Detailed DSSR results for the G-quadruplex: PDB entry 2lyg
Created and maintained by Xiang-Jun Lu <xiangjun@x3dna.org>
Citation: Please cite the NAR'20 DSSR-PyMOL schematics paper and/or the NAR'15 DSSR method paper.
Summary information
- PDB id
- 2lyg
- Class
- DNA
- Method
- NMR
- Summary
- Fuc_tba
- Reference
- Gomez-Pinto I, Vengut-Climent E, Lucas R, Avino A, Eritja R, Gonzalez C, Morales JC (2013): "Carbohydrate-DNA interactions at G-quadruplexes: folding and stability changes by attaching sugars at the 5'-end." Chemistry, 19, 1920-1927. doi: 10.1002/chem.201203902.
- Abstract
- Quadruplex DNA structures are attracting an enormous interest in many areas of chemistry, ranging from chemical biology, supramolecular chemistry to nanoscience. We have prepared carbohydrate-DNA conjugates containing the oligonucleotide sequences of G-quadruplexes (thrombin binding aptamer (TBA) and human telomere (TEL)), measured their thermal stability and studied their structure in solution by using NMR and molecular dynamics. The solution structure of a fucose-TBA conjugate shows stacking interactions between the carbohydrate and the DNA G-tetrad in addition to hydrogen bonding and hydrophobic contacts. We have also shown that attaching carbohydrates at the 5'-end of a quadruplex telomeric sequence can