Detailed DSSR results for the G-quadruplex: PDB entry 2m6w
Created and maintained by Xiang-Jun Lu <xiangjun@x3dna.org>
Citation: Please cite the NAR'20 DSSR-PyMOL schematics paper and/or the NAR'15 DSSR method paper.
Summary information
- PDB id
- 2m6w
- Class
- DNA
- Method
- NMR
- Summary
- Solution NMR structure of the d(ggggttggggttttggggaagggg) quadruplex in sodium conditions
- Reference
- Dvorkin SA, Karsisiotis AI, Webba da Silva M (2018): "Encoding canonical DNA quadruplex structure." Sci Adv, 4, eaat3007. doi: 10.1126/sciadv.aat3007.
- Abstract
- The main challenge in DNA quadruplex design is to encode a three-dimensional structure into the primary sequence, despite its multiple, repetitive guanine segments. We identify and detail structural elements describing all 14 feasible canonical quadruplex scaffolds and demonstrate their use in control of design. This work outlines a new roadmap for implementation of targeted design of quadruplexes for material, biotechnological, and therapeutic applications.
- G4 notes
- 4 G-tetrads, 1 G4 helix, 1 G4 stem, 4(-LwD+Ln), basket(2+2), UDDU
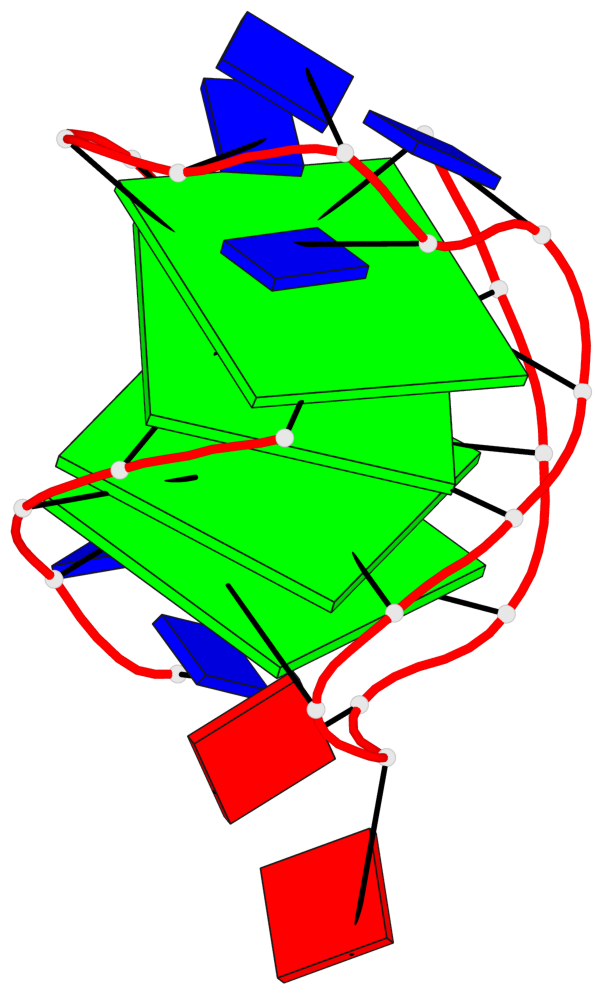
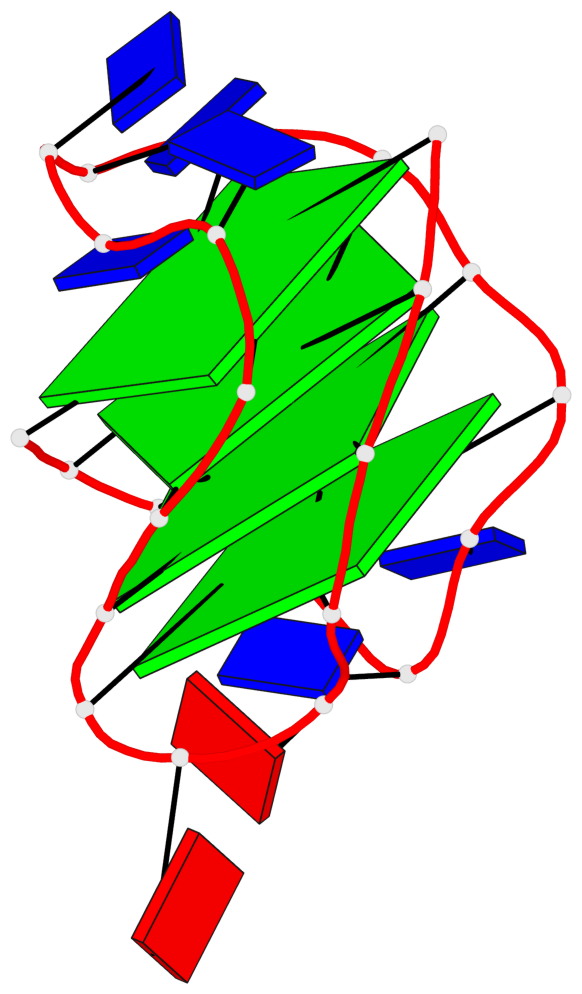
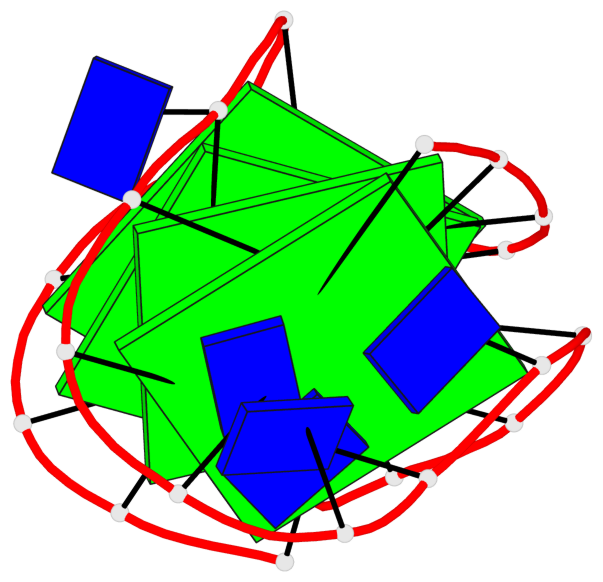
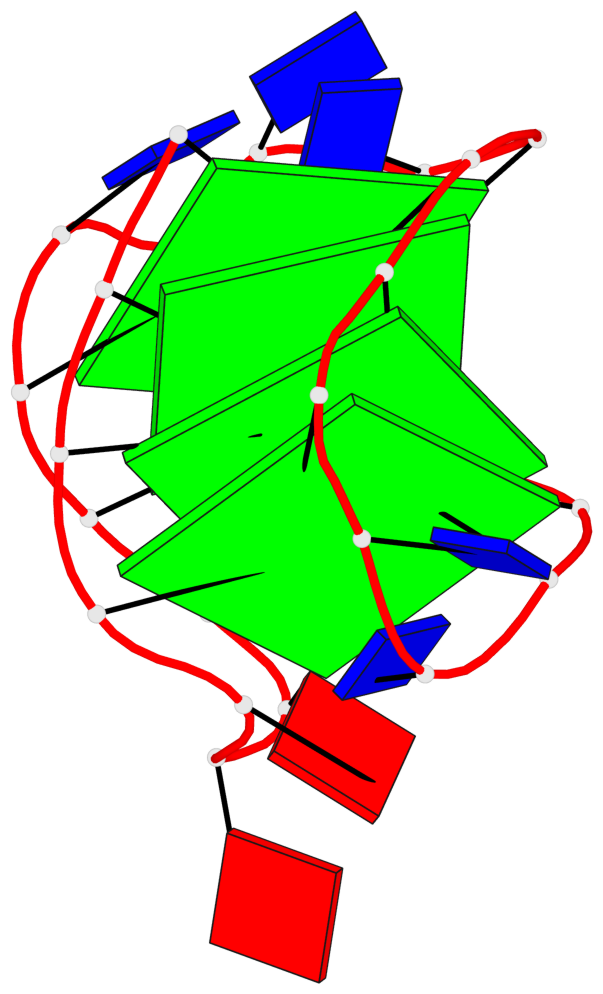
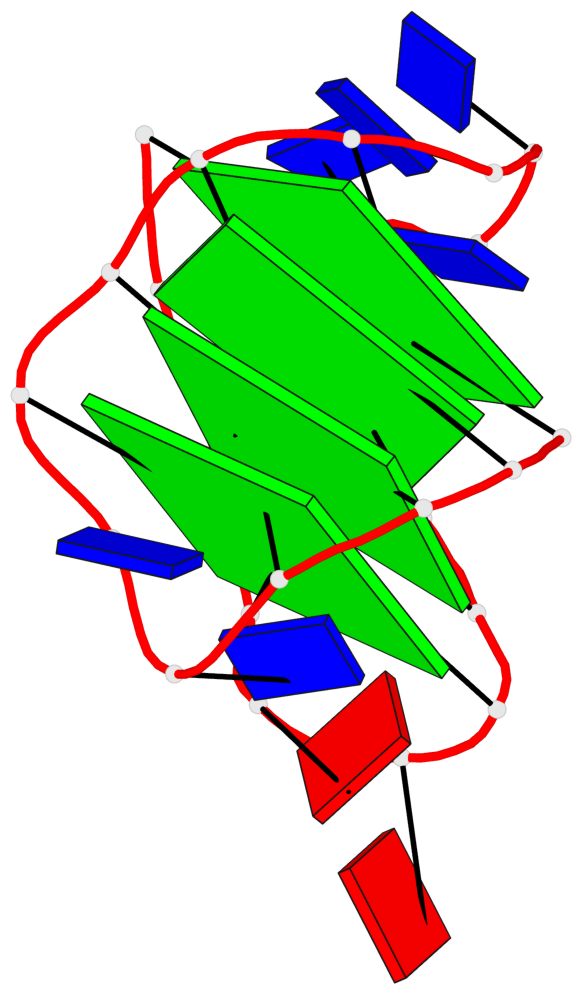
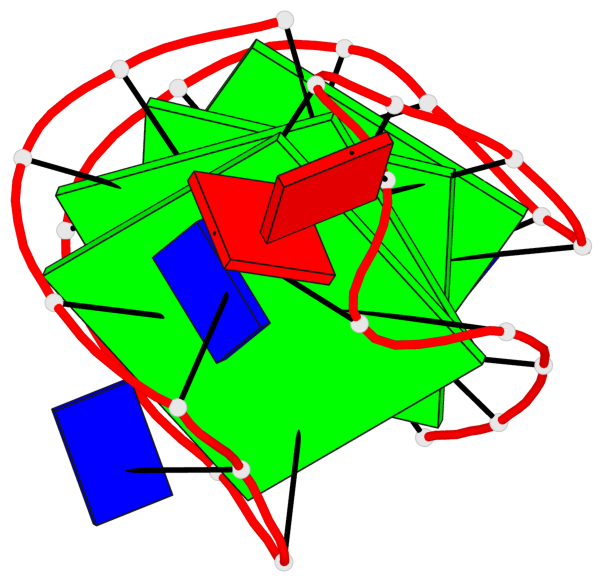
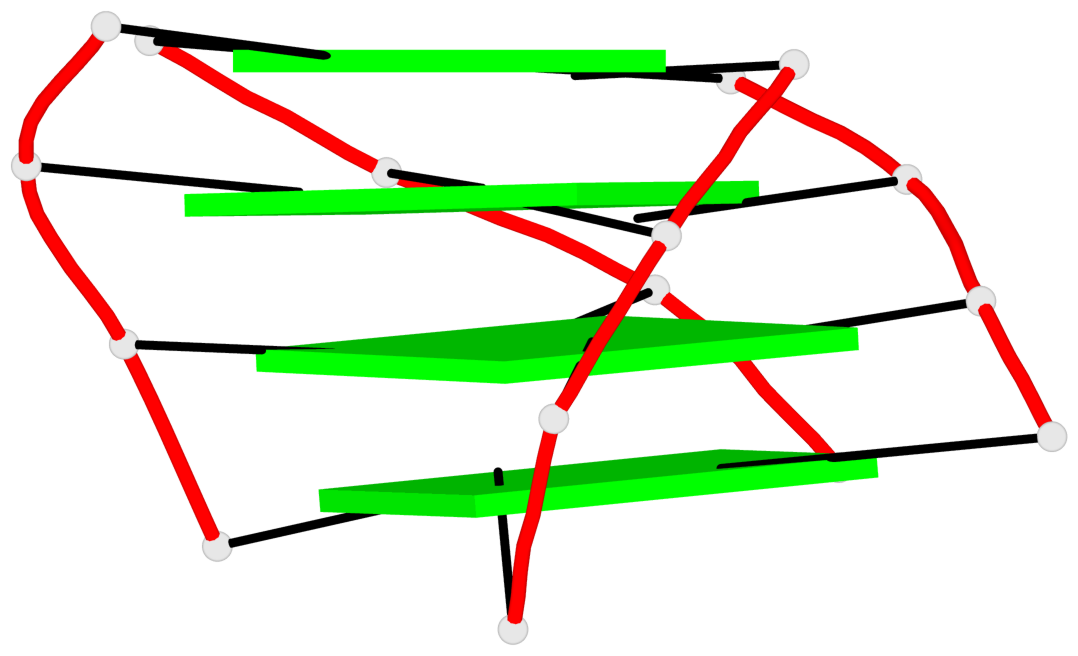
