Detailed DSSR results for the G-quadruplex: PDB entry 352d
Created and maintained by Xiang-Jun Lu <xiangjun@x3dna.org>
Citation: Please cite the NAR'20 DSSR-PyMOL schematics paper and/or the NAR'15 DSSR method paper.
Summary information
- PDB id
- 352d
- Class
- DNA
- Method
- X-ray (0.95 Å)
- Summary
- The crystal structure of a parallel-stranded parallel-stranded guanine tetraplex at 0.95 angstrom resolution
- Reference
- Phillips K, Dauter Z, Murchie AI, Lilley DM, Luisi B (1997): "The crystal structure of a parallel-stranded guanine tetraplex at 0.95 A resolution." J.Mol.Biol., 273, 171-182. doi: 10.1006/jmbi.1997.1292.
- Abstract
- In both DNA and RNA, stretches of guanine bases can form stable four-stranded helices in the presence of sodium or potassium ions. Sequences with a propensity to form guanine tetraplexes have been found in chromosomal telomers, immunoglobulin switch regions, and recombination sites. We report the crystal structure at 0.95 A resolution of a parallel-stranded tetraplex formed by the hexanucleotide d(TG4T) in the presence of sodium ions. The four strands form a right-handed helix that is stabilized by hydrogen-bonding tetrads of co-planar guanine bases. Well-resolved sodium ions