Detailed DSSR results for the G-quadruplex: PDB entry 2o4f
Created and maintained by Xiang-Jun Lu <xiangjun@x3dna.org>
Citation: Please cite the NAR'20 DSSR-PyMOL schematics paper and/or the NAR'15 DSSR method paper.
Summary information
- PDB id
- 2o4f
- Class
- DNA
- Method
- X-ray (1.5 Å)
- Summary
- Structure of a parallel-stranded guanine tetraplex crystallised with monovalent ions
- Reference
- Creze C, Rinaldi B, Haser R, Bouvet P, Gouet P (2007): "Structure of a d(TGGGGT) quadruplex crystallized in the presence of Li+ ions." Acta Crystallogr.,Sect.D, 63, 682-688. doi: 10.1107/S0907444907013315.
- Abstract
- A parallel 5'-d(TGGGGT)-3' quadruplex was formed in Na(+) solution and crystallized using lithium sulfate as the main precipitating agent. The X-ray structure was determined to 1.5 A resolution in space group P2(1) by molecular replacement. The asymmetric unit consists of a characteristic motif of two quadruplexes stacked at their 5' ends. All nucleotides are clearly defined in the density and could be positioned. A single bound Li(+) ion is observed at the surface of the column formed by the two joined molecules. Thus, this small alkali metal ion appears to be unsuitable as a replacement for the Na(+) ion in the central channel of G-quartets, unlike K(+) or Tl(+) ions. A well conserved constellation of water molecules is observed in the grooves of the dimeric structure.
