Detailed DSSR results for the G-quadruplex: PDB entry 2kze
Created and maintained by Xiang-Jun Lu <xiangjun@x3dna.org>
Citation: Please cite the NAR'20 DSSR-PyMOL schematics paper and/or the NAR'15 DSSR method paper.
Summary information
- PDB id
- 2kze
- Class
- DNA
- Method
- NMR
- Summary
- Structure of an all-parallel-stranded g-quadruplex formed by htert promoter sequence
- Reference
- Lim KW, Lacroix L, Yue DJE, Lim JKC, Lim JMW, Phan AT (2010): "Coexistence of two distinct G-quadruplex conformations in the hTERT promoter." J.Am.Chem.Soc., 132, 12331-12342. doi: 10.1021/ja101252n.
- Abstract
- The catalytic subunit of human telomerase, hTERT, actively elongates the 3' end of the telomere in most cancer cells. The hTERT promoter, which contains many guanine-rich stretches on the same DNA strand, exhibits an exceptional potential for G-quadruplex formation. Here we show that one particular G-rich sequence in this region coexists in two G-quadruplex conformations in potassium solution: a (3 + 1) and a parallel-stranded G-quadruplexes. We present the NMR solution structures of both conformations, each comprising several robust structural elements, among which include the (3 + 1) and all-parallel G-tetrad cores, single-residue double-chain-reversal loops, and a capping A.T base pair. A combination of NMR and CD techniques, complemented with sequence modifications and variations of experimental