alter its folding topology. These results suggest the possibility of modulating the folding of the G-quadruplex by linking carbohydrates and have clear implications in molecular recognition and the design of new G-quadruplex ligands.
- G4 notes
- 2 G-tetrads, 1 G4 helix, 1 G4 stem, 2(+Ln+Lw+Ln), chair(2+2), UDUD
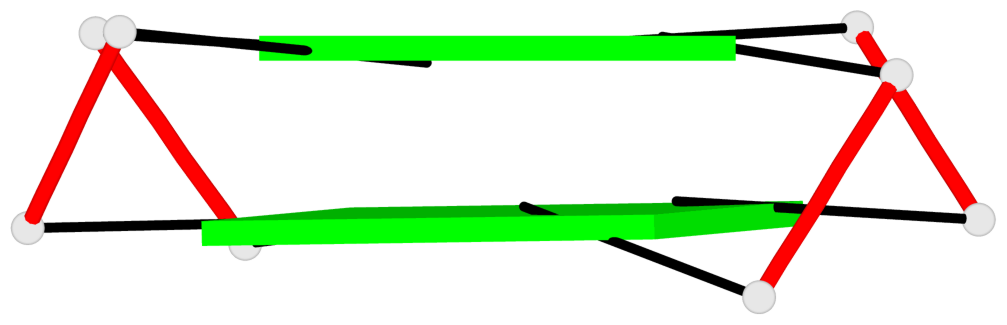
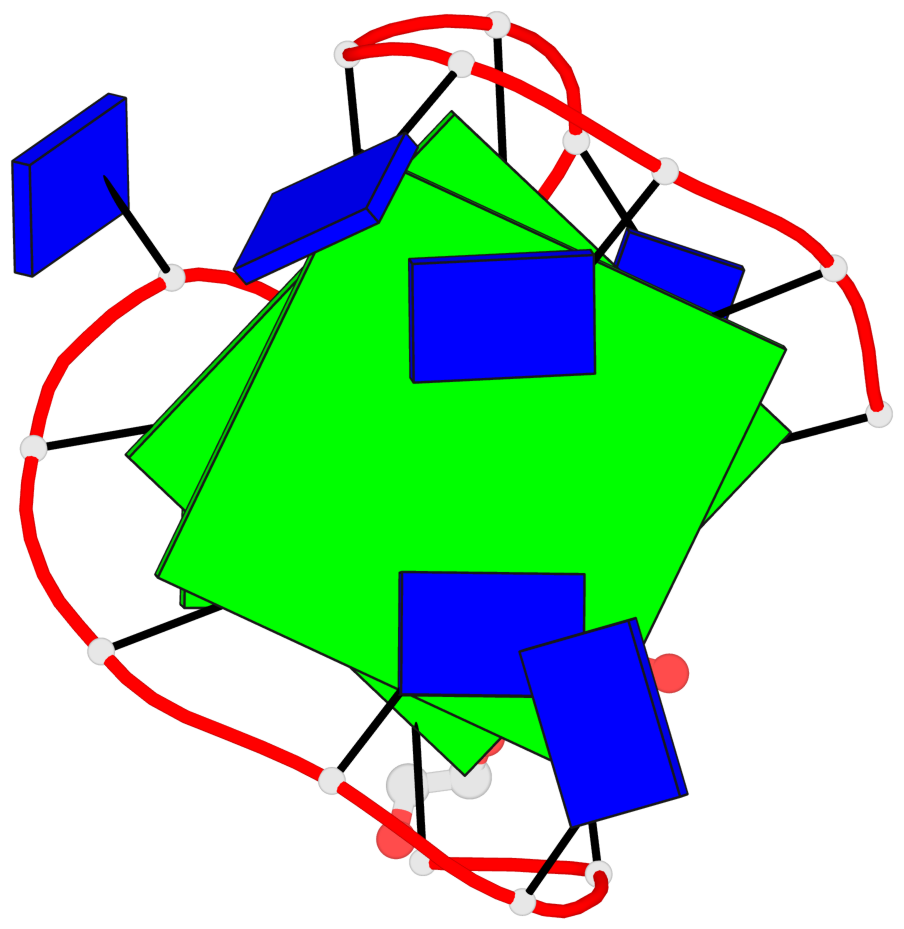
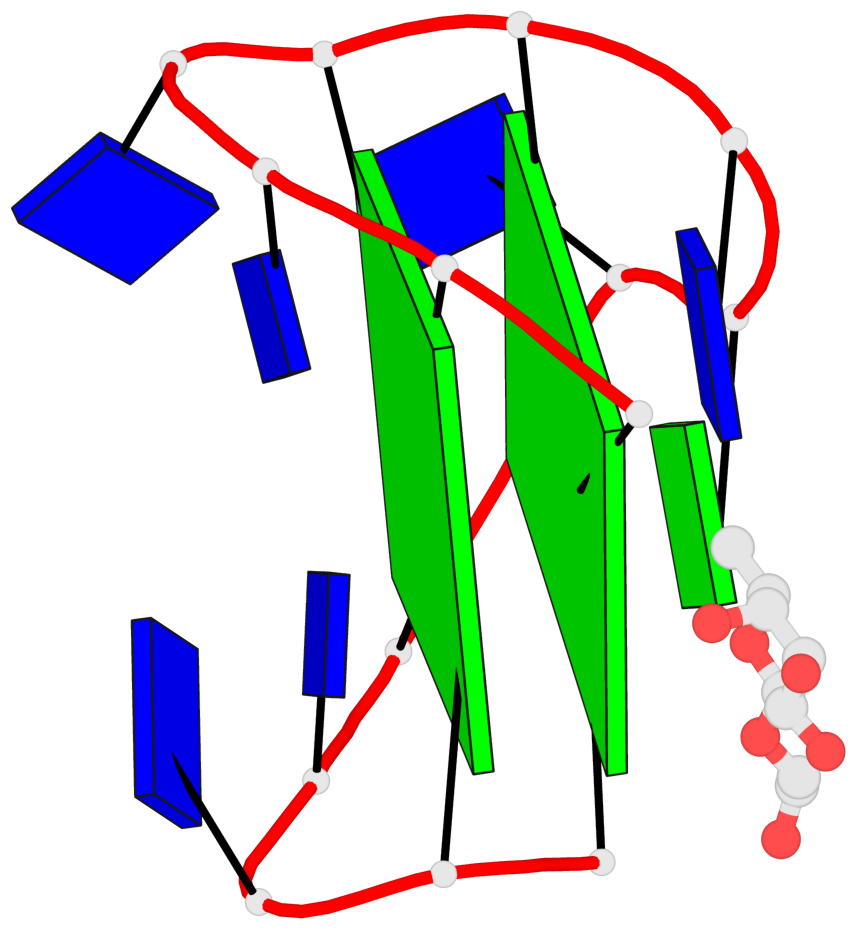