- G4 notes
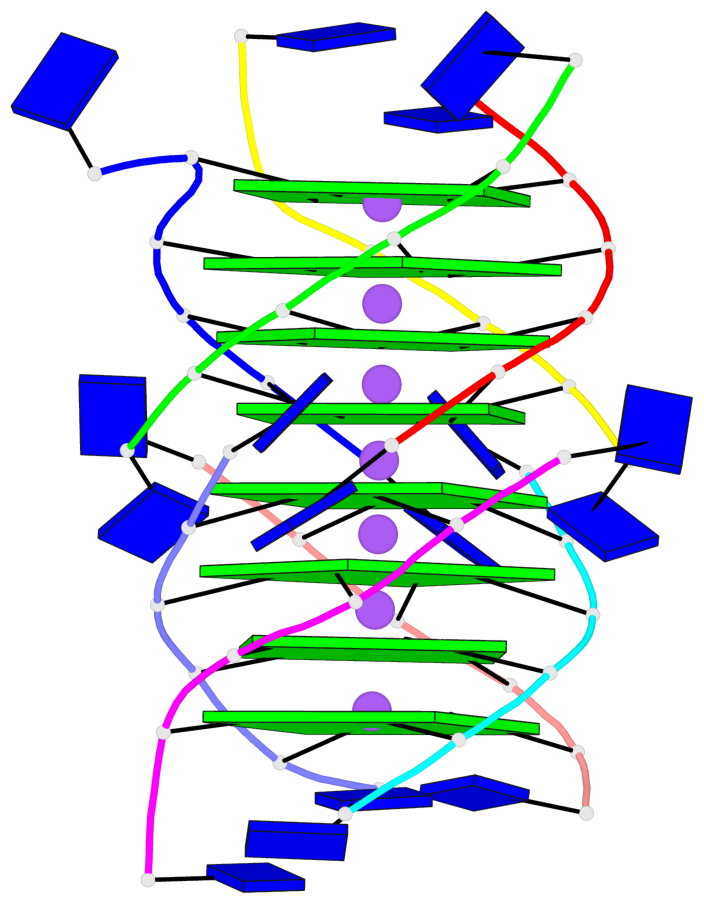
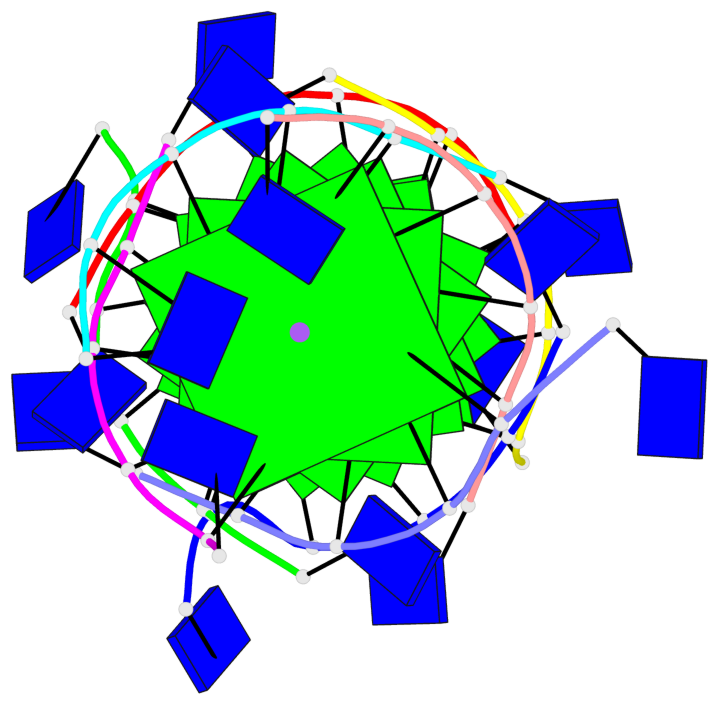
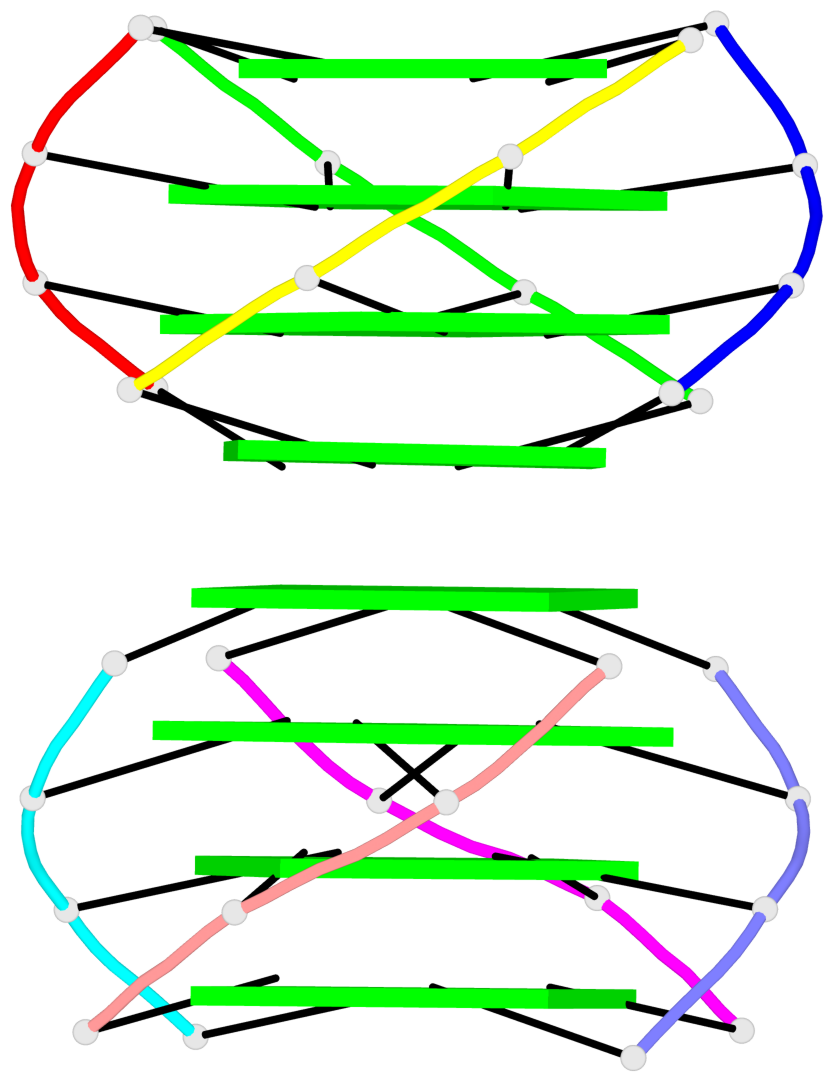
- 8 G-tetrads, 1 G4 helix, 2 G4 stems, 1 G4 coaxial stack, parallel(4+0), UUUU, coaxial interfaces: 5'/5'
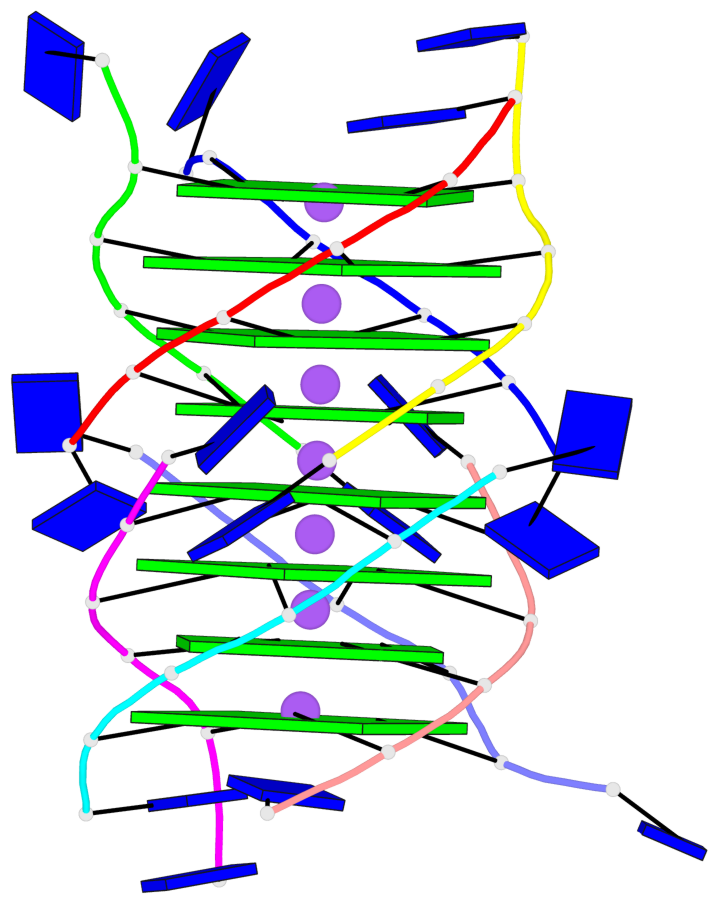
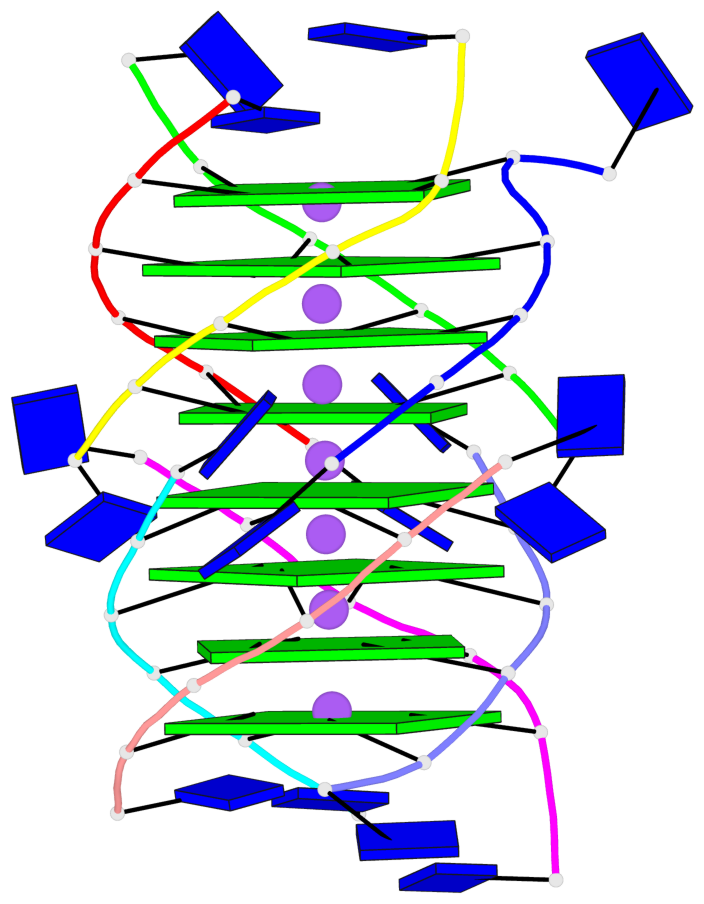
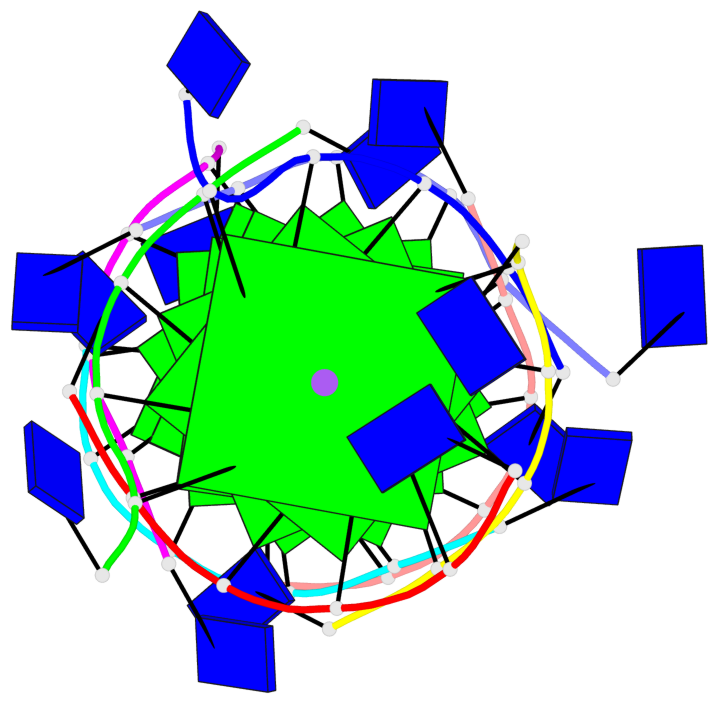