condition, allowed us to better understand the coexistence of the two G-quadruplex conformations in equilibrium and how different structural elements conspire to favor a particular form.
- G4 notes
- 3 G-tetrads, 1 G4 helix, 1 G4 stem, 3(-P-P-P), parallel(4+0), UUUU
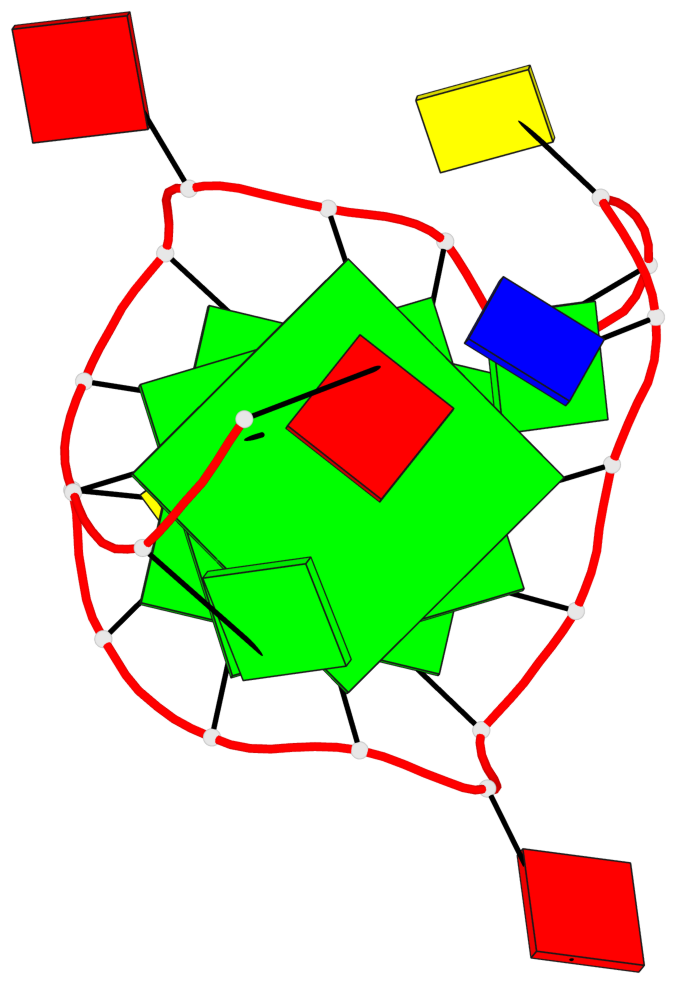
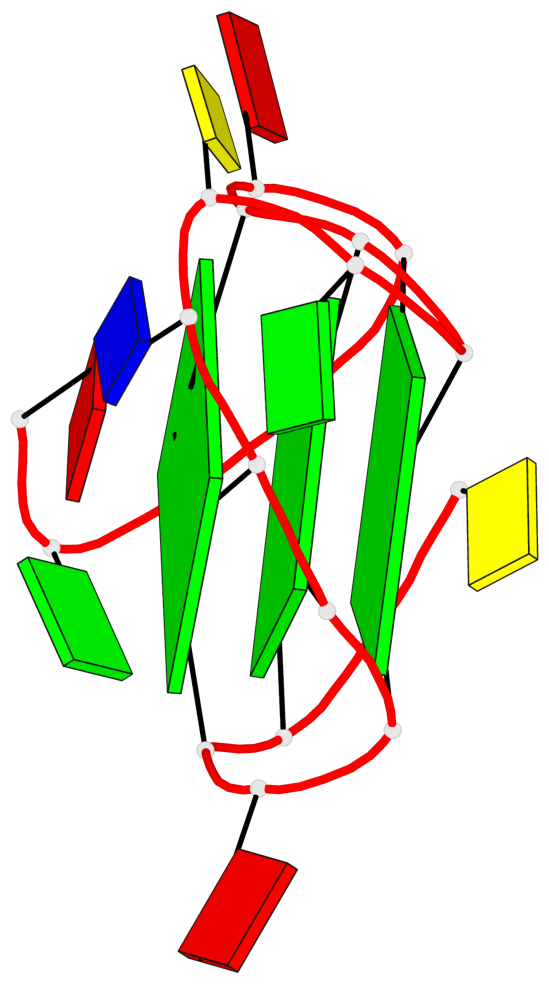
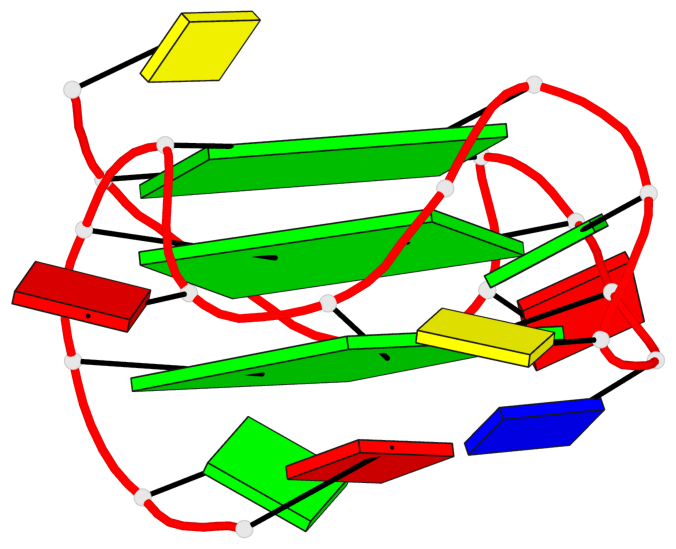
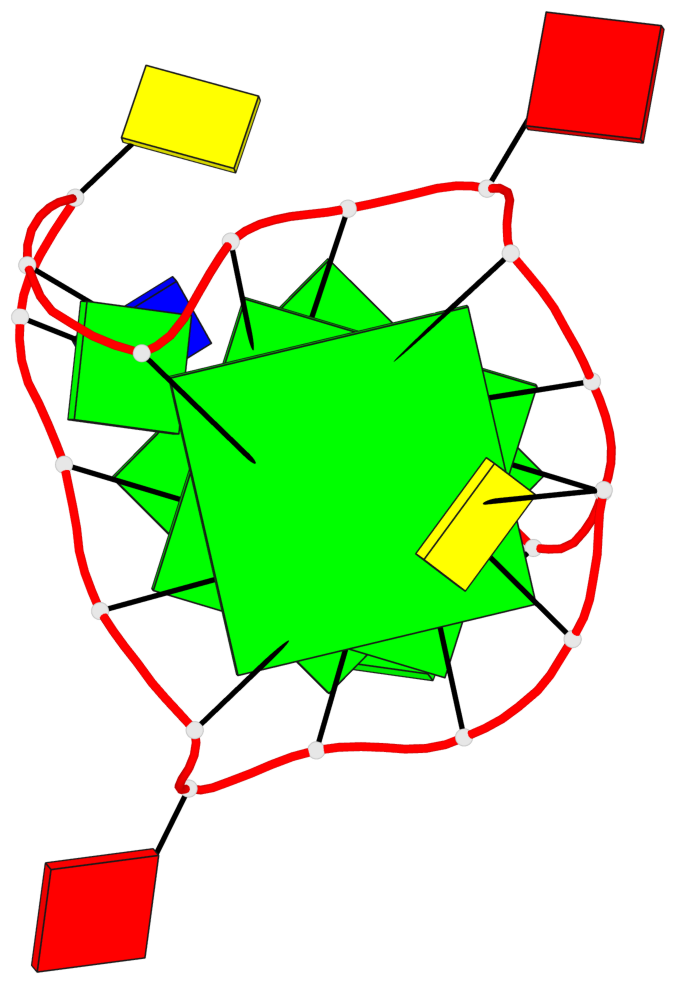
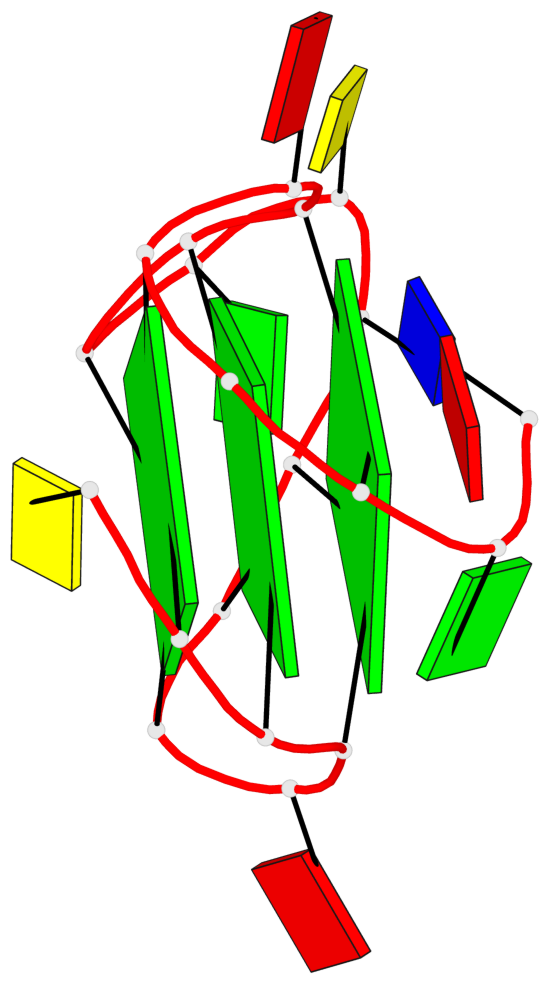
