Detailed DSSR results for the G-quadruplex: PDB entry 2km3
Created and maintained by Xiang-Jun Lu <xiangjun@x3dna.org>
Citation: Please cite the NAR'20 DSSR-PyMOL schematics paper and/or the NAR'15 DSSR method paper.
Summary information
- PDB id
- 2km3
- Class
- DNA
- Method
- NMR
- Summary
- Structure of an intramolecular g-quadruplex containing a g.c.g.c tetrad formed by human telomeric variant ctaggg repeats
- Reference
- Lim KW, Alberti P, Guedin A, Lacroix L, Riou JF, Royle NJ, Mergny JL, Phan AT (2009): "Sequence variant (CTAGGG)n in the human telomere favors a G-quadruplex structure containing a G.C.G.C tetrad." Nucleic Acids Res., 37, 6239-6248. doi: 10.1093/nar/gkp630.
- Abstract
- Short contiguous arrays of variant CTAGGG repeats in the human telomere are unstable in the male germline and somatic cells, suggesting formation of unusual structures by this repeat type. Here, we report on the structure of an intramolecular G-quadruplex formed by DNA sequences containing four human telomeric variant CTAGGG repeats in potassium solution. Our results reveal a new robust antiparallel G-quadruplex fold involving two G-tetrads sandwiched between a G.C base pair and a G.C.G.C tetrad, which could represent a new platform for drug design targeted to human telomeric DNA.
- G4 notes
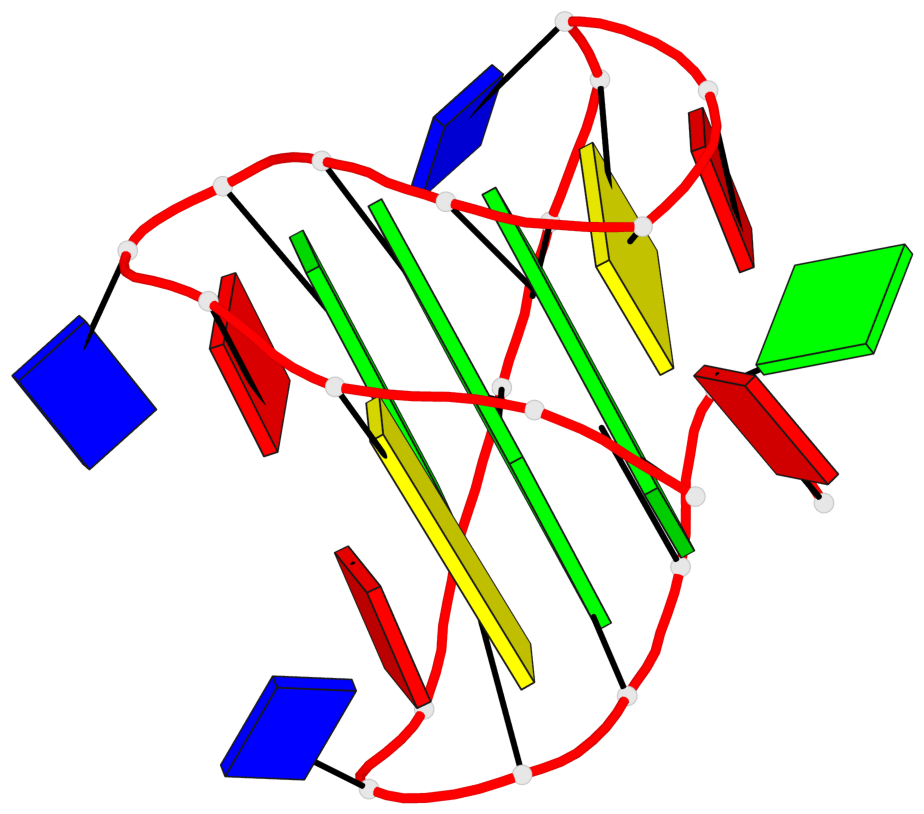
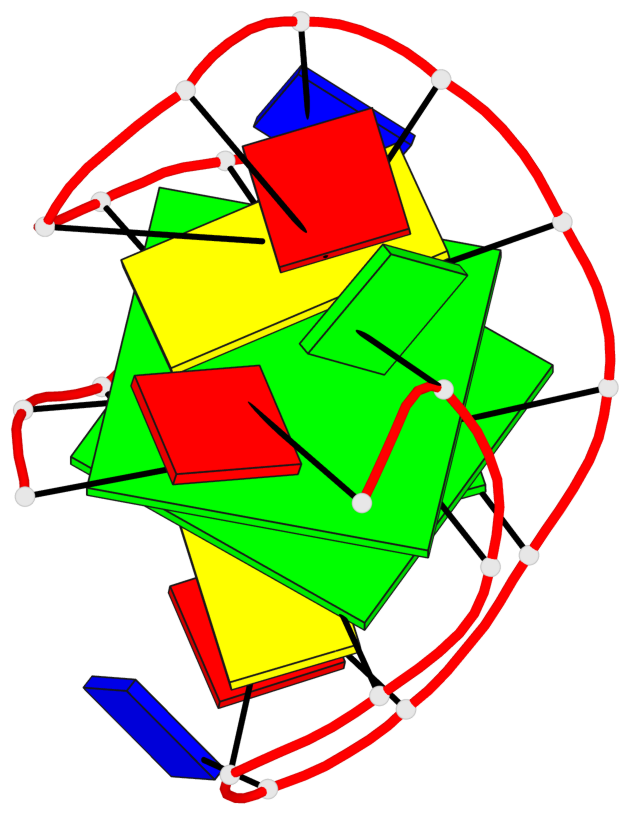
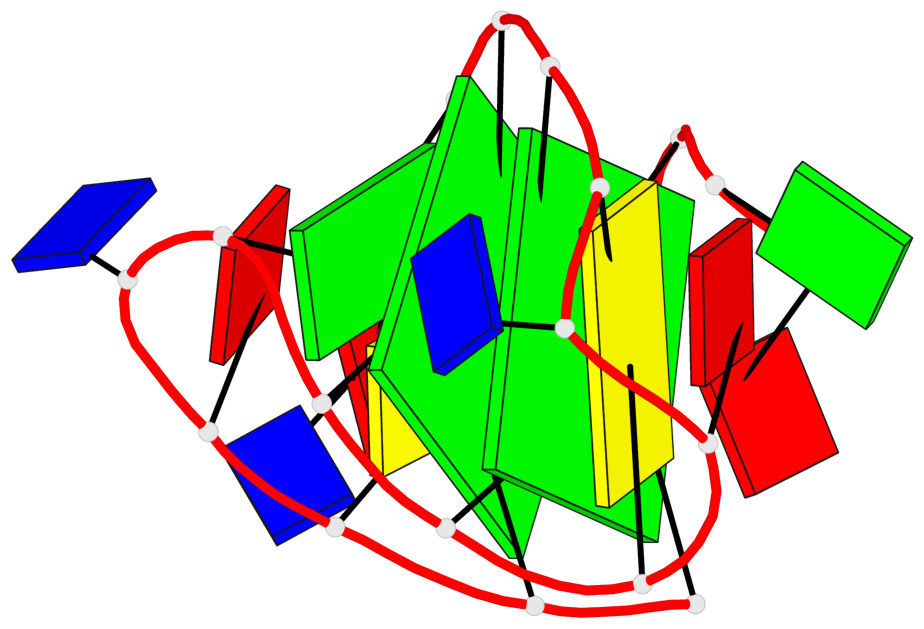