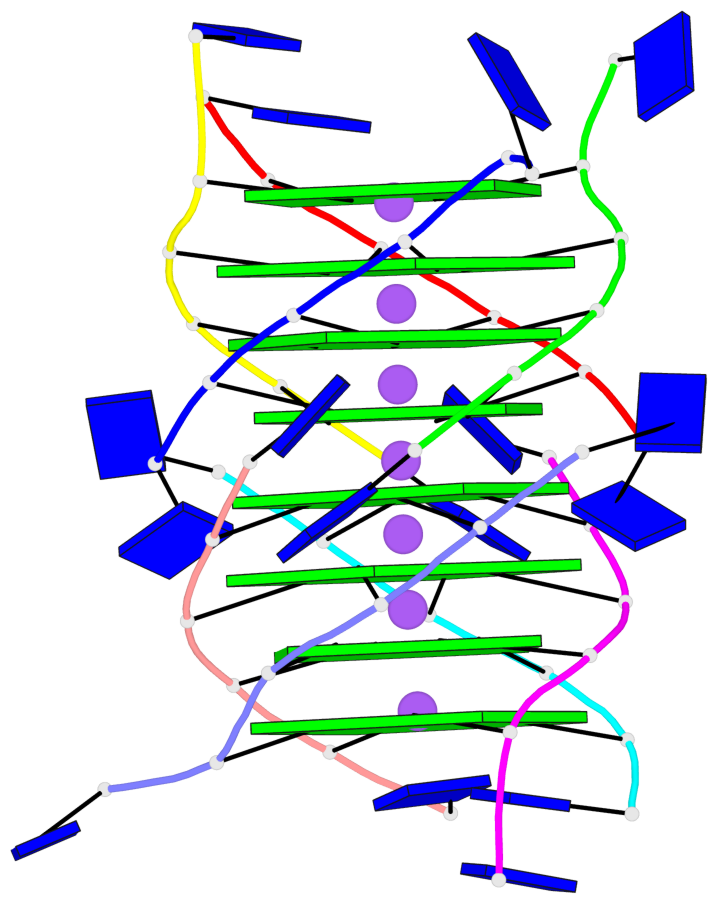
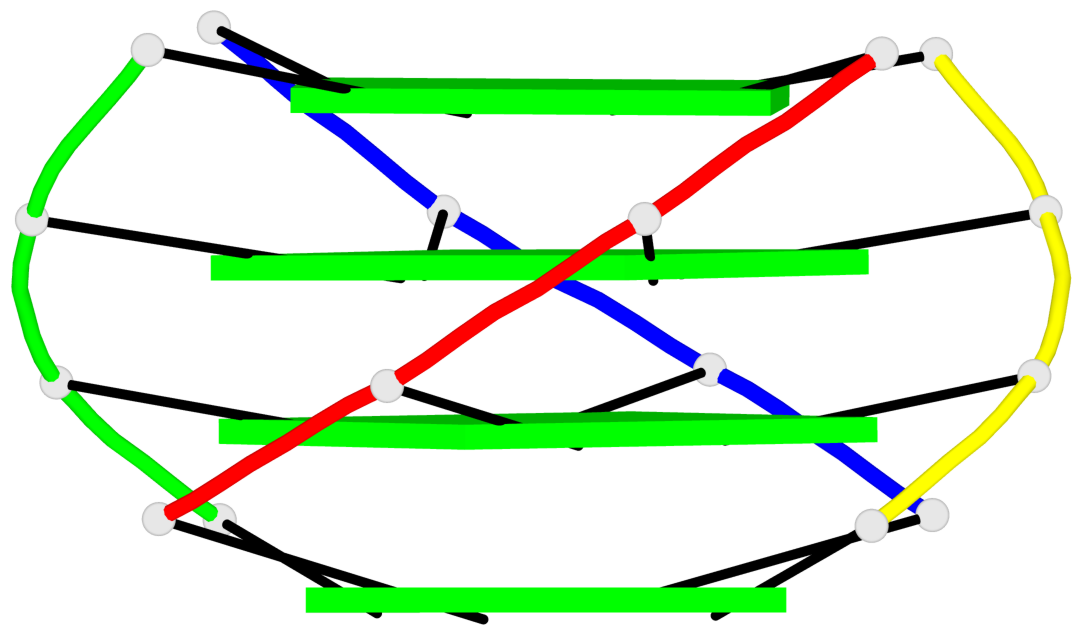
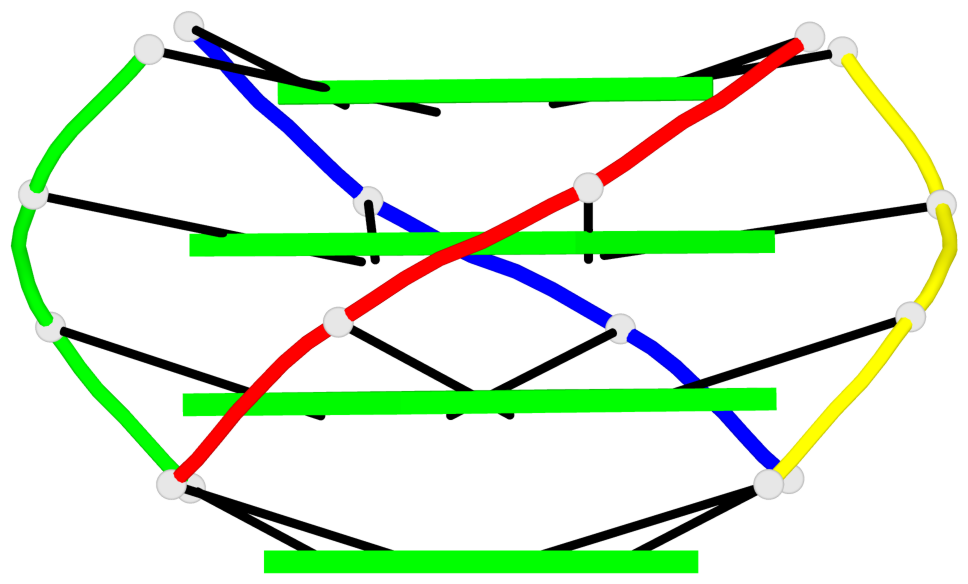
Base-block schematics in six views
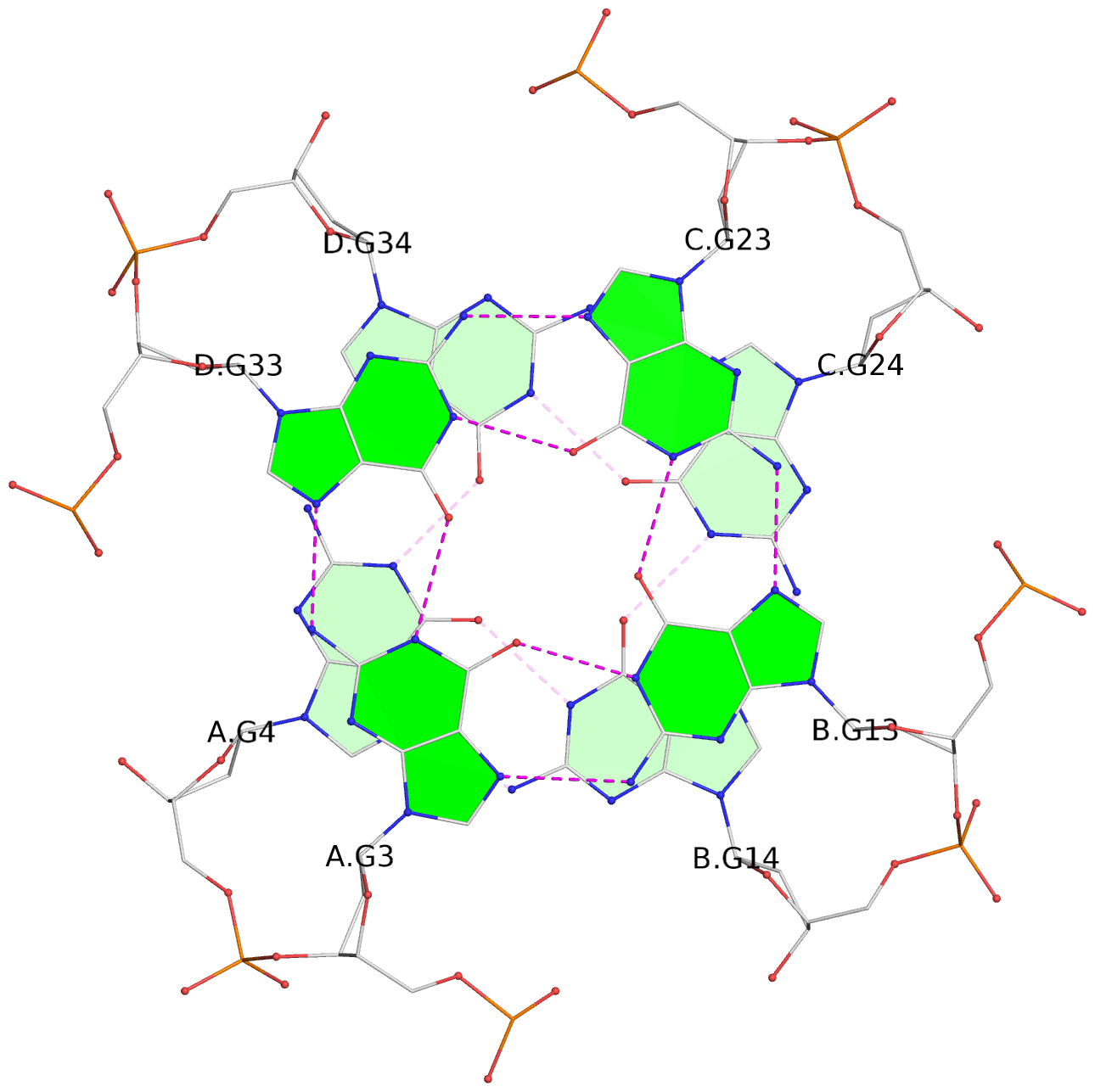
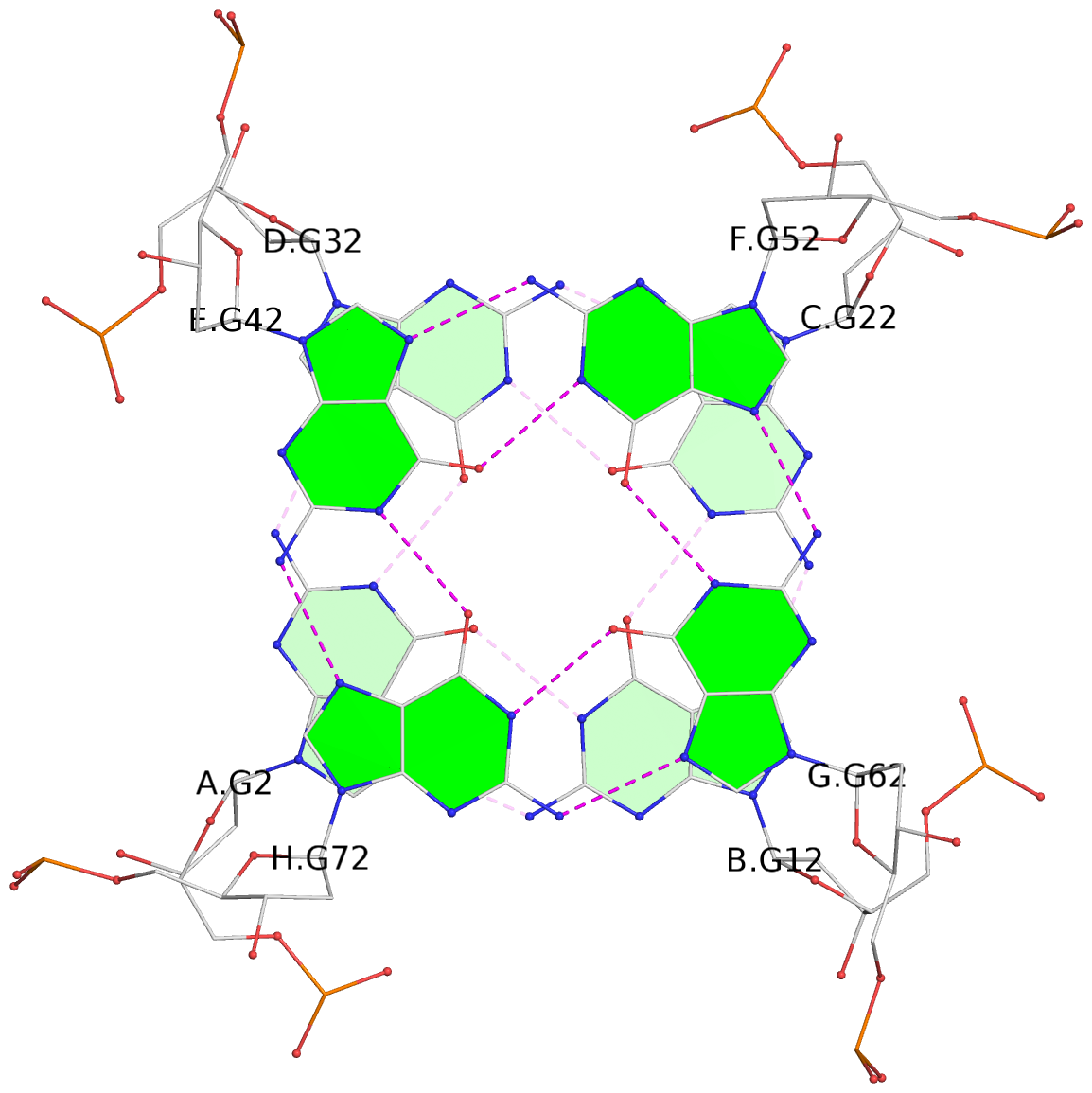
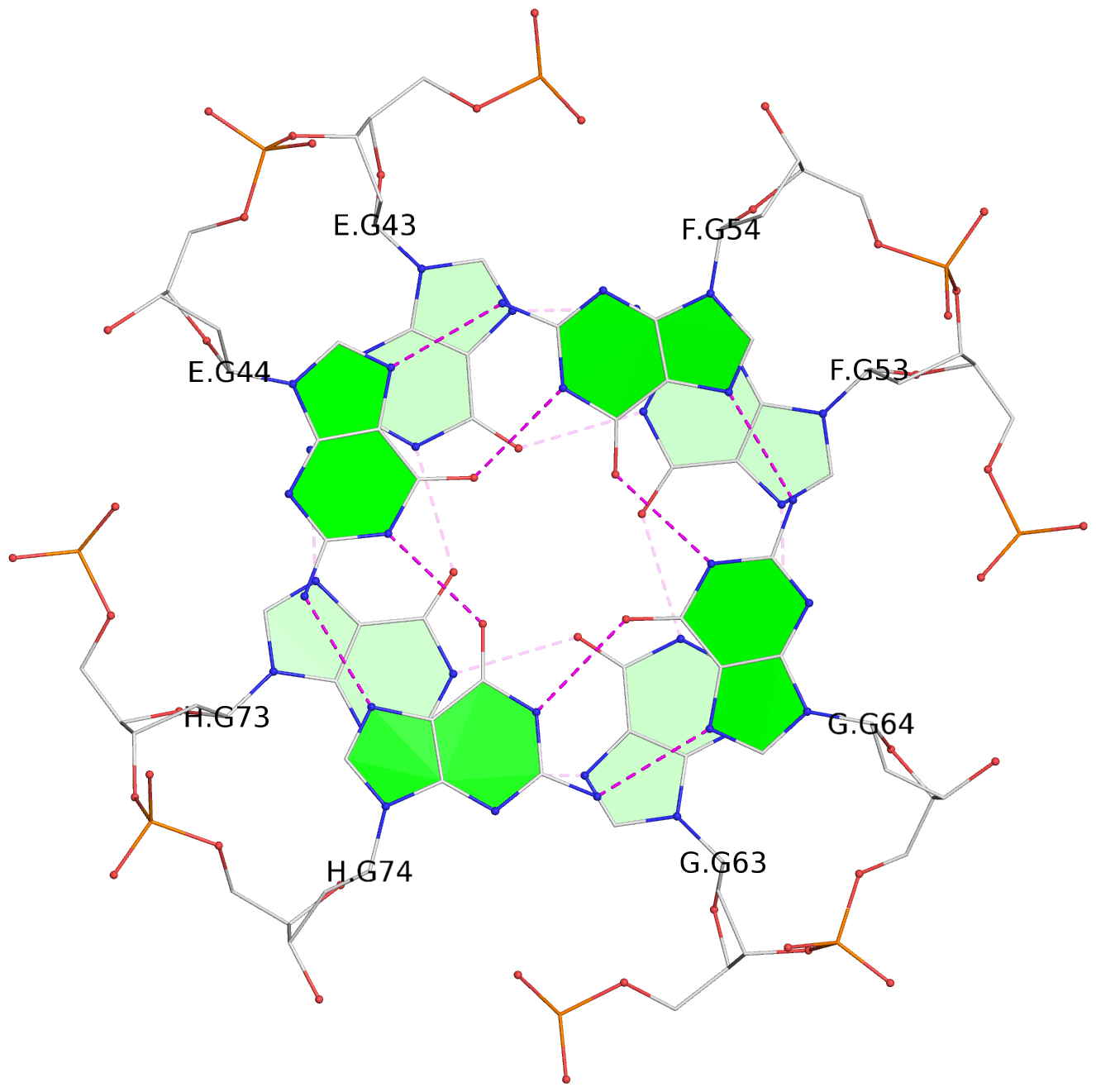
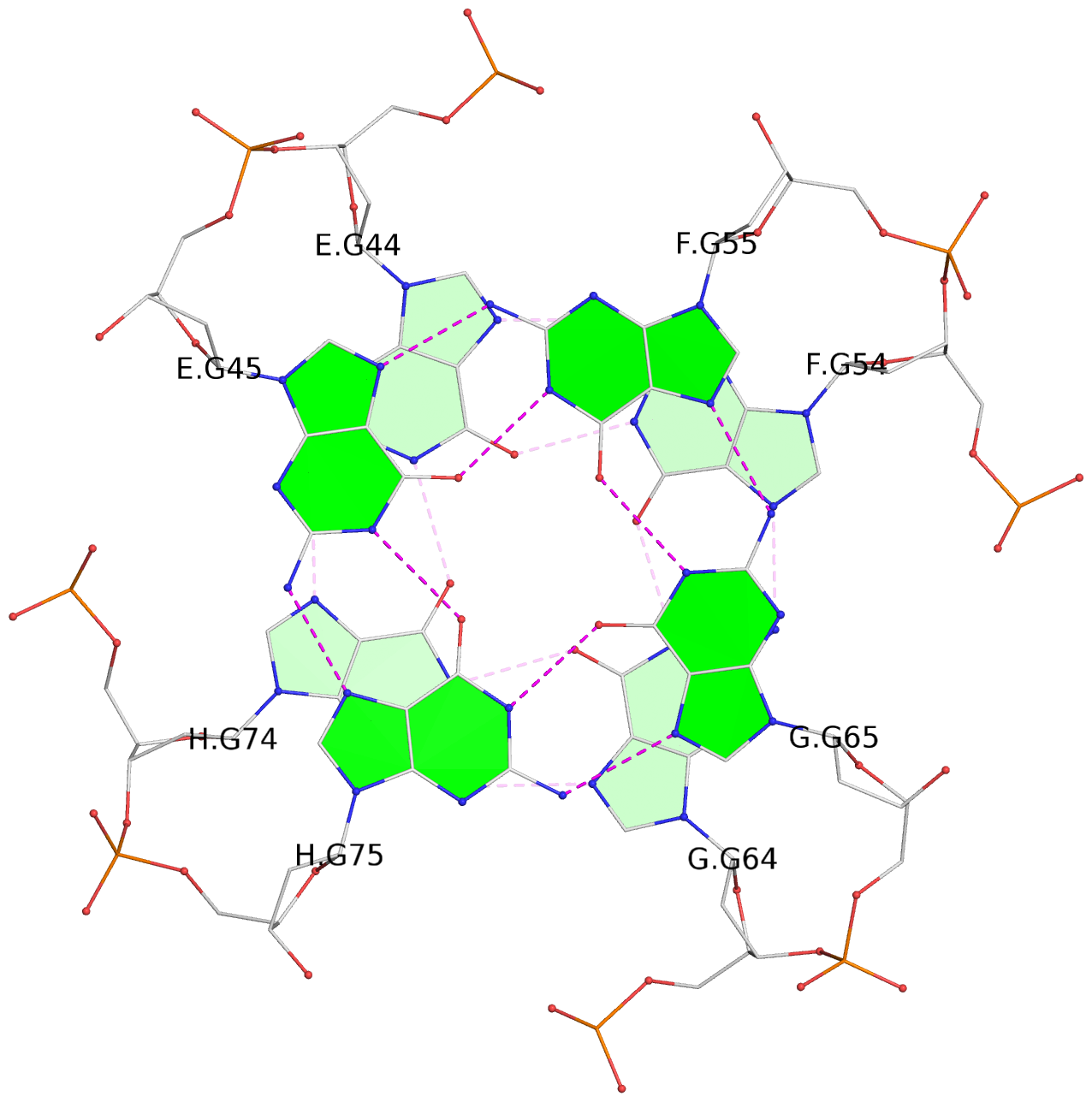
List of 8 G-tetrads
1 glyco-bond=---- sugar=---- groove=---- planarity=0.277 type=bowl nts=4 GGGG A.DG2,D.DG32,C.DG22,B.DG12 2 