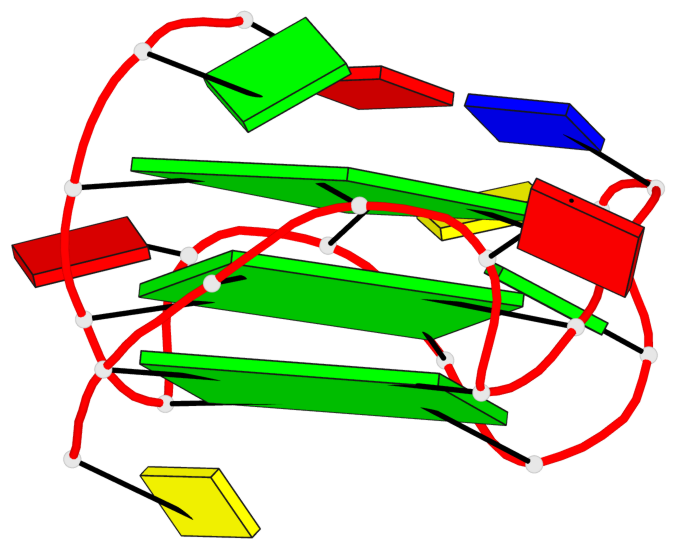
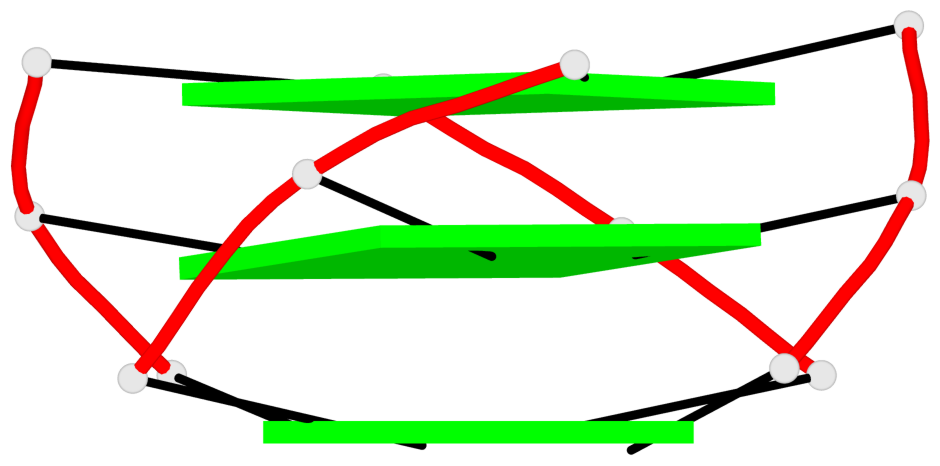
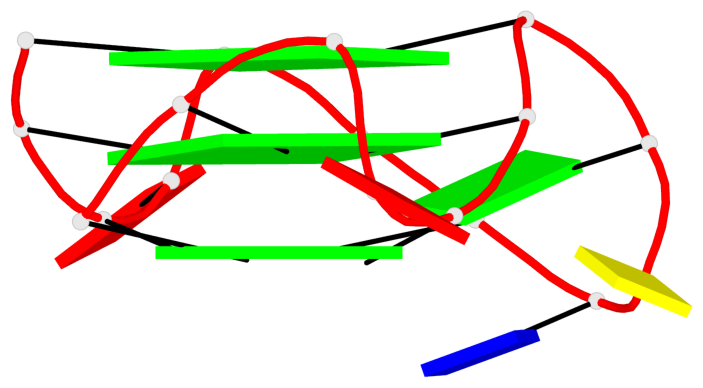
Base-block schematics in six views
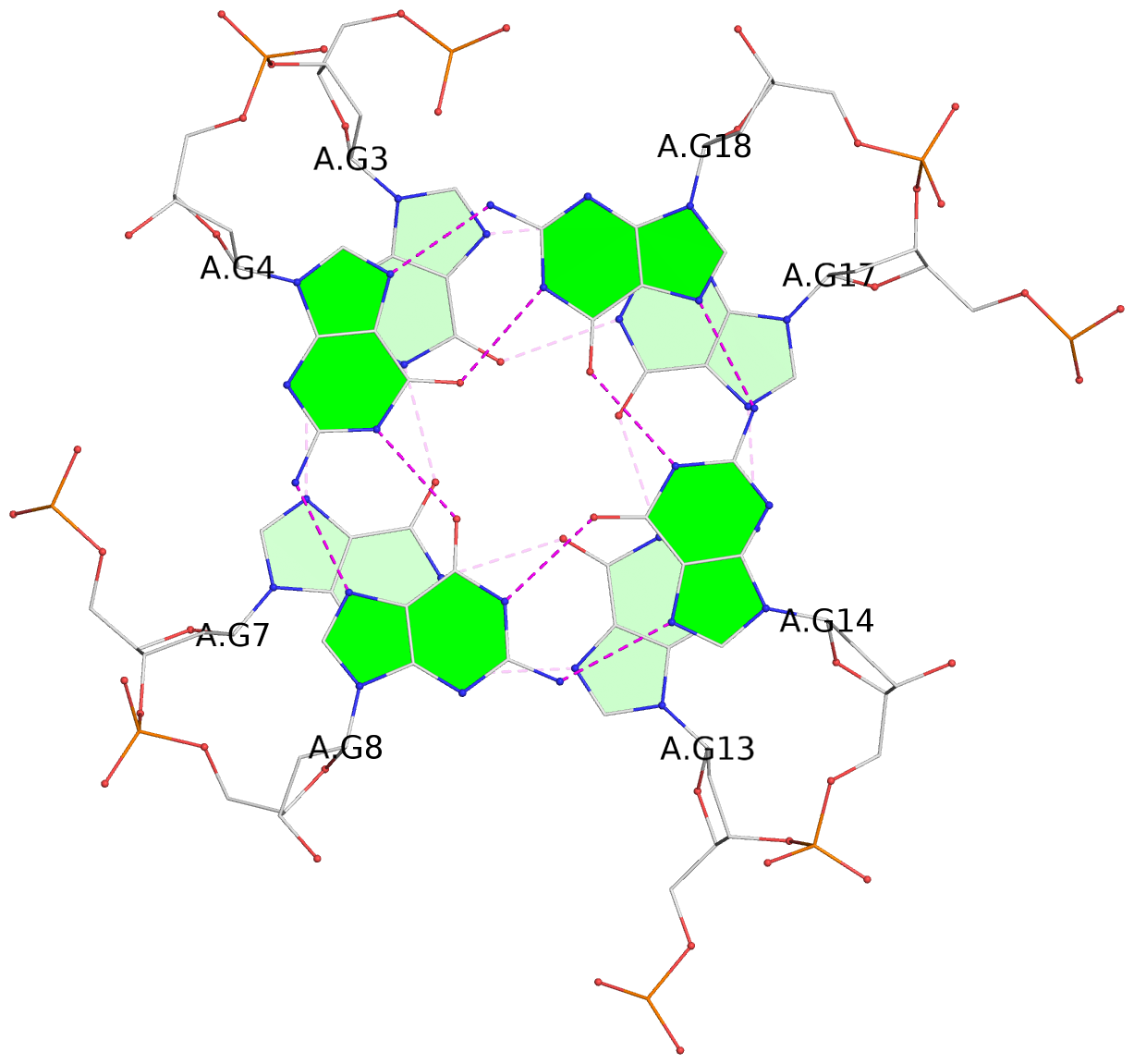
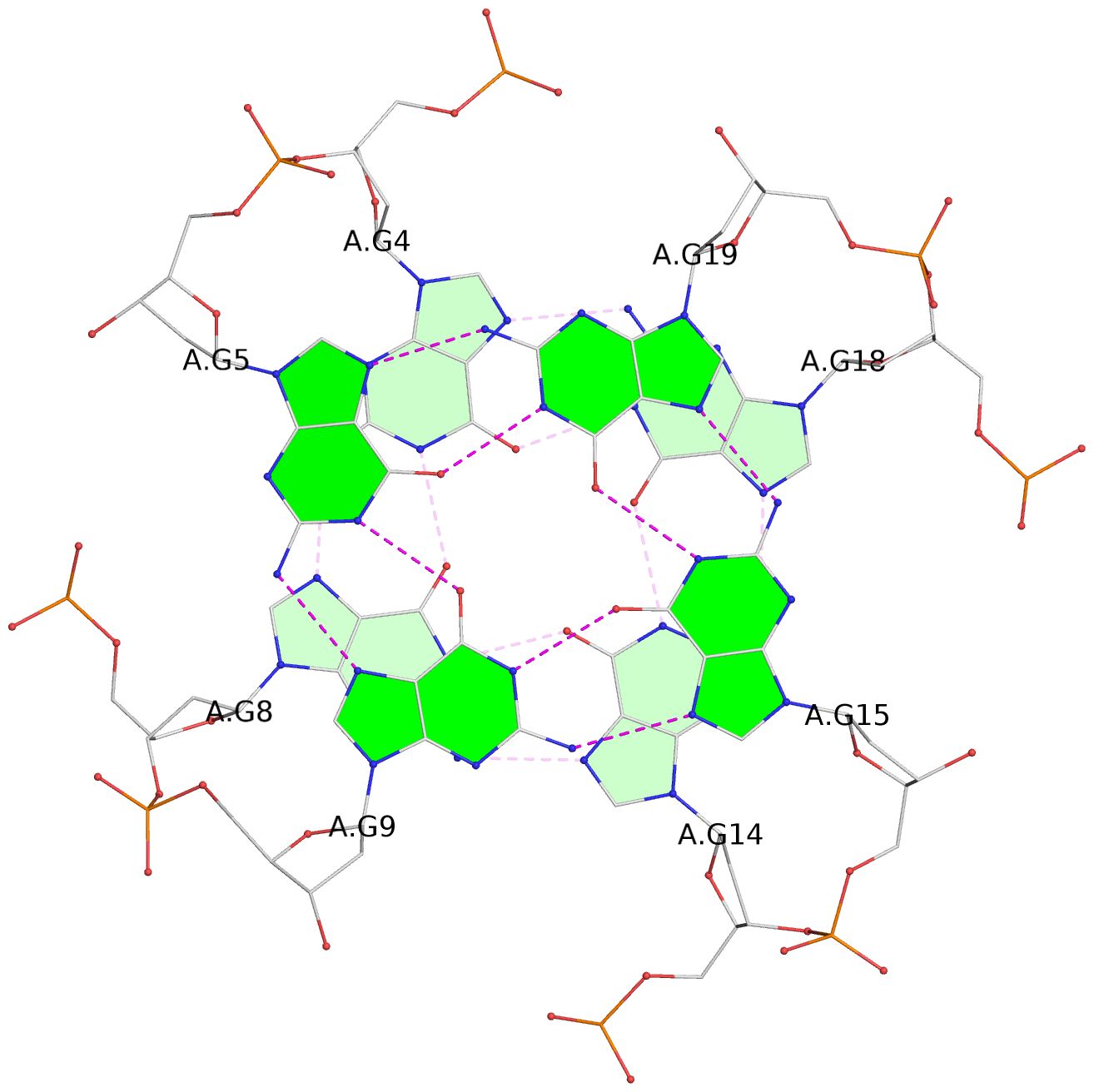
List of 3 G-tetrads
1 glyco-bond=---- sugar=---- groove=---- planarity=0.218 type=bowl nts=4 GGGG A.DG3,A.DG7,A.DG13,A.DG17 2 glyco-bond=---- sugar=--.- groove=---- planarity=0.193 type=other nts=4 GGGG A.DG4,A.DG8,A.DG14,A.DG18 3 glyco-bond=---- sugar=---- groove=---- planarity=0.137 type=planar nts=4 GGGG A.DG5,A.DG9,A.DG15,A.DG19
List of 1 G4-helix
In DSSR, a G4-helix is defined by stacking interactions of G-tetrads, regardless of backbone connectivity, and may contain more than one G4-stem.
Helix#1, 3 G-tetrad layers, INTRA-molecular, with 1 stem
List of 1 G4-stem
In DSSR, a G4-stem is defined as a G4-helix with backbone connectivity. Bulges are also allowed along each of the four strands.