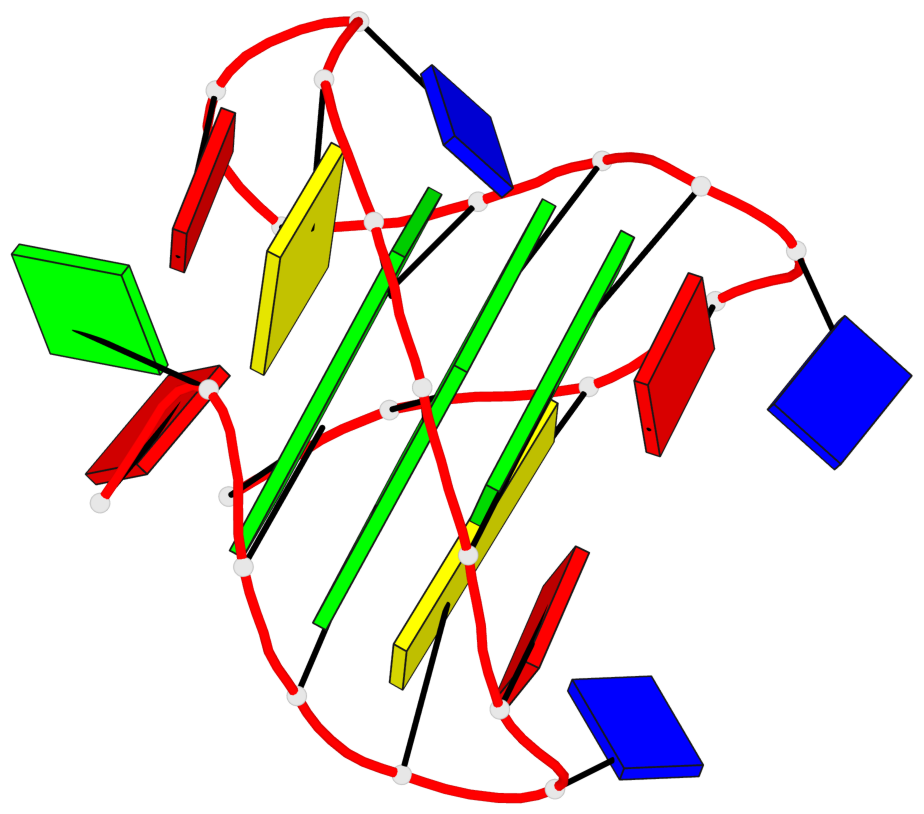
- 2 G-tetrads, 1 G4 helix, 1 G4 stem, 2(+Ln+Lw+Ln), chair(2+2), UDUD
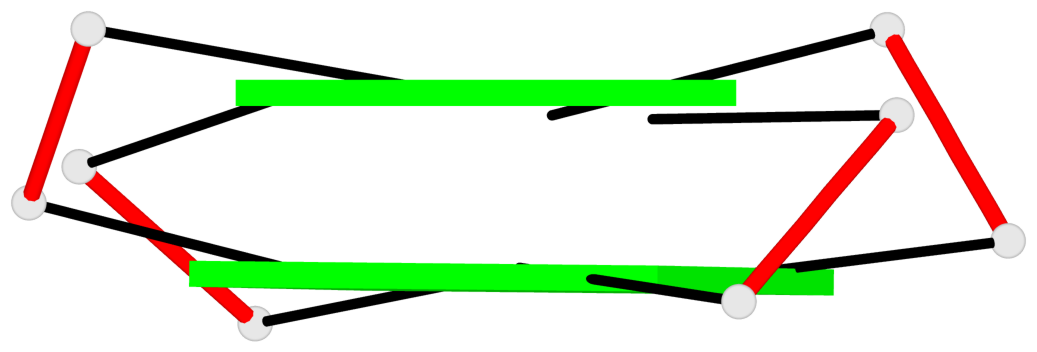
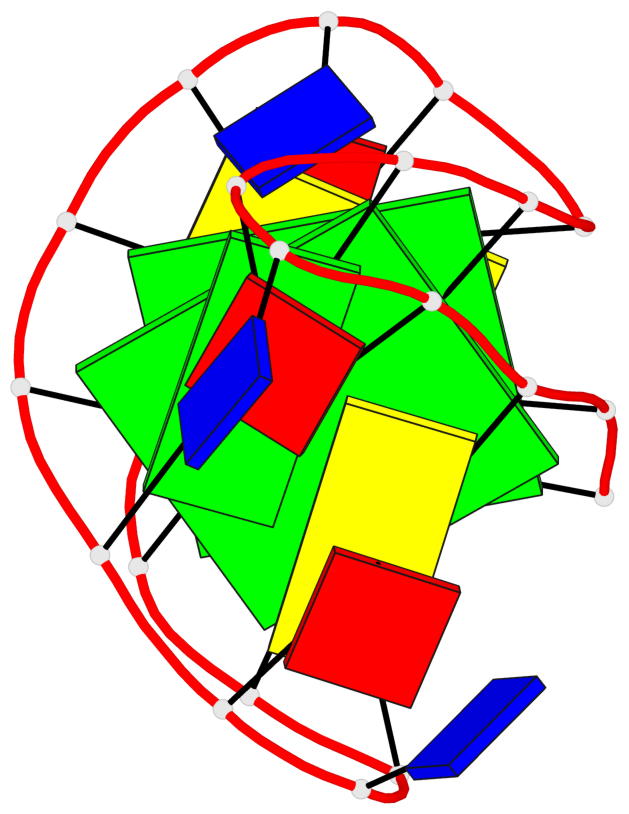
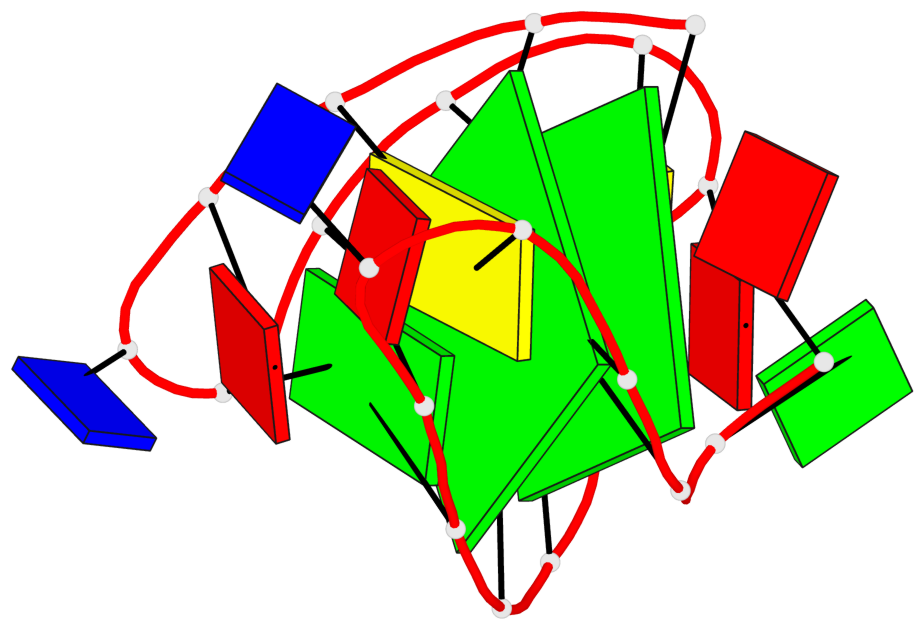
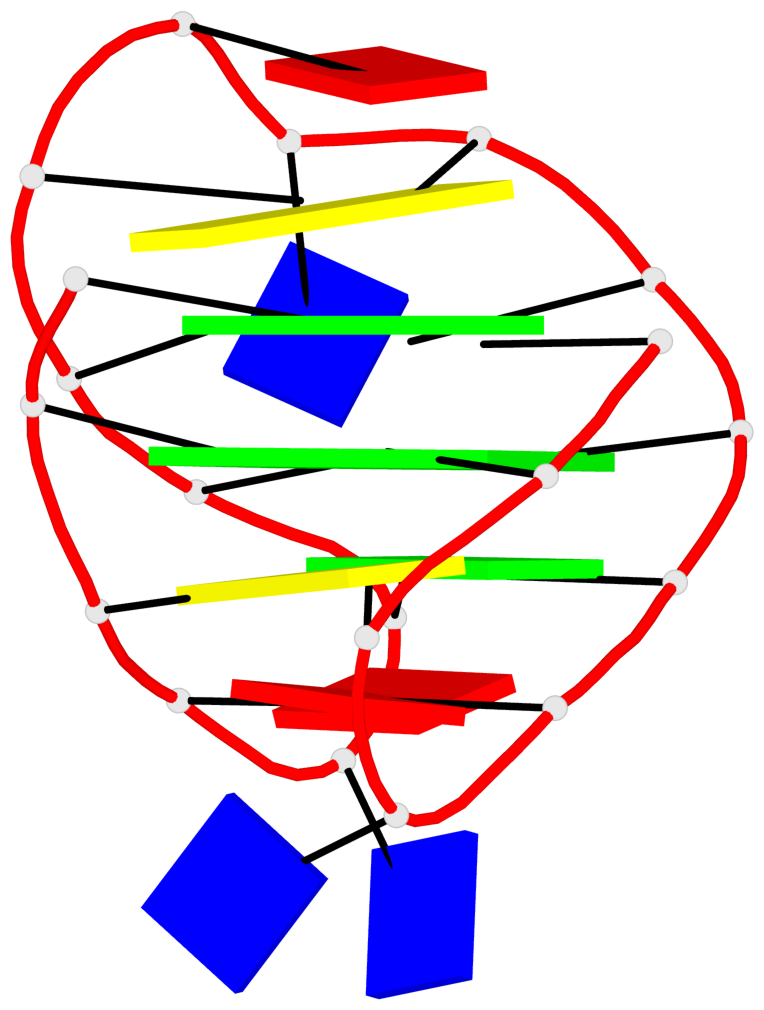
Base-block schematics in six views
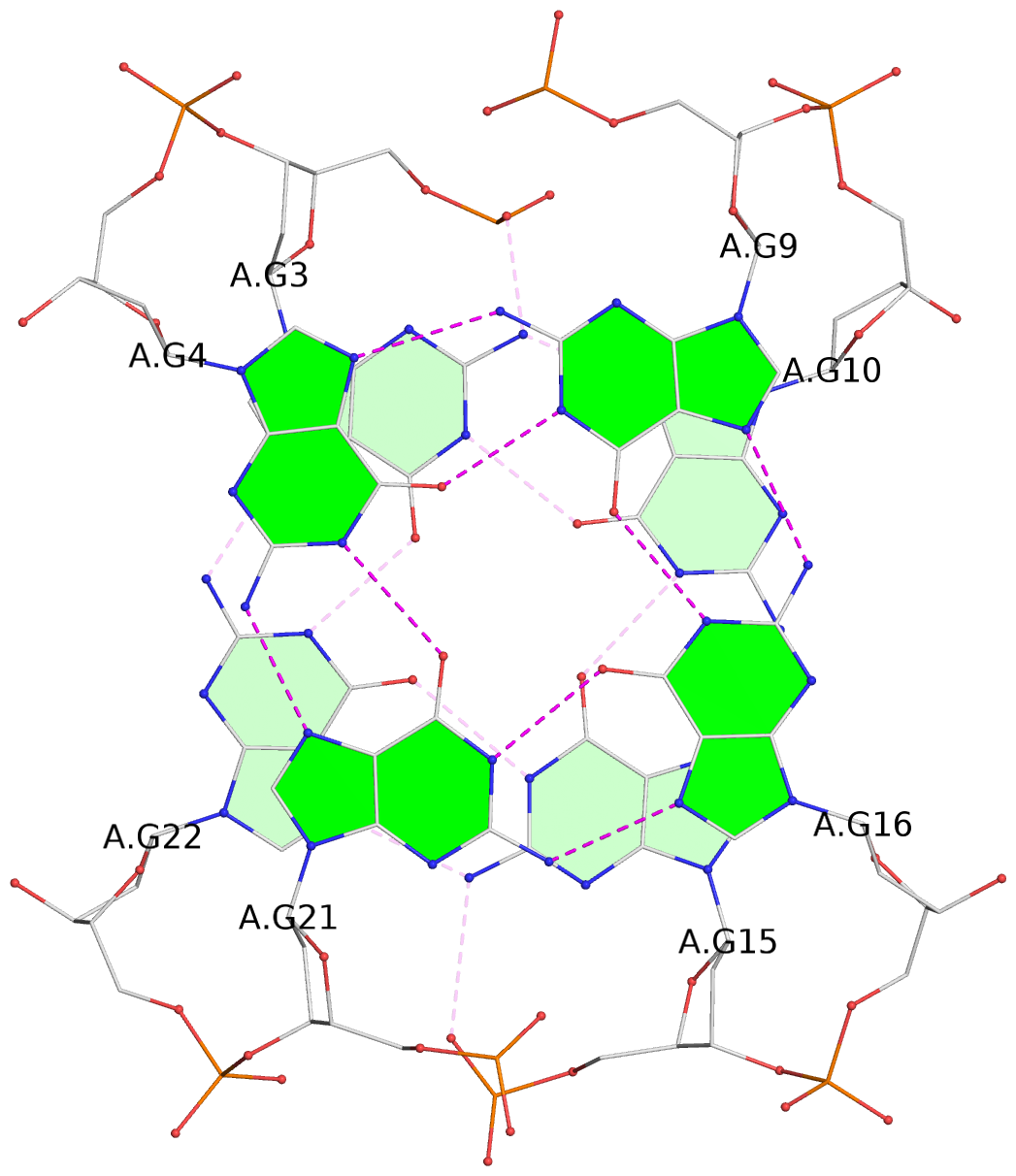
List of 2 G-tetrads
1 glyco-bond=s-s- sugar=---- groove=wnwn planarity=0.294 type=other nts=4 GGGG A.DG3,A.DG22,A.DG15,A.DG10 2 glyco-bond=-s-s sugar=---- groove=wnwn planarity=0.167 type=other nts=4 GGGG A.DG4,A.DG21,A.DG16,A.DG9
List of 1 G4-helix
In DSSR, a G4-helix is defined by stacking interactions of G-tetrads, regardless of backbone connectivity, and may contain more than one G4-stem.
Helix#1, 2 G-tetrad layers, INTRA-molecular, with 1 stem
List of 1 G4-stem
In DSSR, a G4-stem is defined as a G4-helix with backbone connectivity. Bulges are also allowed along each of the four strands.