are found between and, at defined points, within tetrad planes and are coordinated with the guanine O6 groups. Nine calcium ions have been identified, each with a well-defined hepta-coordinate hydration shell. Hydrogen-bonding water patterns are observed within the tetraplex's helical grooves and clustered about the phosphate groups. Water molecules in the groove may form a hydrogen bond with the O4', and may affect the stacking behavior of guanine. Two distinct stacking arrangements are noted for the guanine tetrads. The thymine bases do not contribute to the four-stranded conformation, but instead stack to stabilize the crystal lattice. We present evidence that the sugar conformation is strained and propose that this originates from forces that optimize guanine base stacking. Discrete conformational disorder is observed at several places in the phosphodiester backbone, which results from a simple crankshaft rotation that requires no net change in the sugar conformation.
- G4 notes
- 16 G-tetrads, 2 G4 helices, 4 G4 stems, 2 G4 coaxial stacks, parallel(4+0), UUUU, coaxial interfaces: 5'/5'
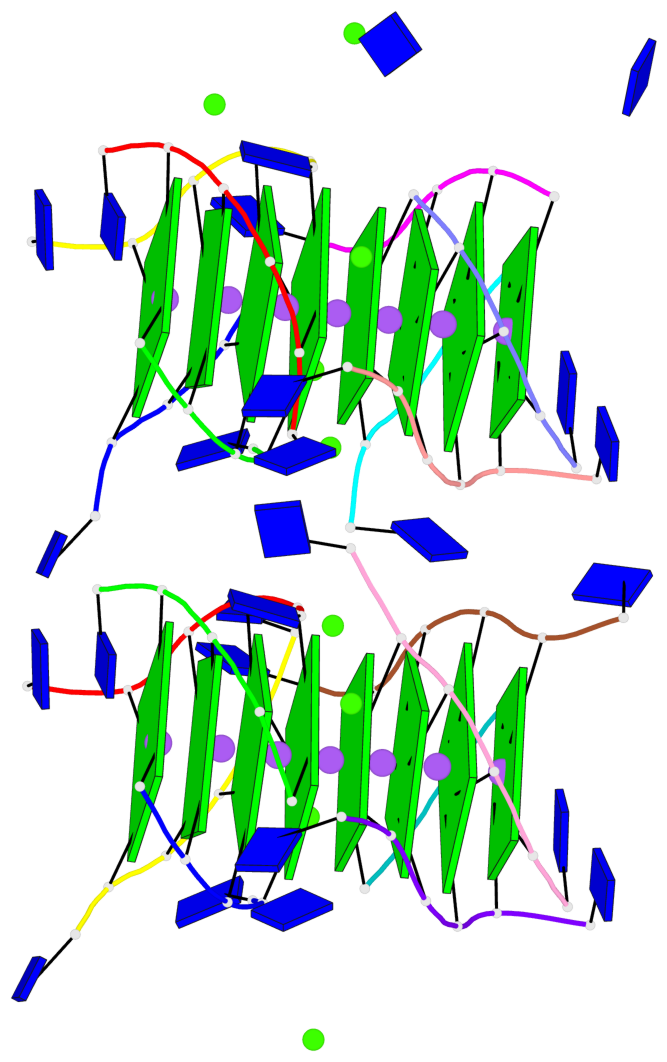
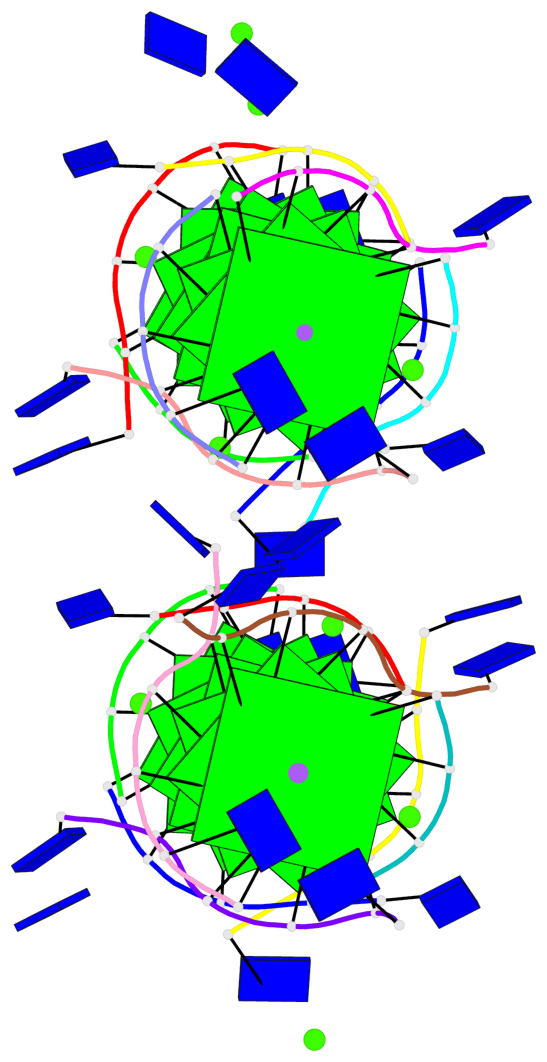
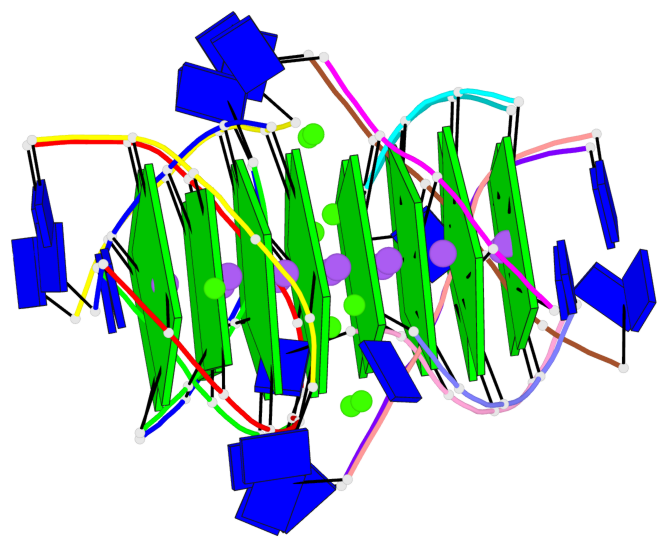
