Detailed DSSR results for the G-quadruplex: PDB entry 2l7v
Created and maintained by Xiang-Jun Lu <xiangjun@x3dna.org>
Citation: Please cite the NAR'20 DSSR-PyMOL schematics paper and/or the NAR'15 DSSR method paper.
Summary information
- PDB id
- 2l7v
- Class
- DNA-inhibitor
- Method
- NMR
- Summary
- Quindoline-g-quadruplex complex
- Reference
- Dai J, Carver M, Hurley LH, Yang D (2011): "Solution Structure of a 2:1 Quindoline-c-MYC G-Quadruplex: Insights into G-Quadruplex-Interactive Small Molecule Drug Design." J.Am.Chem.Soc., 133, 17673-17680. doi: 10.1021/ja205646q.
- Abstract
- Unimolecular parallel-stranded G-quadruplex structures are found to be prevalent in gene promoters. The nuclease hypersensitivity element III(1) (NHE III(1)) of the c-MYC promoter can form transcriptionally active and silenced forms, and the formation of DNA G-quadruplex structures has been shown to be critical for c-MYC transcriptional silencing. The solution structure of a 2:1 quindoline-G-quadruplex complex has been solved and shows unexpected features, including the drug-induced reorientation of the flanking sequences to form a new binding pocket. While both 3' and 5' complexes show overall similar features, there are identifiable differences that emphasize the importance of both stacking and electronic interactions. For the first time, we describe the importance of the shape of the ligand as well as the two flanking bases in determining drug binding specificity. These structures provide important insights for the structure-based rational design of drugs that bind to unimolecular parallel G-quadruplexes commonly found in promoter elements.
- G4 notes
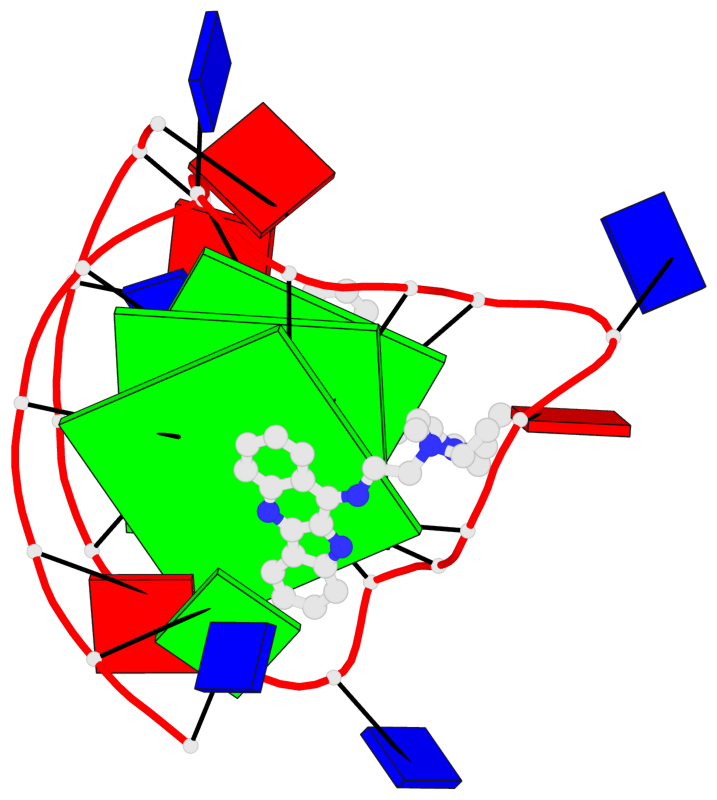
