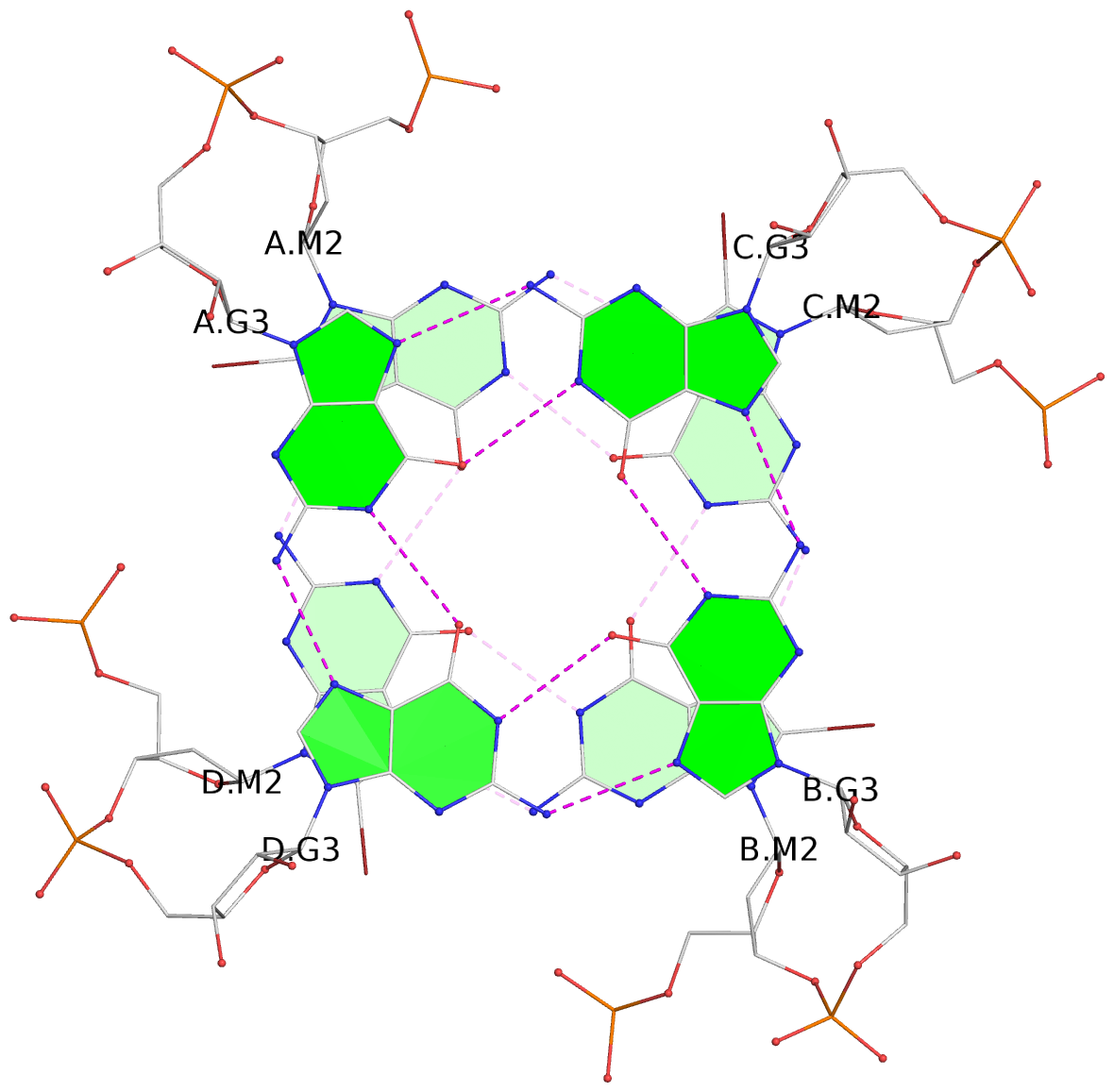
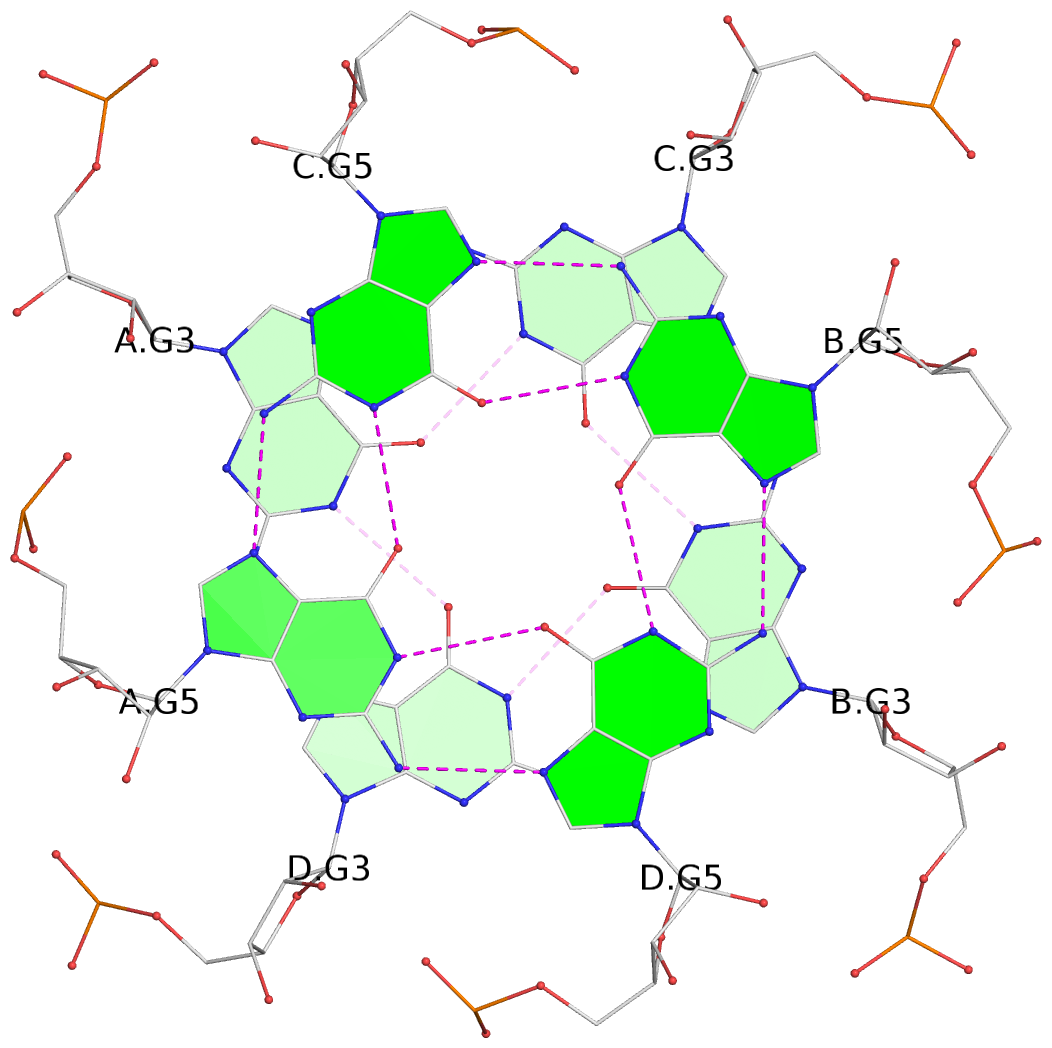
List of 6 G-tetrads
1 glyco-bond=ssss sugar=---- groove=---- planarity=0.163 type=other nts=4 gggg A.BGM2,D.BGM2,B.BGM2,C.BGM2 2 glyco-bond=---- sugar=-.-- groove=---- planarity=0.255 type=bowl nts=4 GGGG A.G3,D.G3,B.G3,C.G3 3 glyco-bond=---- sugar=3333 groove=---- planarity=0.268 type=bowl nts=4 GGGG A.G5,D.G5,B.G5,C.G5 4 glyco-bond=ssss sugar=---- groove=---- planarity=0.176 type=other nts=4 gggg E.BGM2,H.BGM2,F.BGM2,G.BGM2 5 glyco-bond=---- sugar=---- groove=---- planarity=0.259 type=bowl nts=4 GGGG E.G3,H.G3,F.G3,G.G3 6 glyco-bond=---- sugar=3333 groove=---- planarity=0.291 type=bowl nts=4 GGGG E.G5,H.G5,F.G5,G.G5
List of 2 G4-helices
In DSSR, a G4-helix is defined by stacking interactions of G-tetrads, regardless of backbone connectivity, and may contain more than one G4-stem.
Helix#1, 3 G-tetrad layers, inter-molecular, with 1 stem
Helix#2, 3 G-tetrad layers, inter-molecular, with 1 stem
List of 2 G4-stems
In DSSR, a G4-stem is defined as a G4-helix with backbone connectivity. Bulges are also allowed along each of the four strands.