glyco-bond=---- sugar=---- groove=---- planarity=0.191 type=bowl nts=4 GGGG A.DG3,D.DG33,C.DG23,B.DG13 3 glyco-bond=---- sugar=---- groove=---- planarity=0.216 type=bowl nts=4 GGGG A.DG4,D.DG34,C.DG24,B.DG14 4 glyco-bond=---- sugar=---- groove=---- planarity=0.228 type=bowl nts=4 GGGG A.DG5,D.DG35,C.DG25,B.DG15 5 glyco-bond=---- sugar=3333 groove=---- planarity=0.106 type=planar nts=4 GGGG E.DG42,H.DG72,G.DG62,F.DG52 6 glyco-bond=---- sugar=---- groove=---- planarity=0.243 type=bowl nts=4 GGGG E.DG43,H.DG73,G.DG63,F.DG53 7 glyco-bond=---- sugar=---- groove=---- planarity=0.296 type=bowl nts=4 GGGG E.DG44,H.DG74,G.DG64,F.DG54 8 glyco-bond=---- sugar=---- groove=---- planarity=0.266 type=bowl nts=4 GGGG E.DG45,H.DG75,G.DG65,F.DG55
List of 1 G4-helix
In DSSR, a G4-helix is defined by stacking interactions of G-tetrads, regardless of backbone connectivity, and may contain more than one G4-stem.
Helix#1, 8 G-tetrad layers, inter-molecular, with 2 stems
List of 2 G4-stems
In DSSR, a G4-stem is defined as a G4-helix with backbone connectivity. Bulges are also allowed along each of the four strands.
Stem#1, 4 G-tetrad layers, 0 loops, inter-molecular, UUUU, parallel, parallel(4+0)
Stem#2, 4 G-tetrad layers, 0 loops, inter-molecular, UUUU, parallel, parallel(4+0)
List of 1 G4 coaxial stack
1 G4 helix#1 contains 2 G4 stems: [#1,#2] [5'/5']