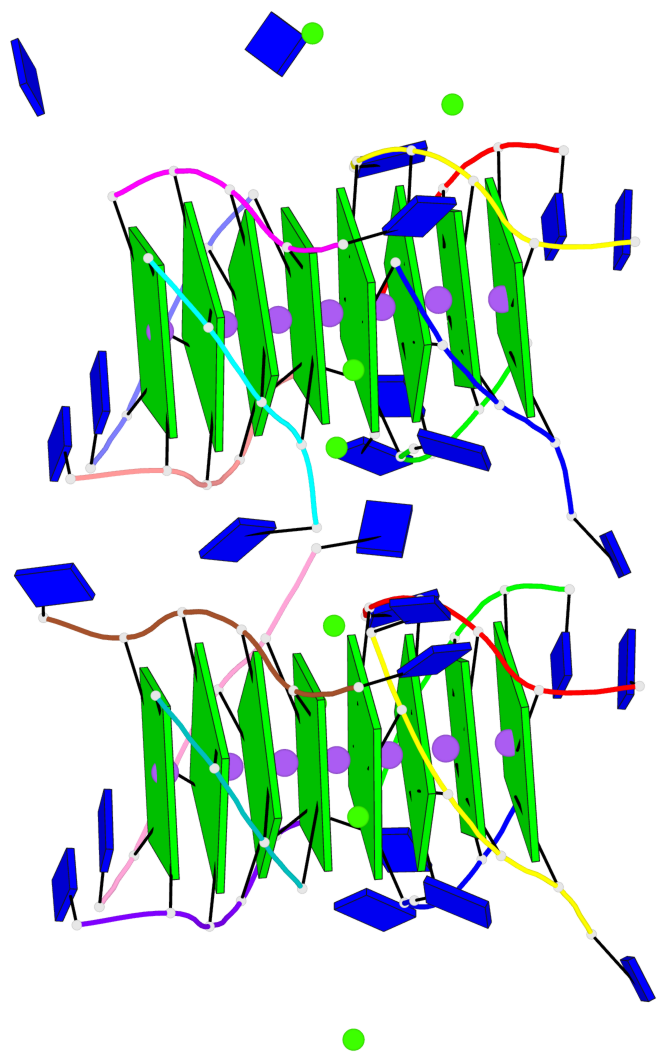
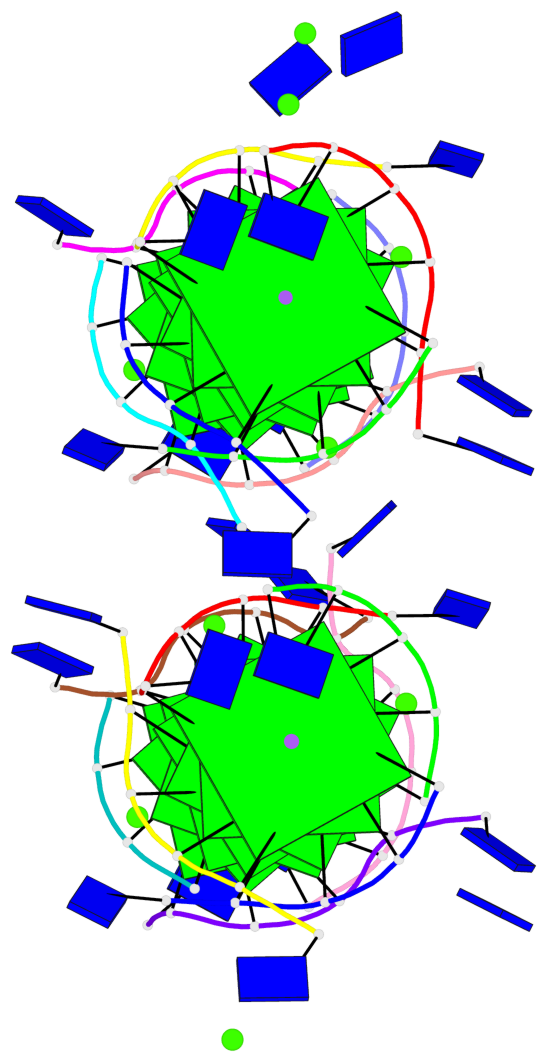
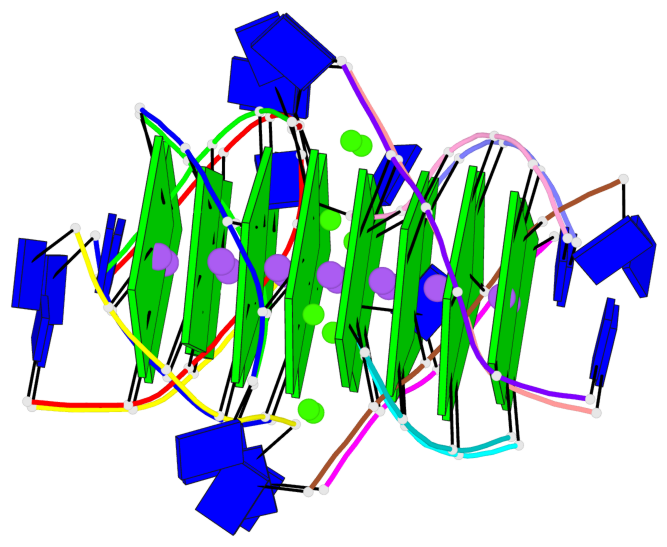
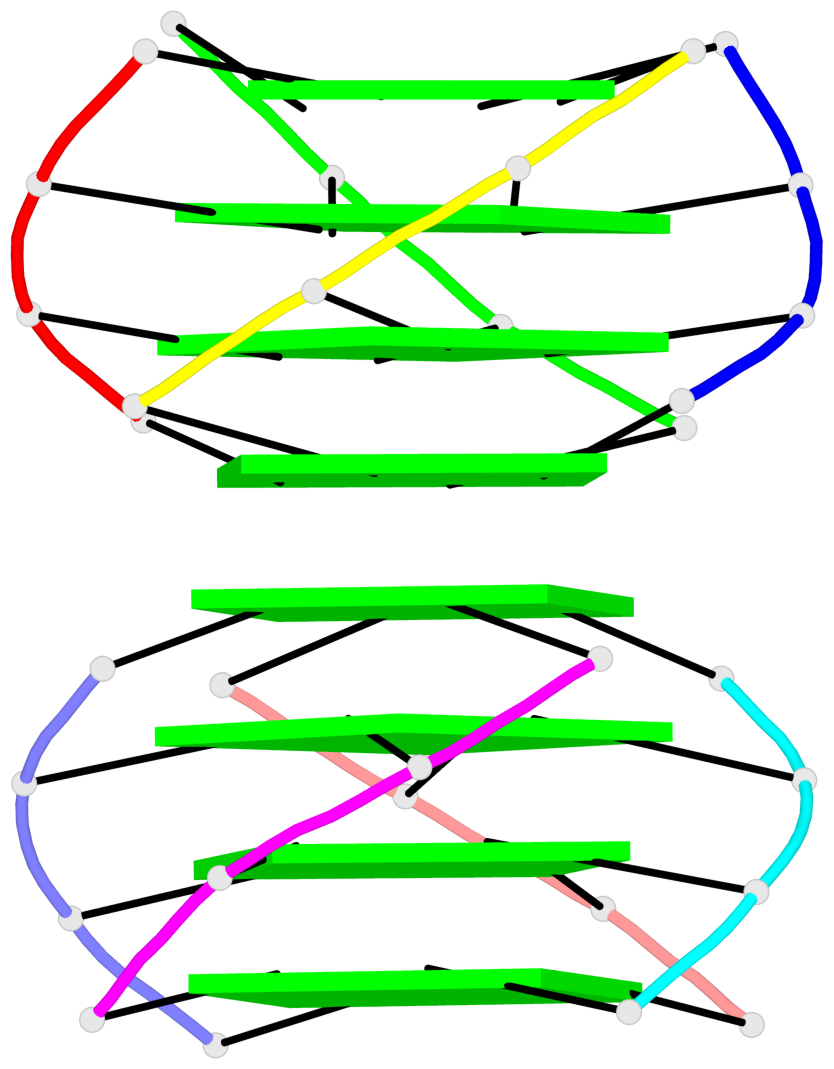
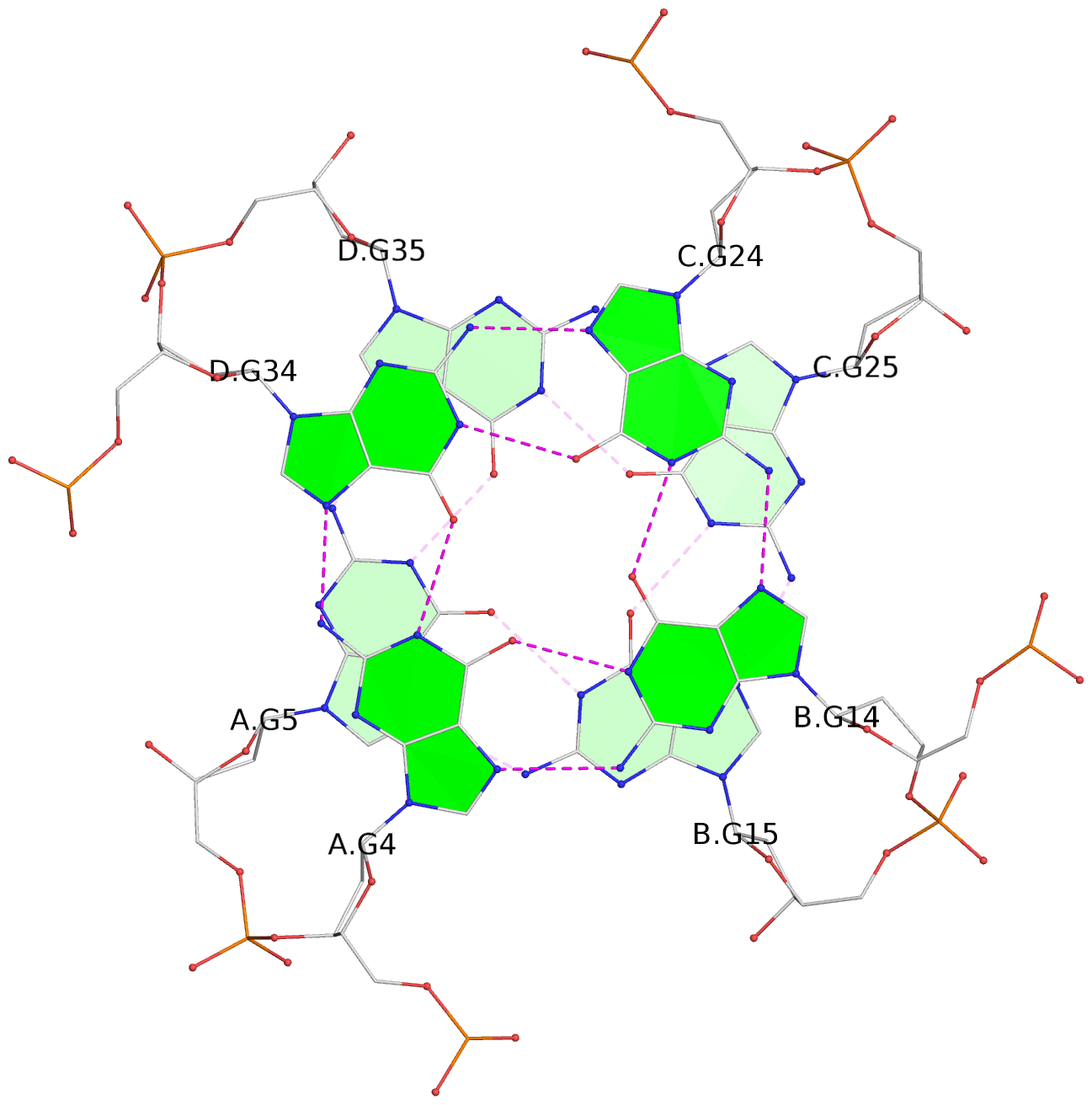
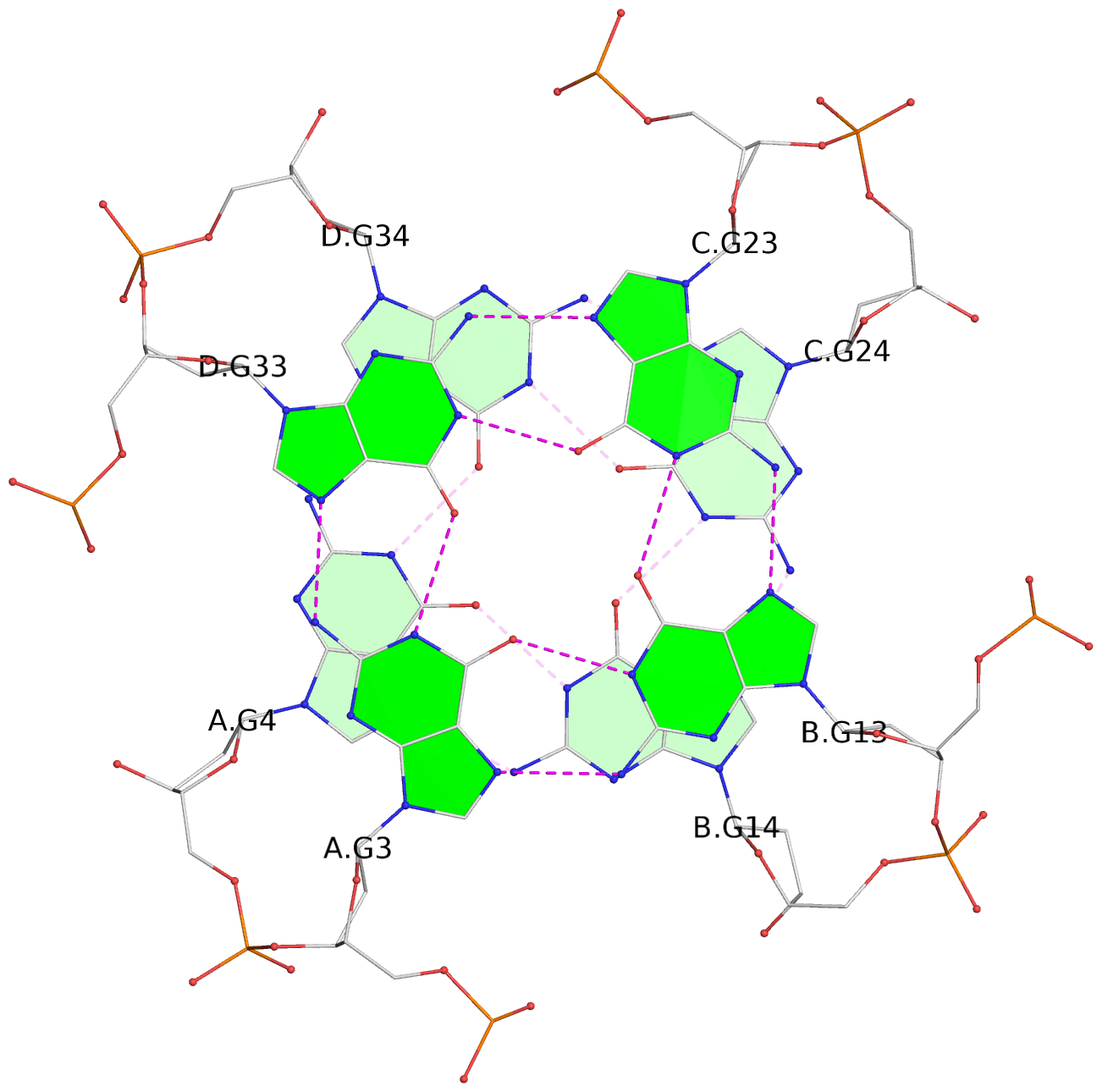
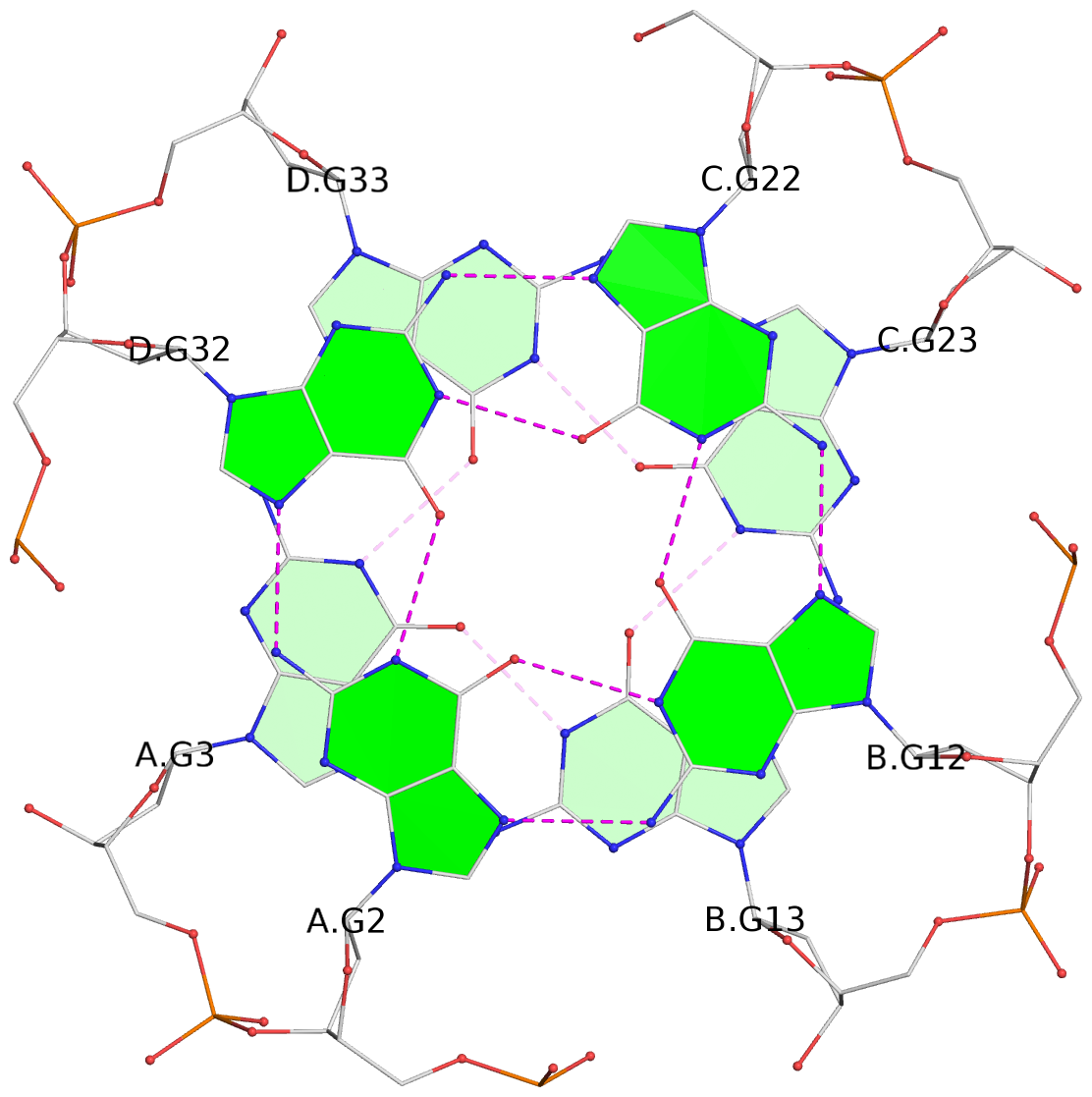
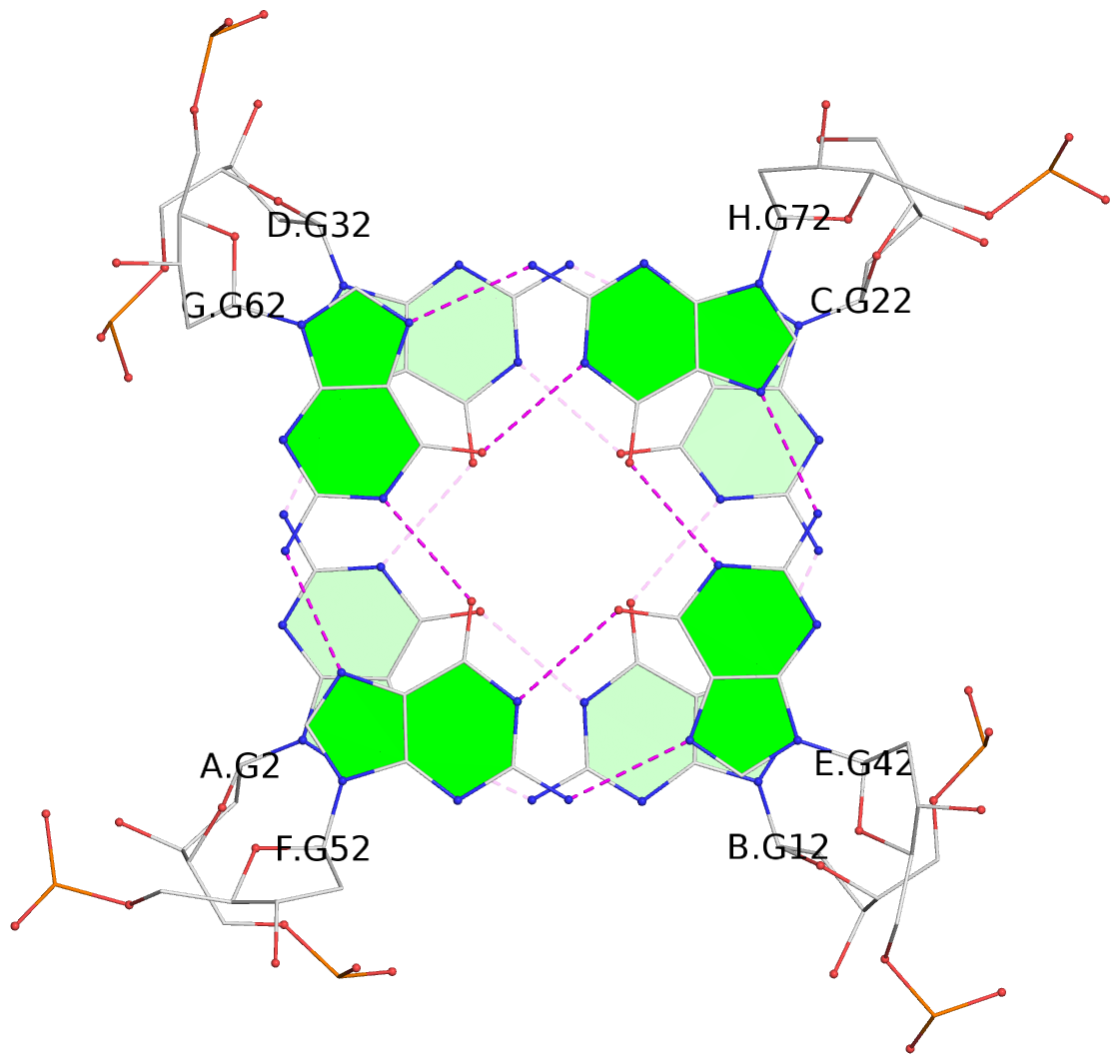
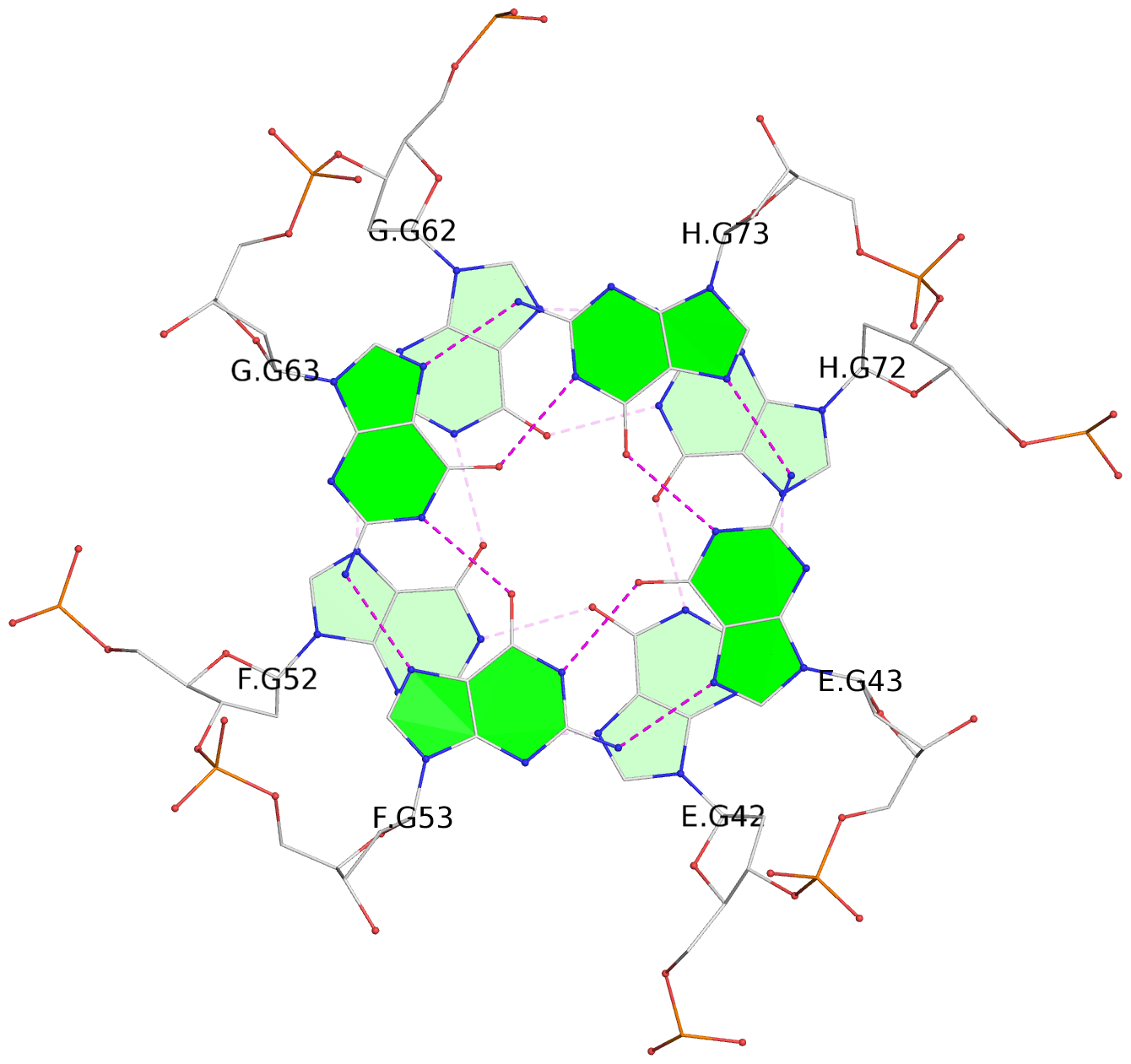
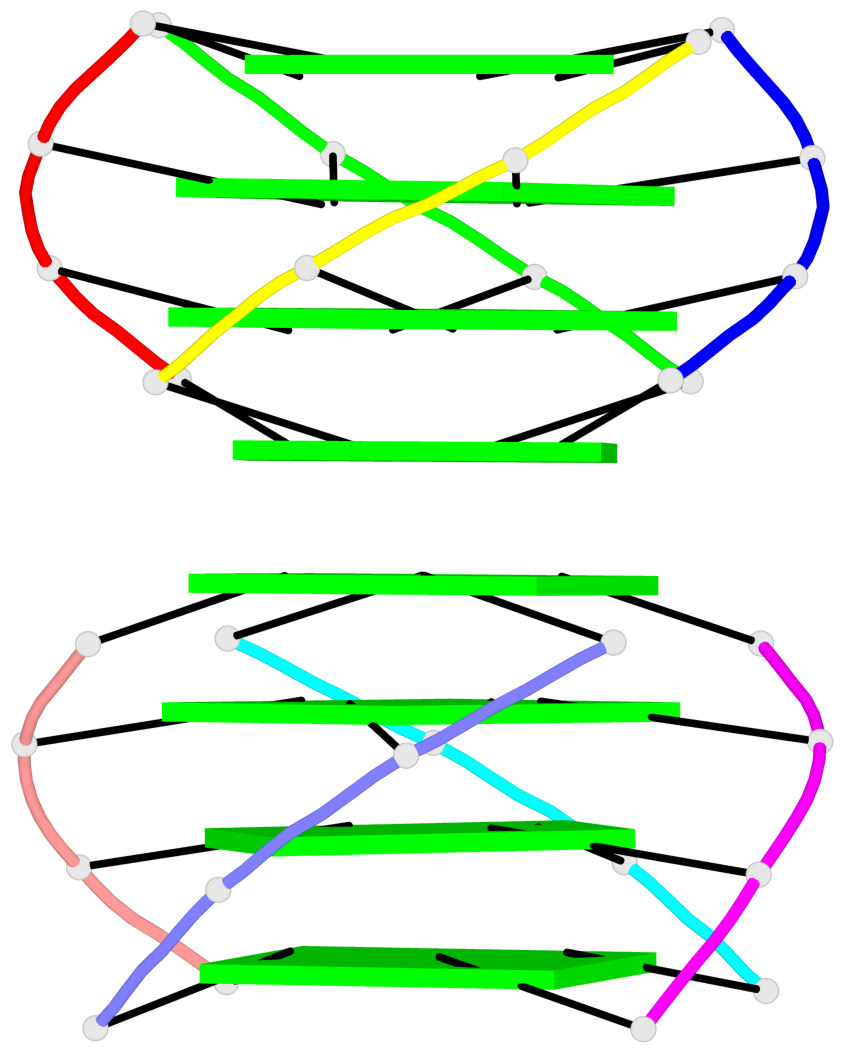