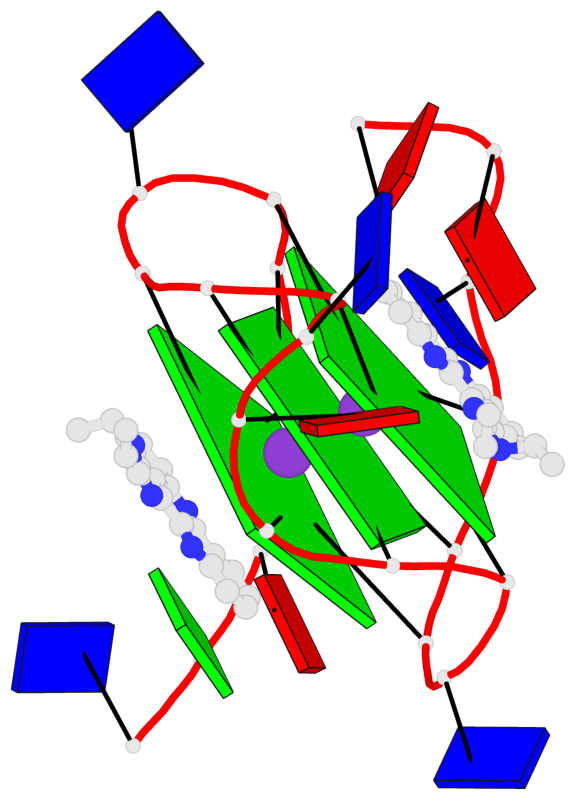
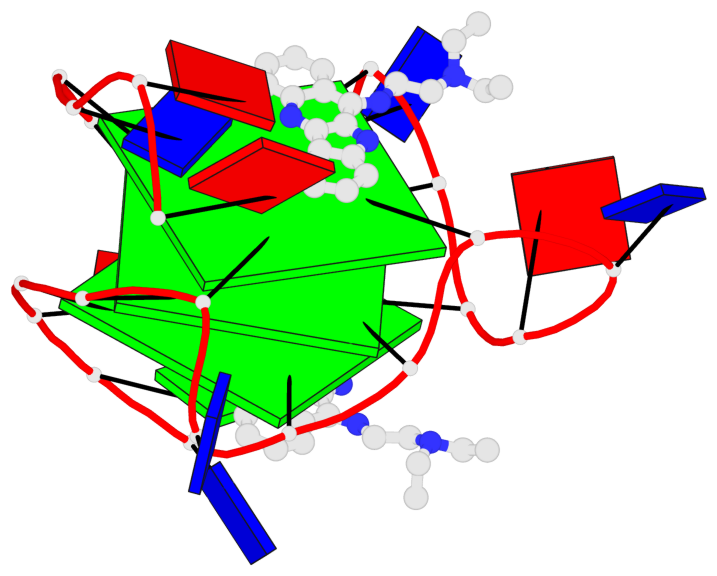
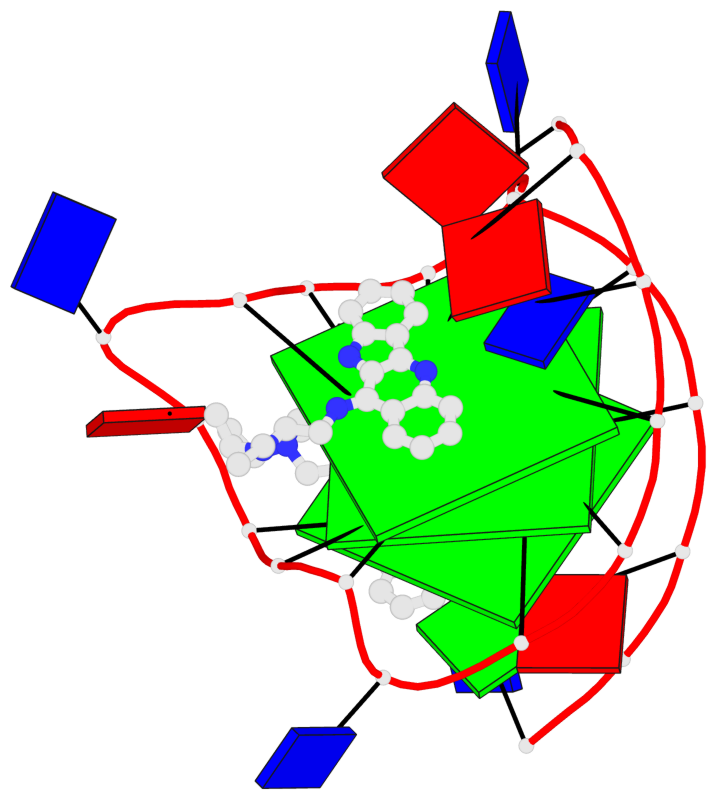
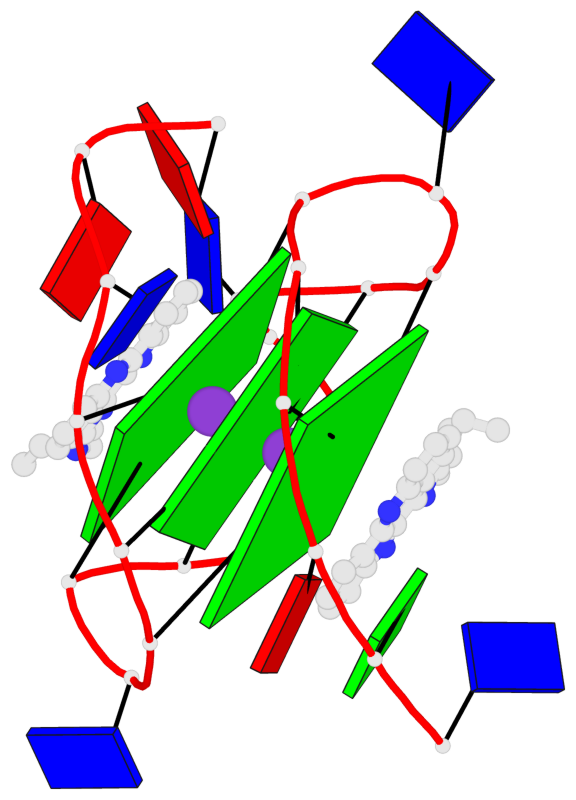
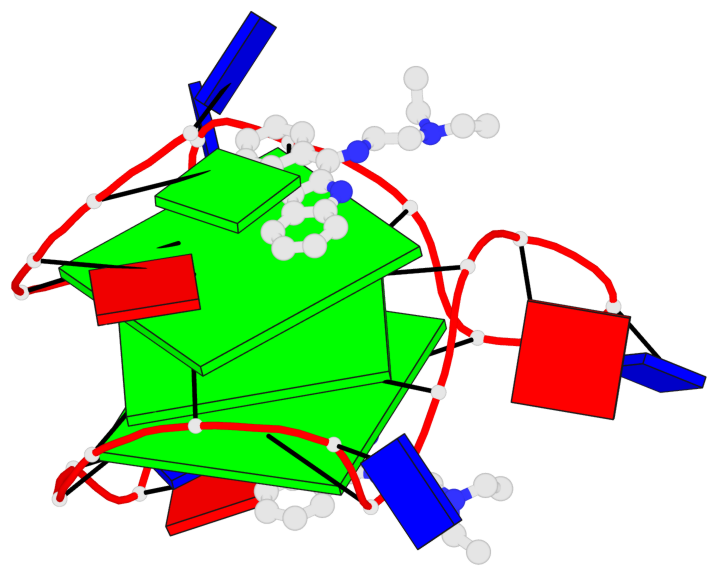
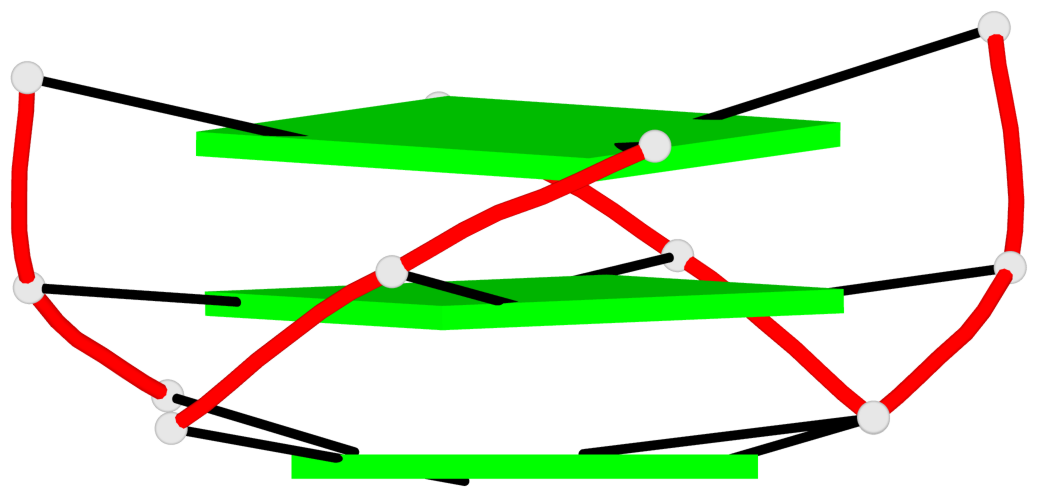
- 3 G-tetrads, 1 G4 helix, 1 G4 stem, 3(-P-P-P), parallel(4+0), UUUU
Base-block schematics in six views
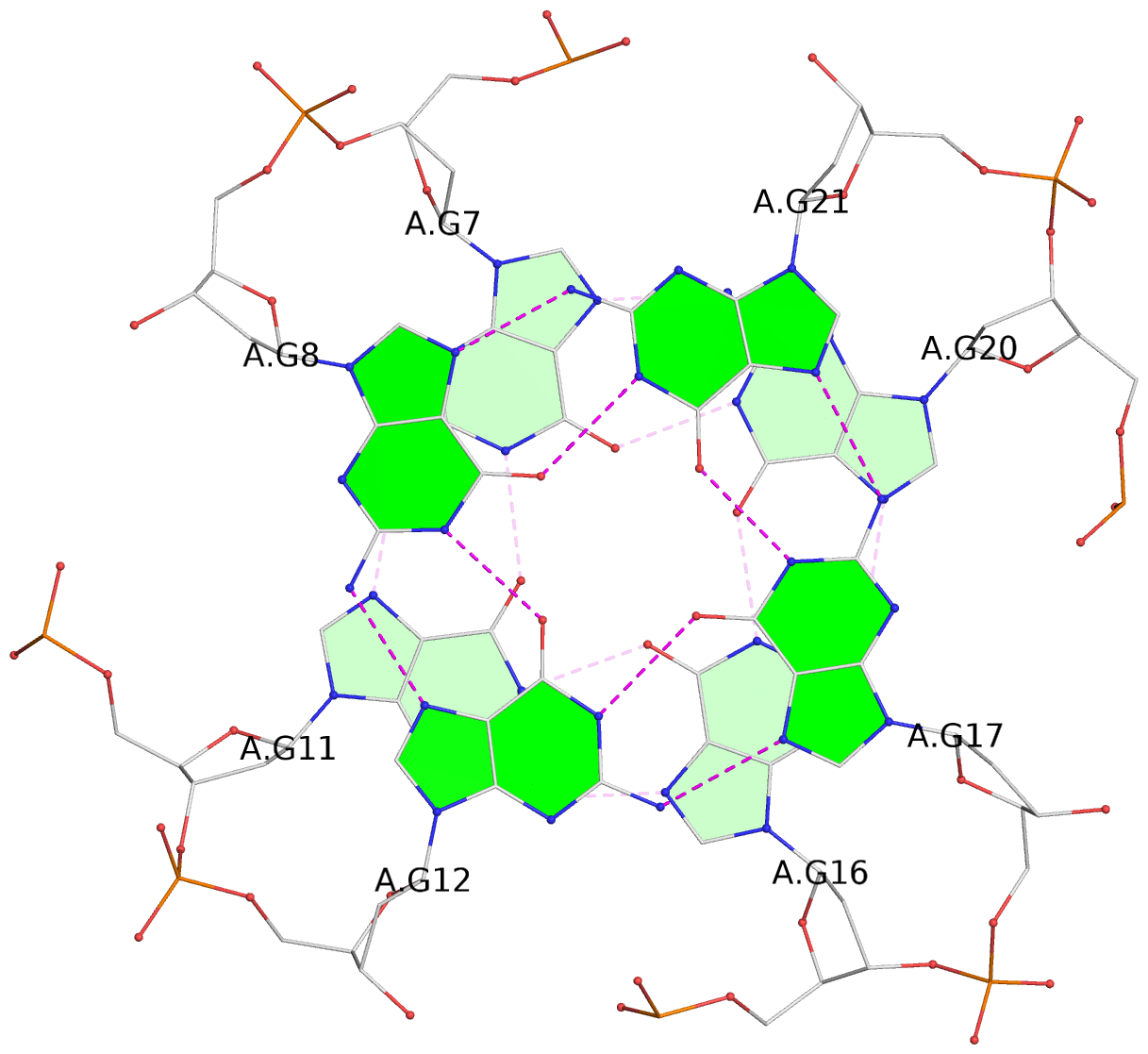
List of 3 G-tetrads
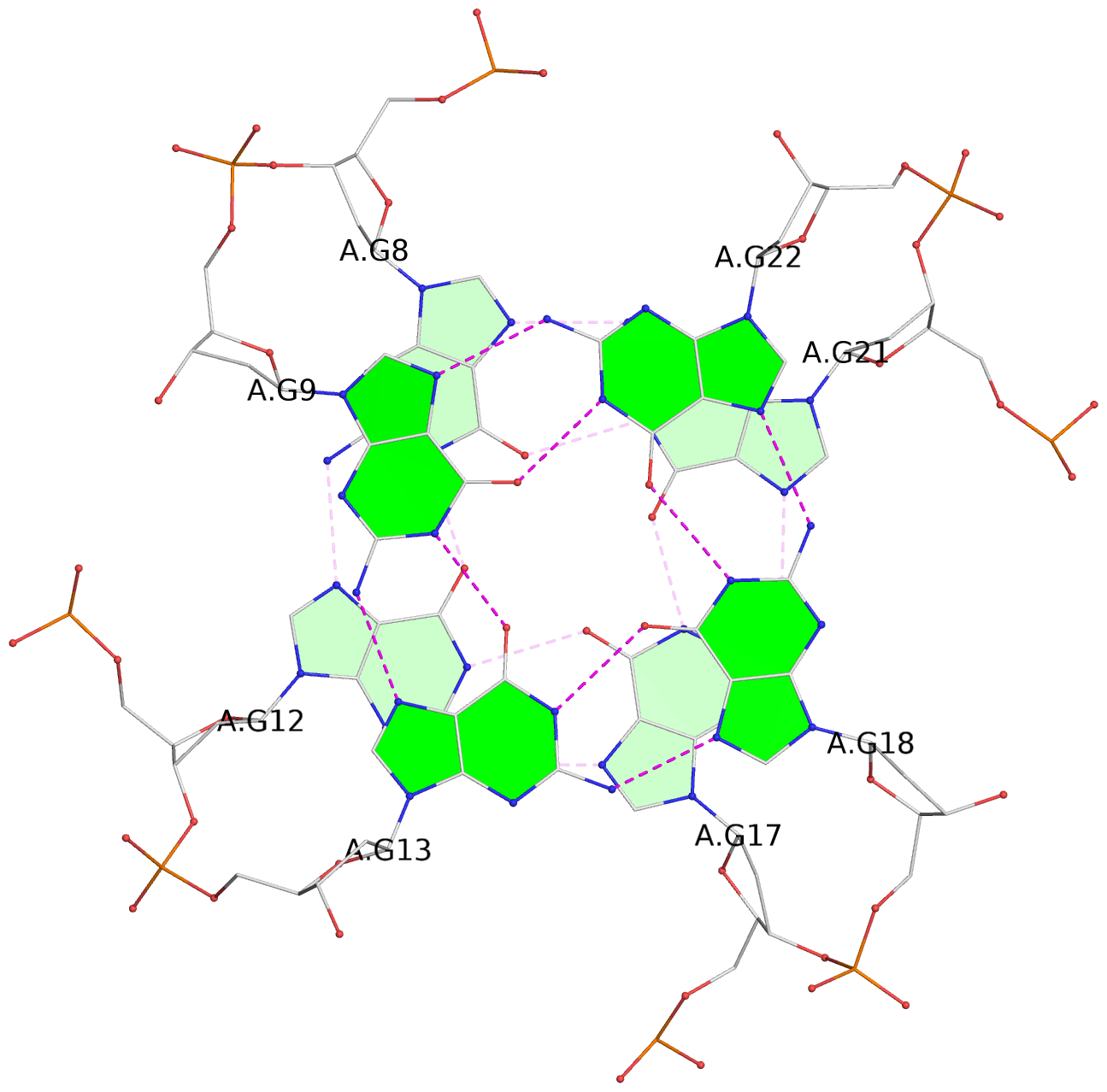
1 glyco-bond=---- sugar=-.-- groove=---- planarity=0.146 type=planar nts=4 GGGG A.DG7,A.DG11,A.DG16,A.DG20 2 glyco-bond=---- sugar=--.- groove=---- planarity=0.163 type=other nts=4 GGGG A.DG8,A.DG12,A.DG17,A.DG21 3 glyco-bond=---- sugar=---- groove=---- planarity=0.178 type=other nts=4 GGGG A.DG9,A.DG13,A.DG18,A.DG22
List of 1 G4-helix
In DSSR, a G4-helix is defined by stacking interactions of G-tetrads, regardless of backbone connectivity, and may contain more than one G4-stem.
Helix#1, 3 G-tetrad layers, INTRA-molecular, with 1 stem
List of 1 G4-stem
In DSSR, a G4-stem is defined as a G4-helix with backbone connectivity. Bulges are also allowed along each of the four strands.