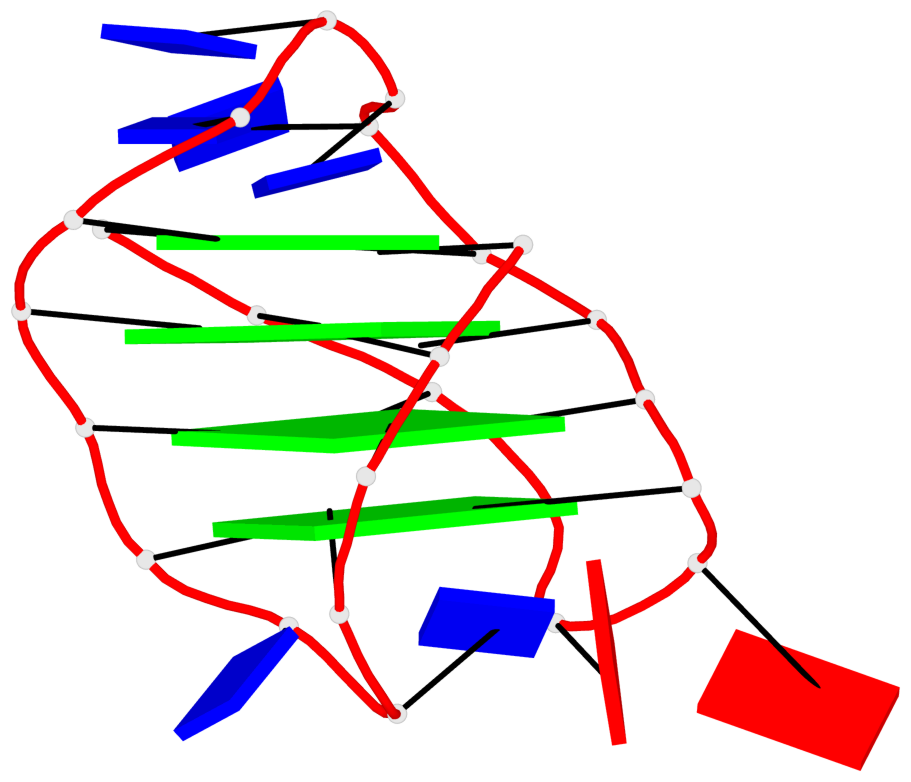
Base-block schematics in six views
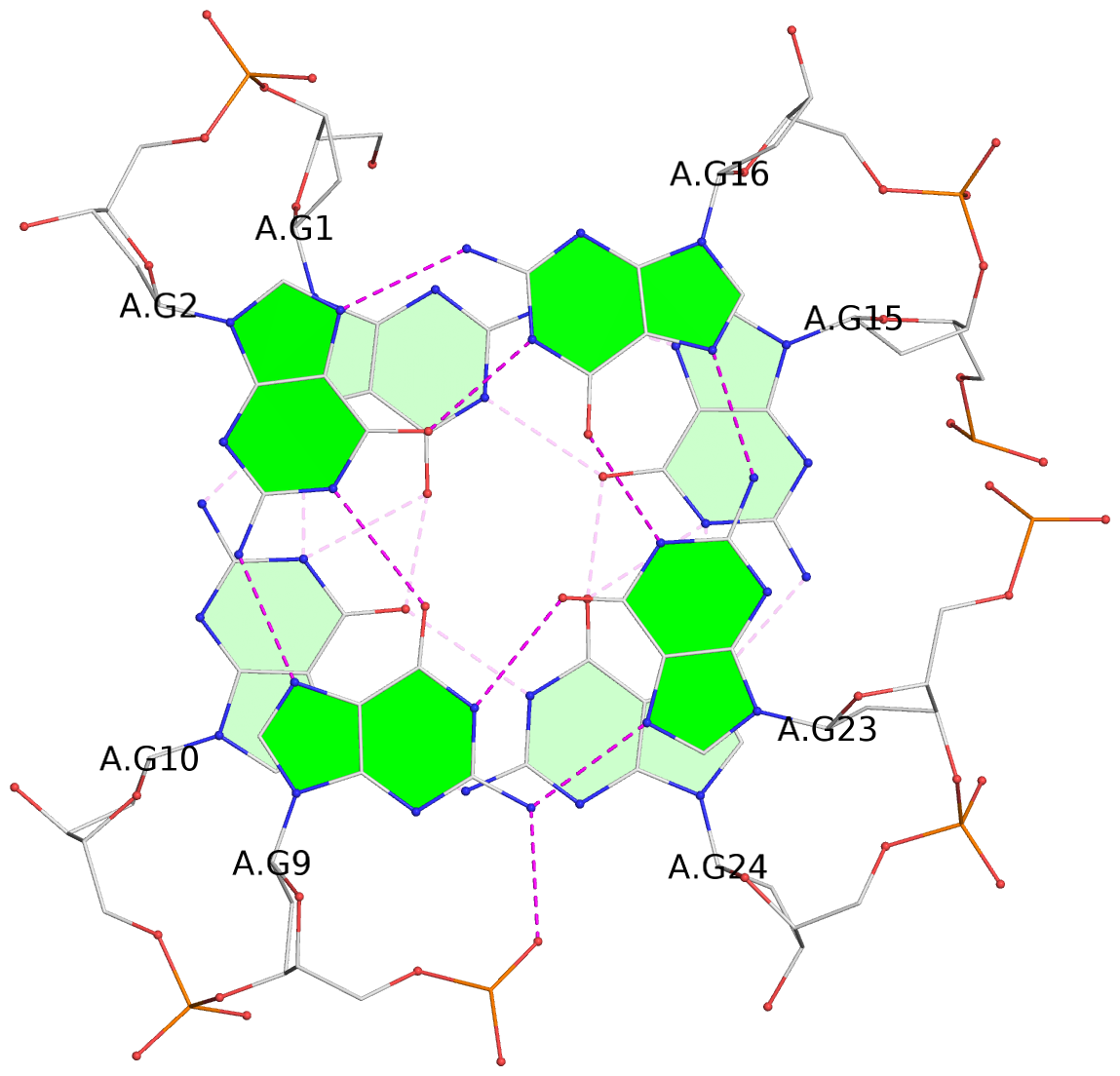
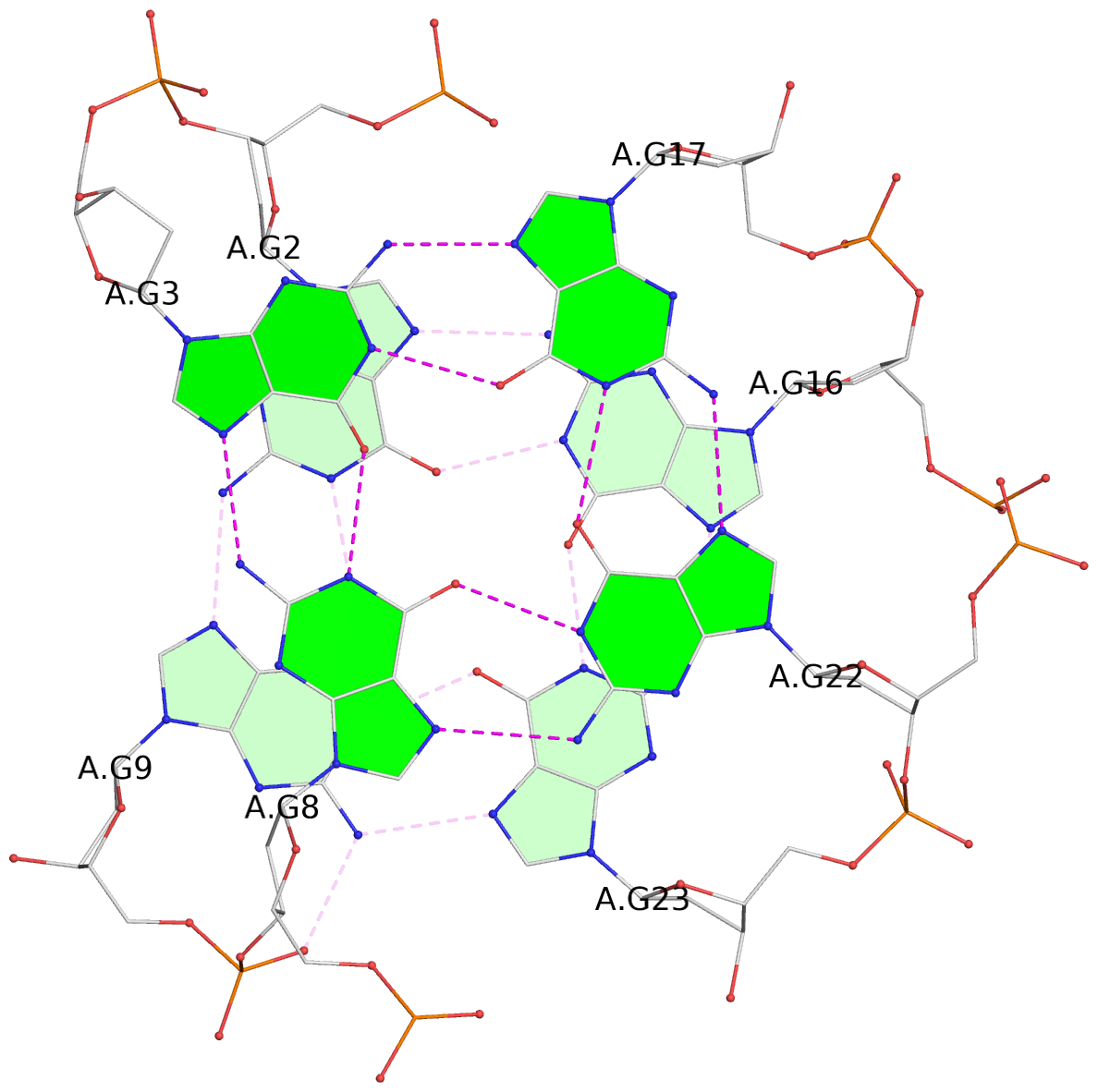
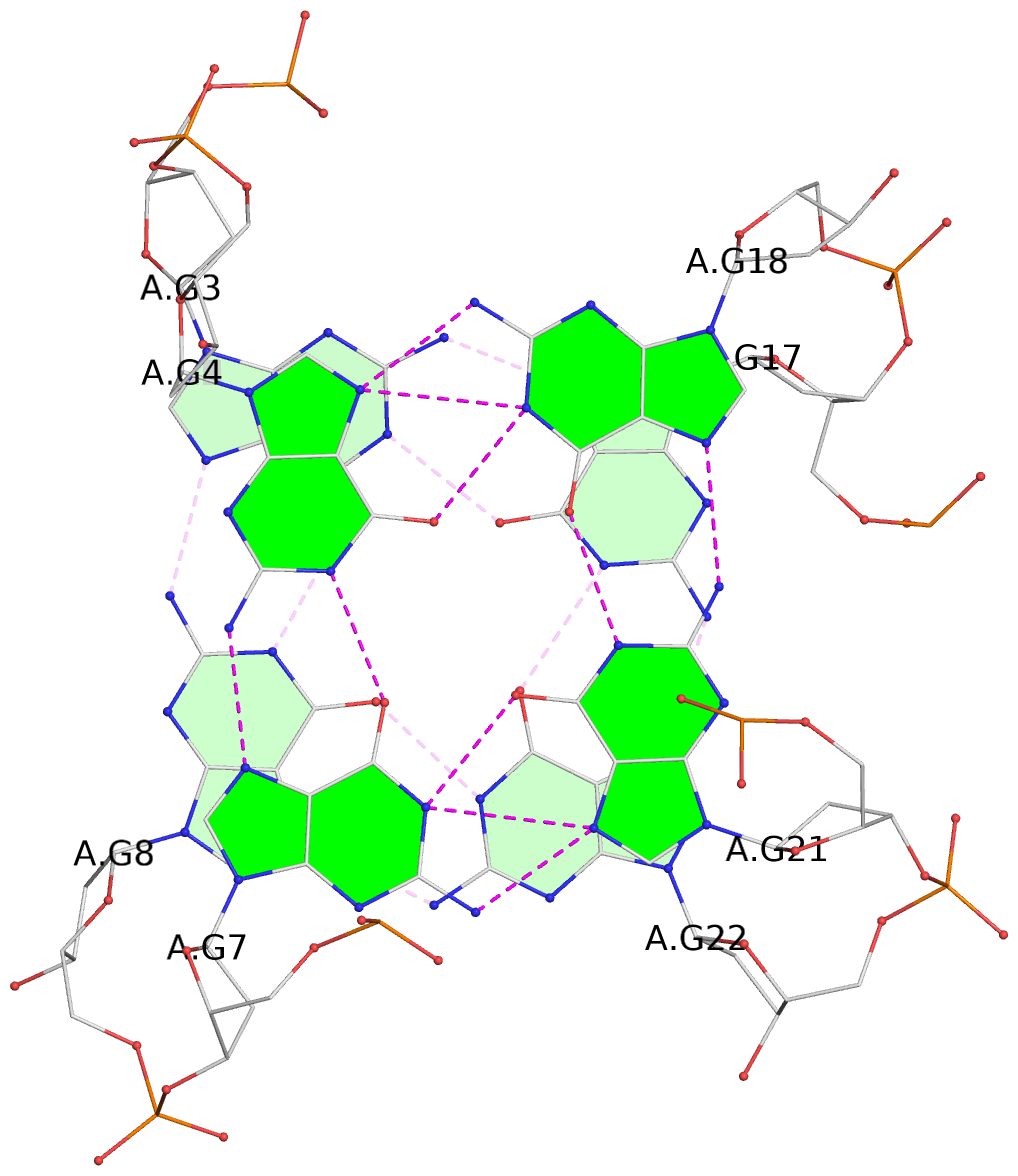
List of 4 G-tetrads
1 glyco-bond=s--s sugar=---- groove=w-n- planarity=0.146 type=planar nts=4 GGGG A.DG1,A.DG10,A.DG24,A.DG15 2 glyco-bond=-ss- sugar=---- groove=w-n- planarity=0.209 type=other nts=4 GGGG A.DG2,A.DG9,A.DG23,A.DG16 3 glyco-bond=s--s sugar=-3-. groove=w-n- planarity=0.260 type=other nts=4 GGGG A.DG3,A.DG8,A.DG22,A.DG17 4 glyco-bond=-ss- sugar=3--. groove=w-n- planarity=0.222 type=other nts=4 GGGG A.DG4,A.DG7,A.DG21,A.DG18
List of 1 G4-helix
In DSSR, a G4-helix is defined by stacking interactions of G-tetrads, regardless of backbone connectivity, and may contain more than one G4-stem.
Helix#1, 4 G-tetrad layers, INTRA-molecular, with 1 stem
List of 1 G4-stem
In DSSR, a G4-stem is defined as a G4-helix with backbone connectivity. Bulges are also allowed along each of the four strands.