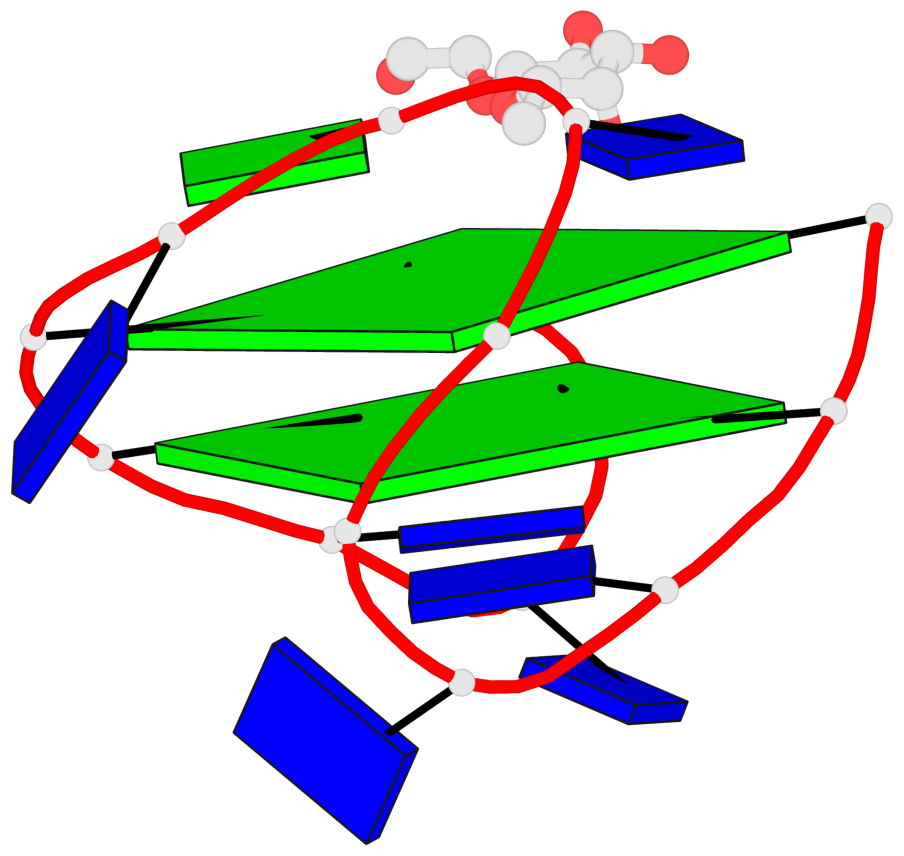
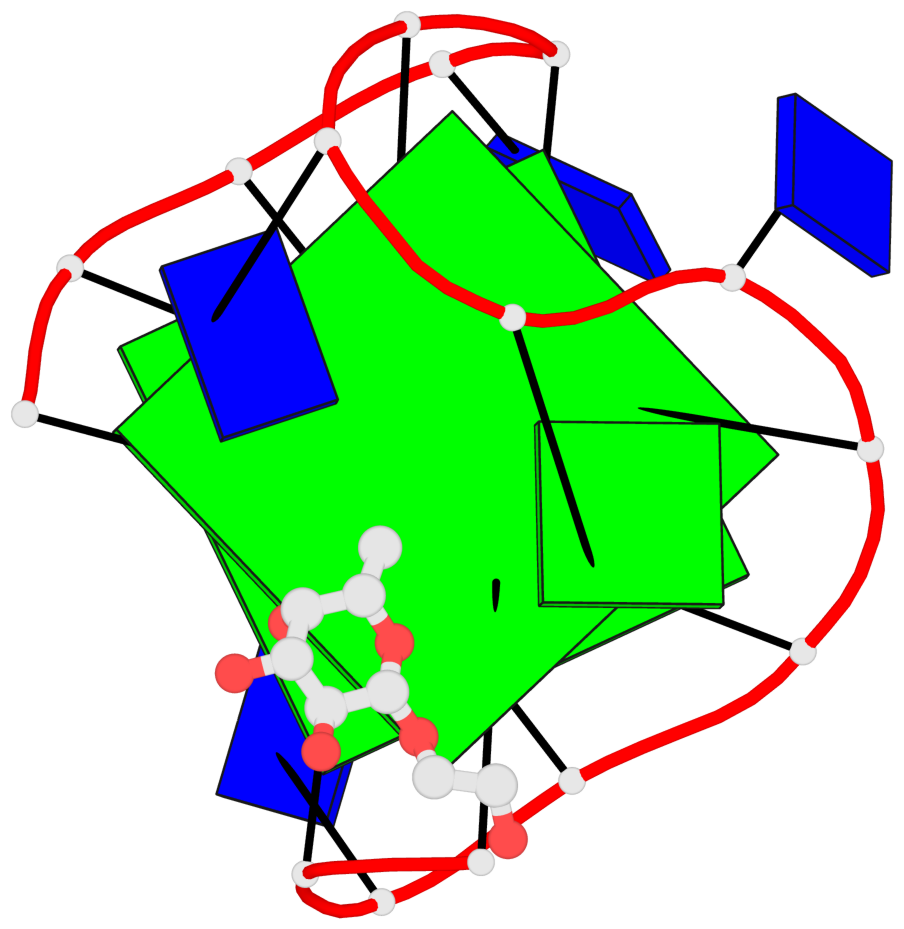
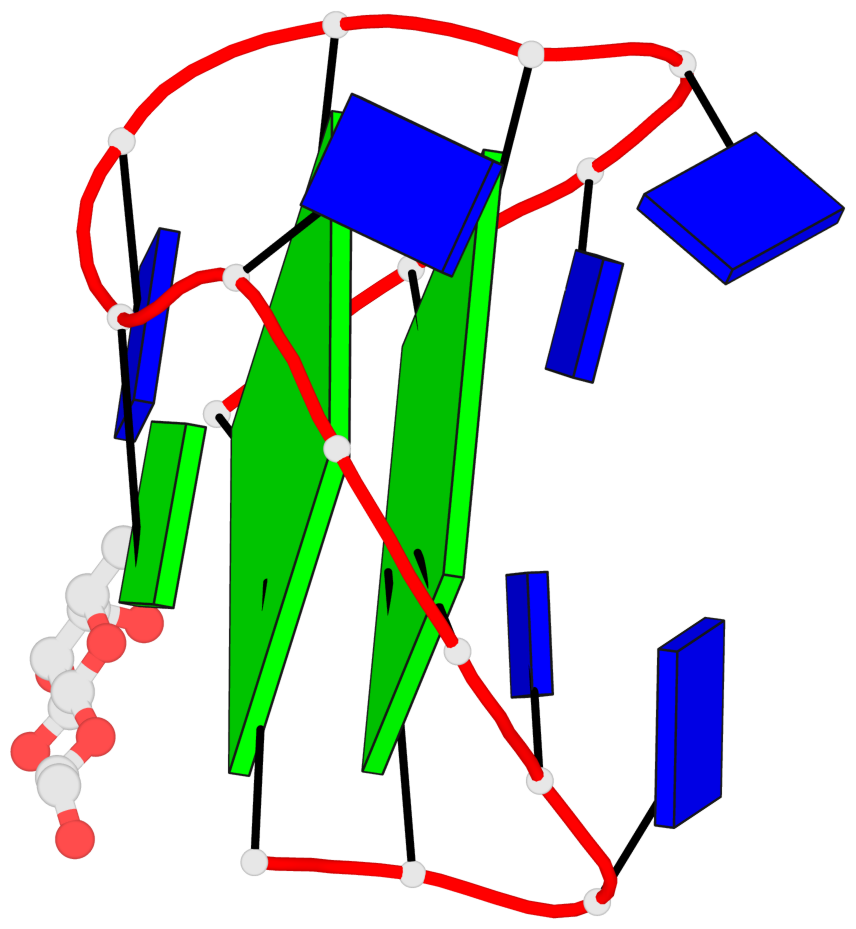
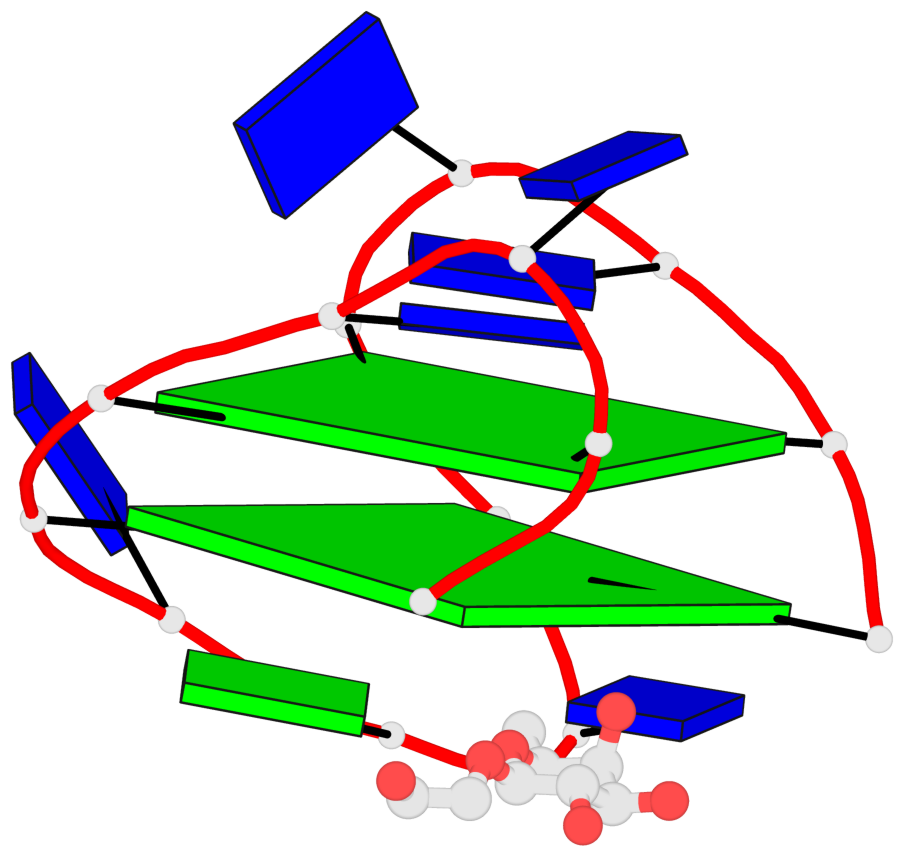
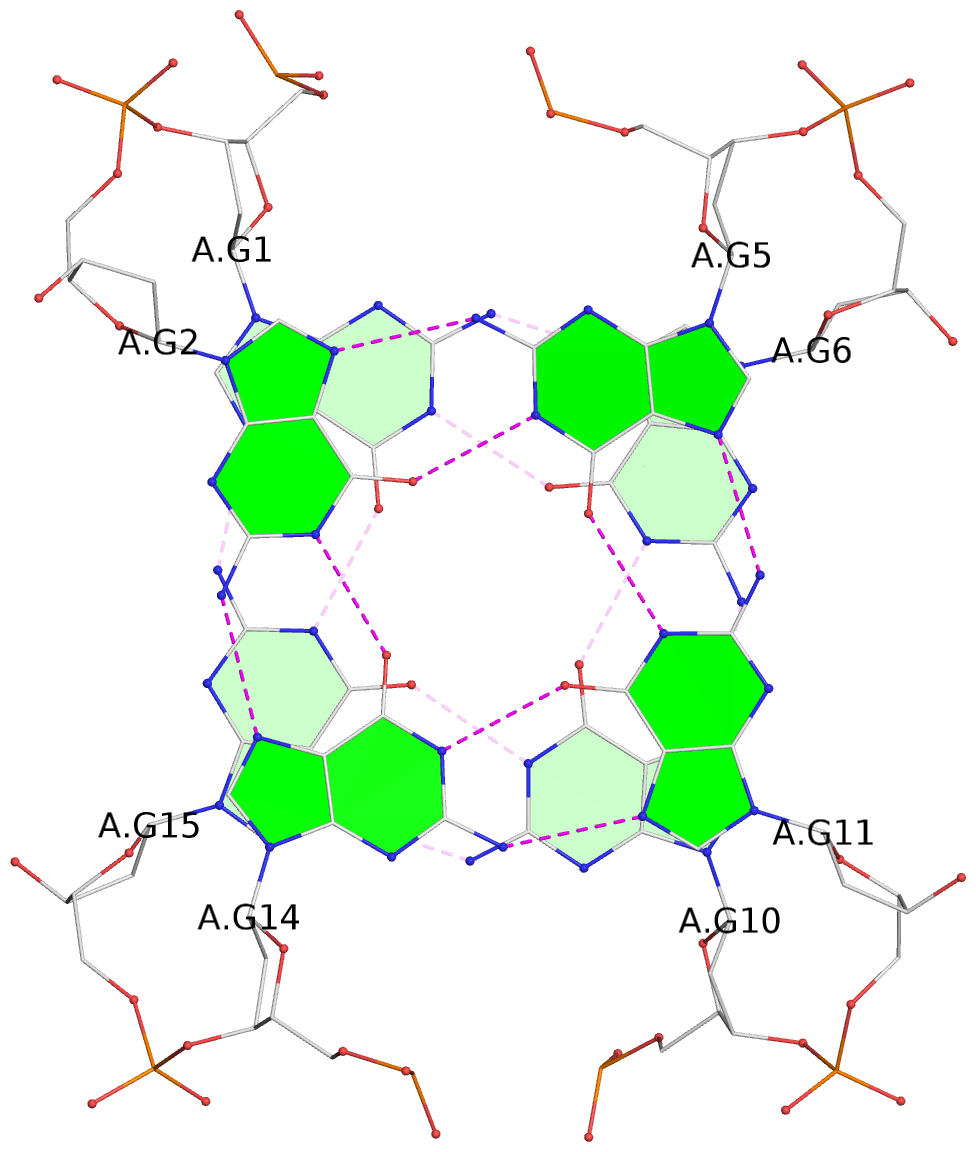
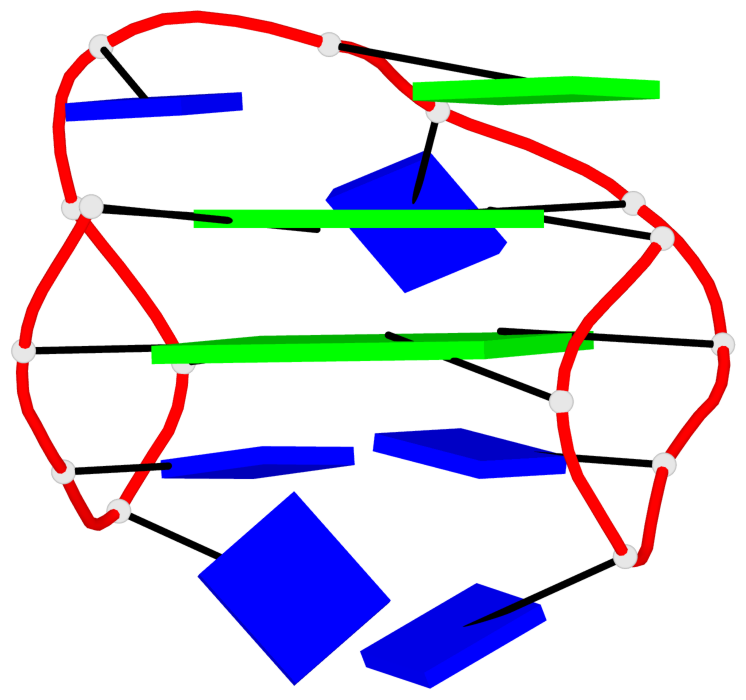
Base-block schematics in six views
List of 2 G-tetrads
1 glyco-bond=s-s- sugar=---- groove=wnwn planarity=0.177 type=other nts=4 GGGG A.DG1,A.DG15,A.DG10,A.DG6 2 glyco-bond=-s-s sugar=---- groove=wnwn planarity=0.261 type=bowl nts=4 GGGG A.DG2,A.DG14,A.DG11,A.DG5
List of 1 G4-helix
In DSSR, a G4-helix is defined by stacking interactions of G-tetrads, regardless of backbone connectivity, and may contain more than one G4-stem.
Helix#1, 2 G-tetrad layers, INTRA-molecular, with 1 stem
List of 1 G4-stem
In DSSR, a G4-stem is defined as a G4-helix with backbone connectivity. Bulges are also allowed along each of the four strands.