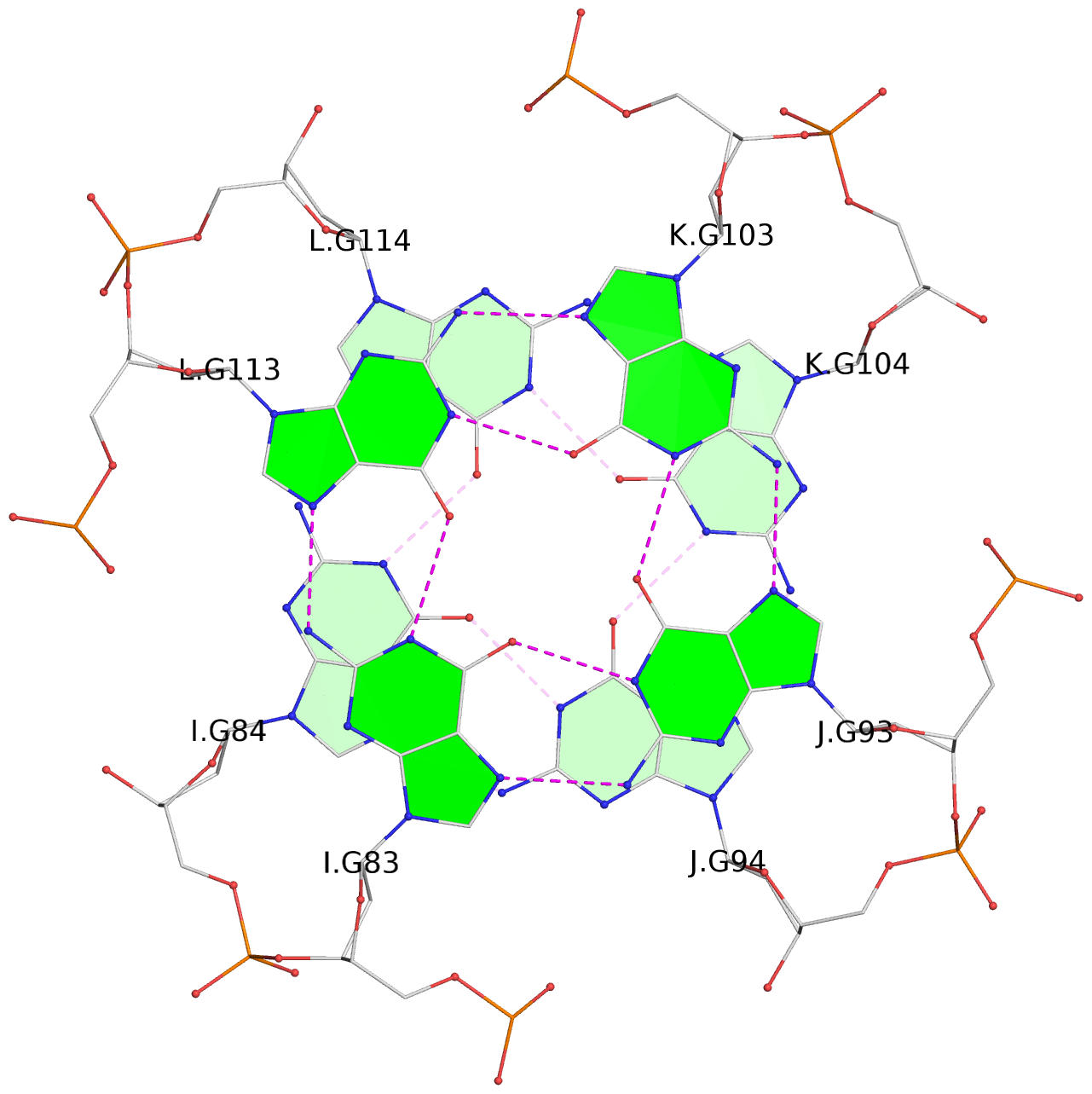
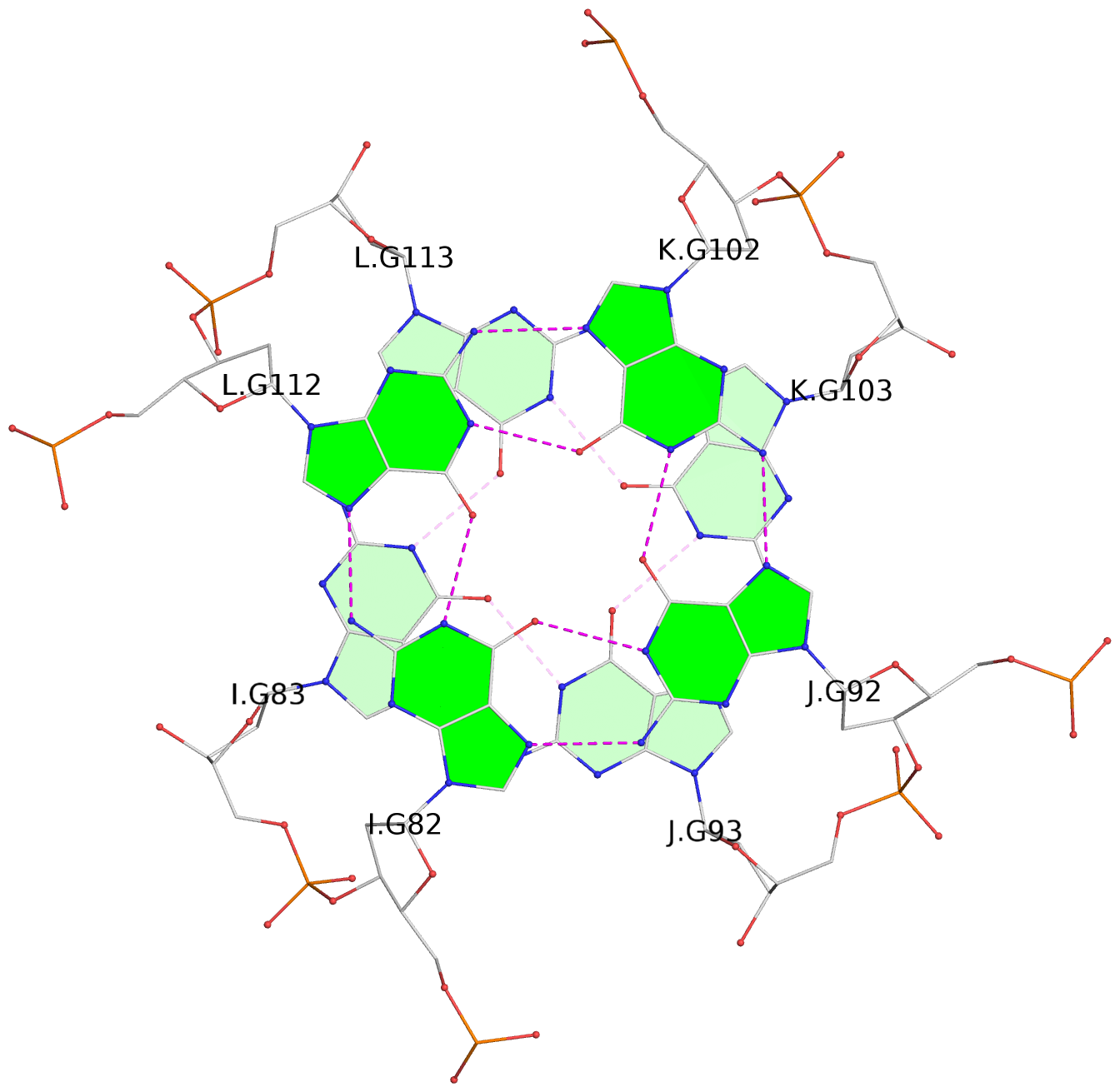
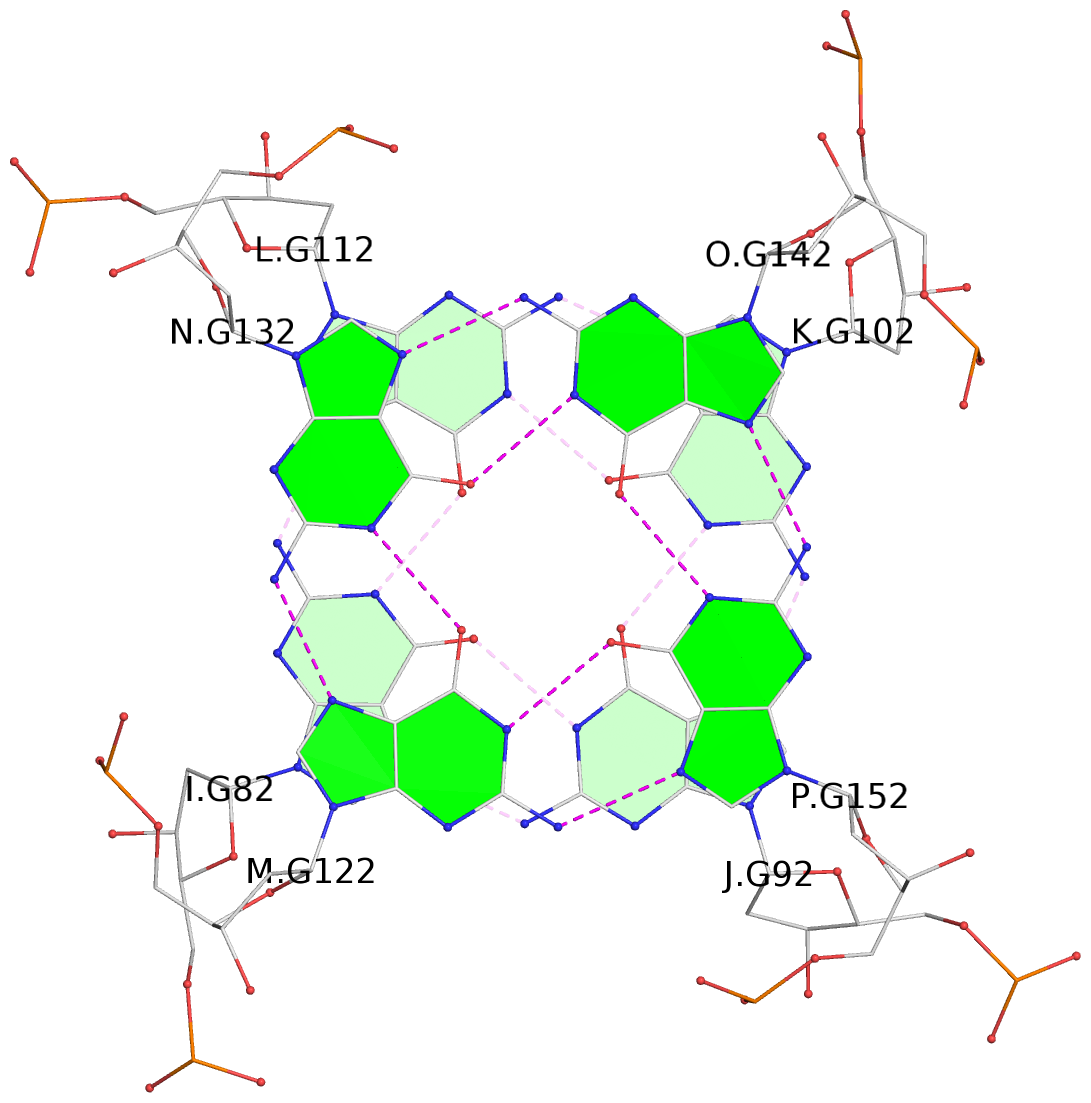
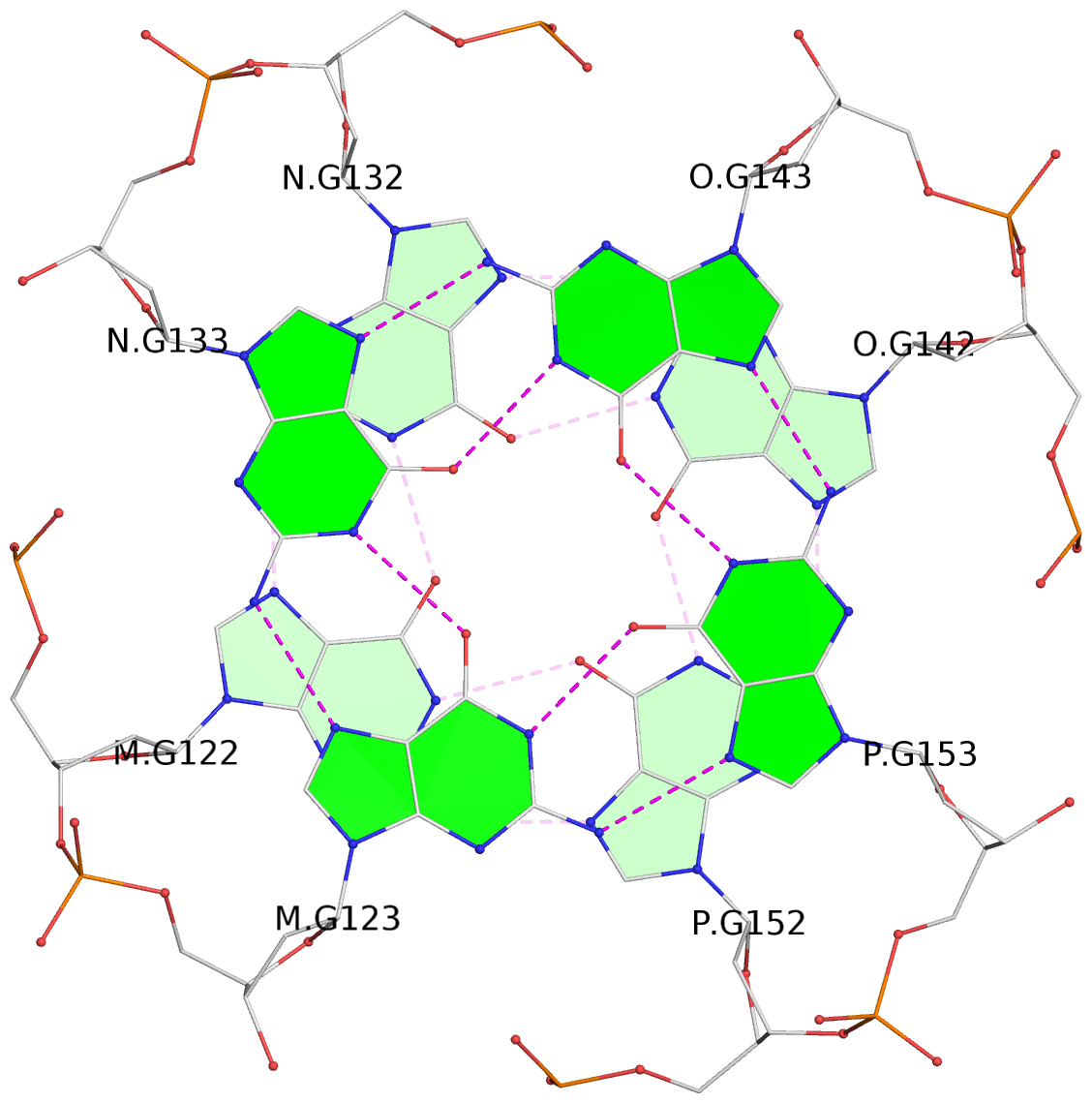
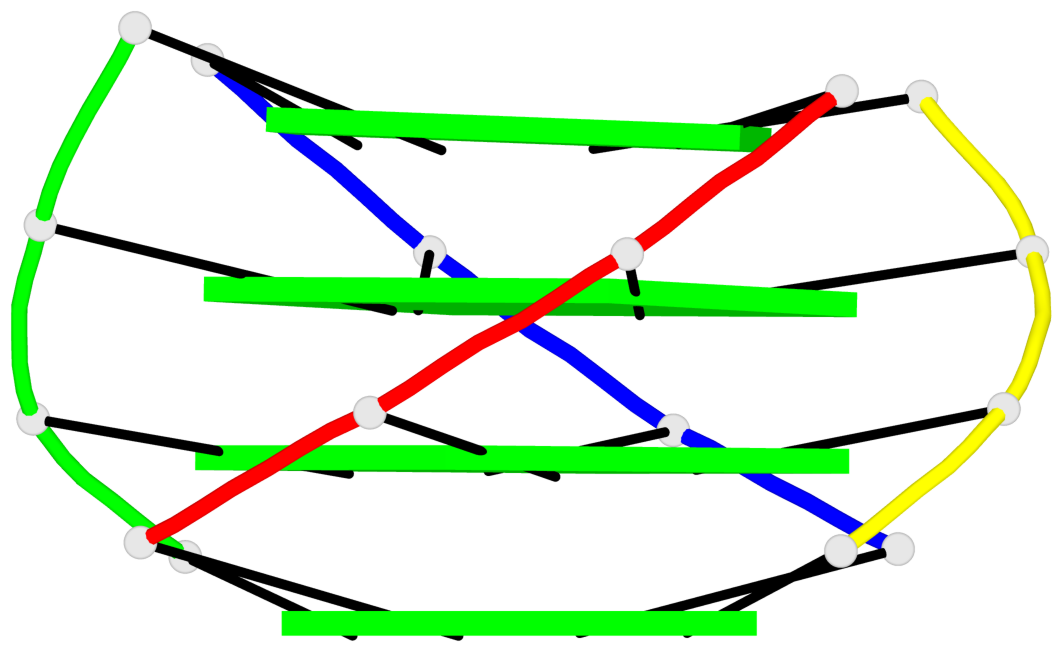
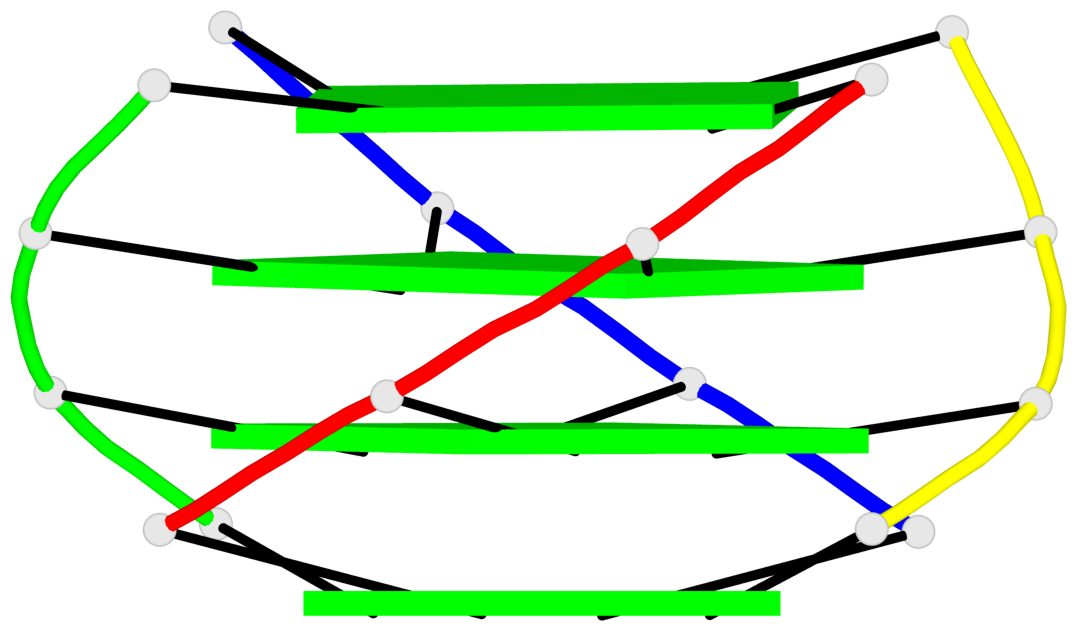
Base-block schematics in six views
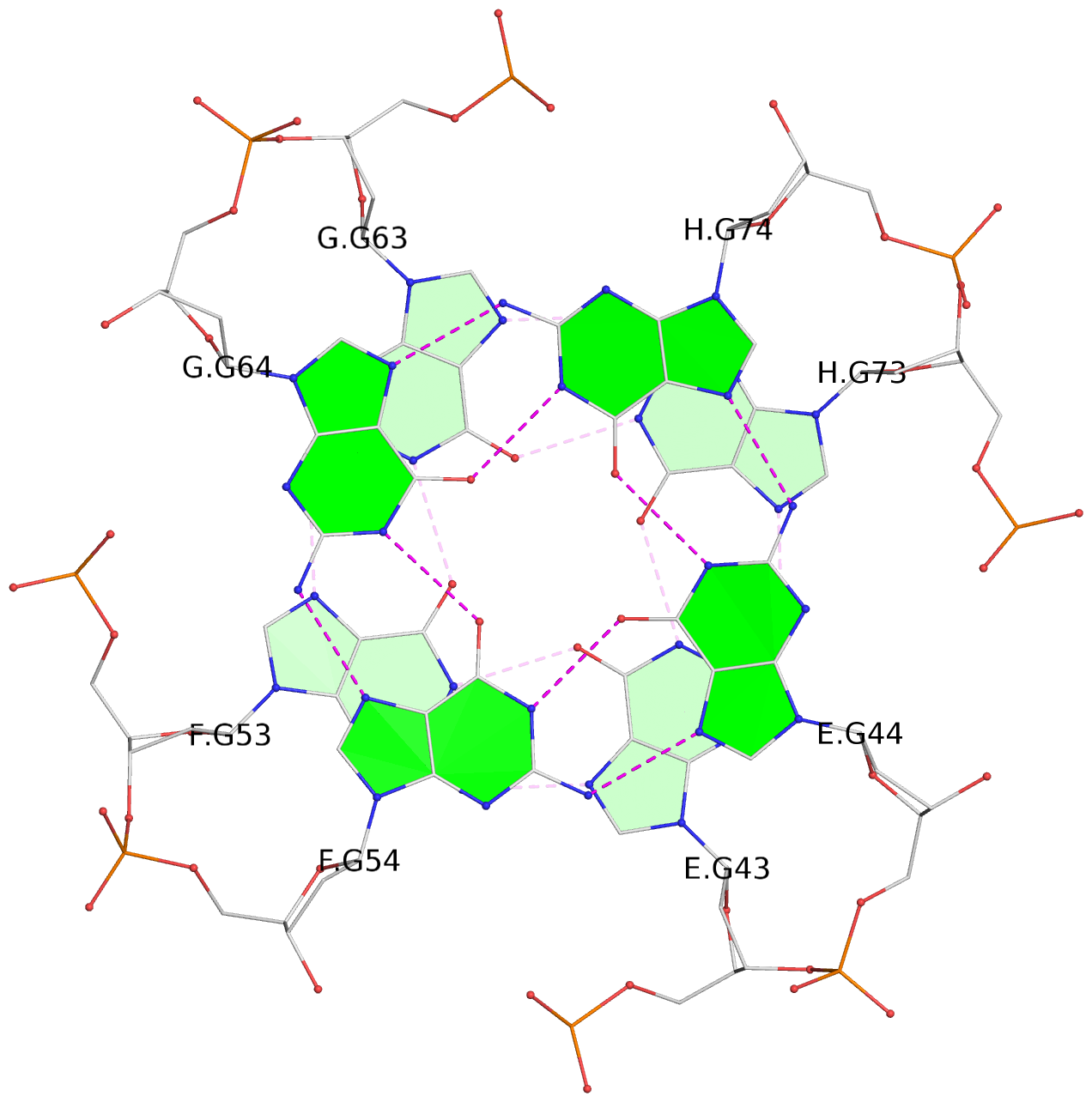
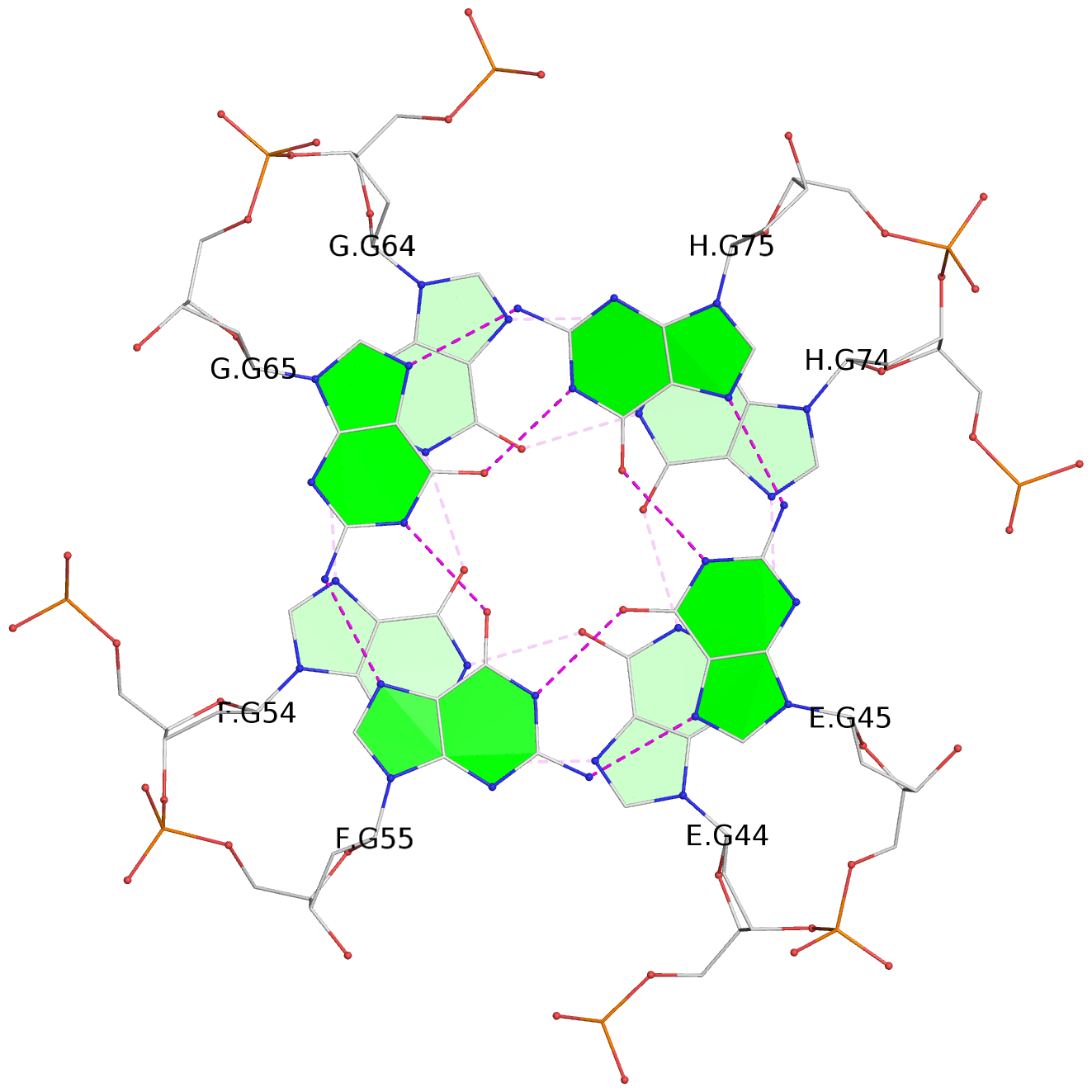
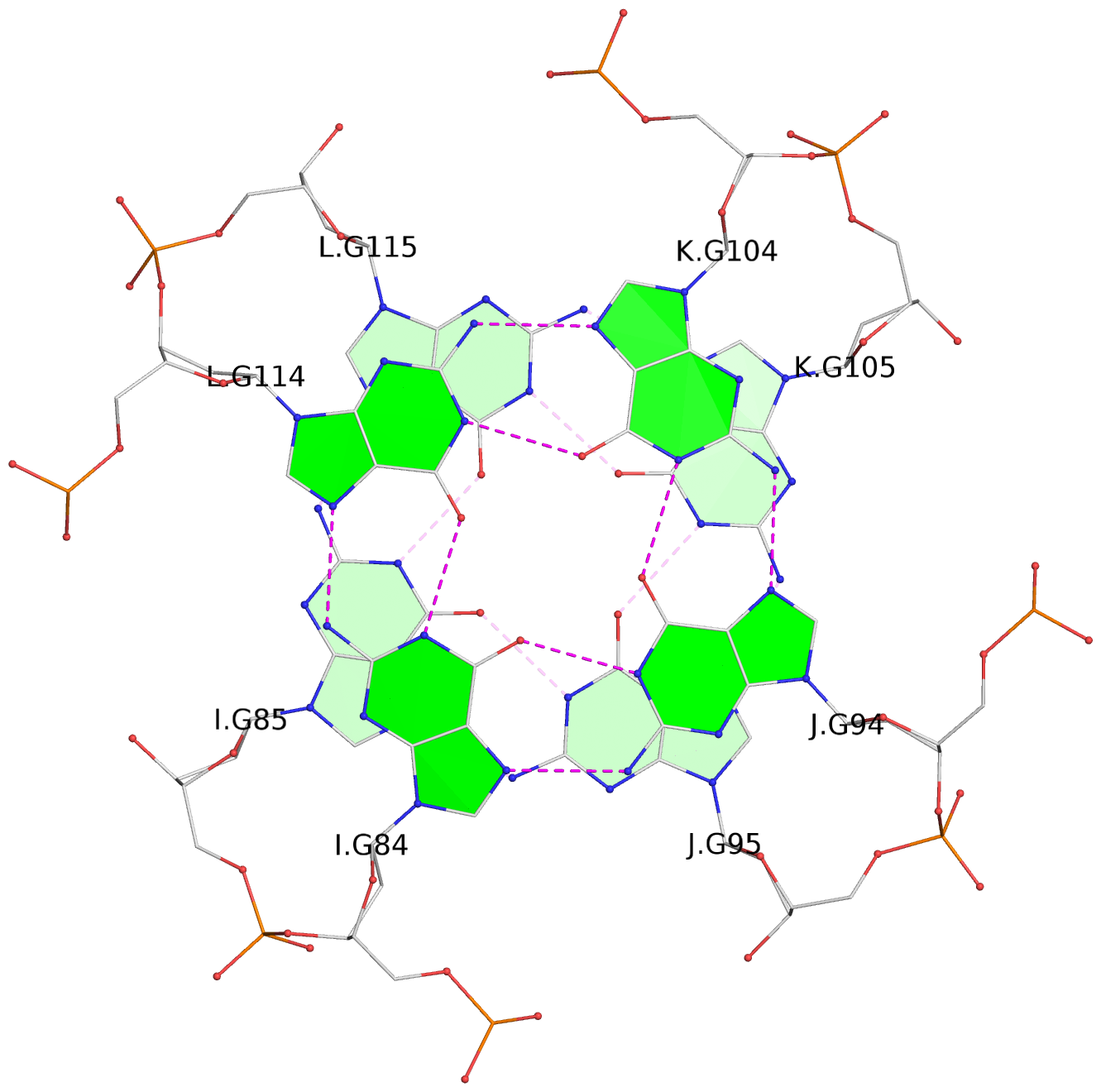
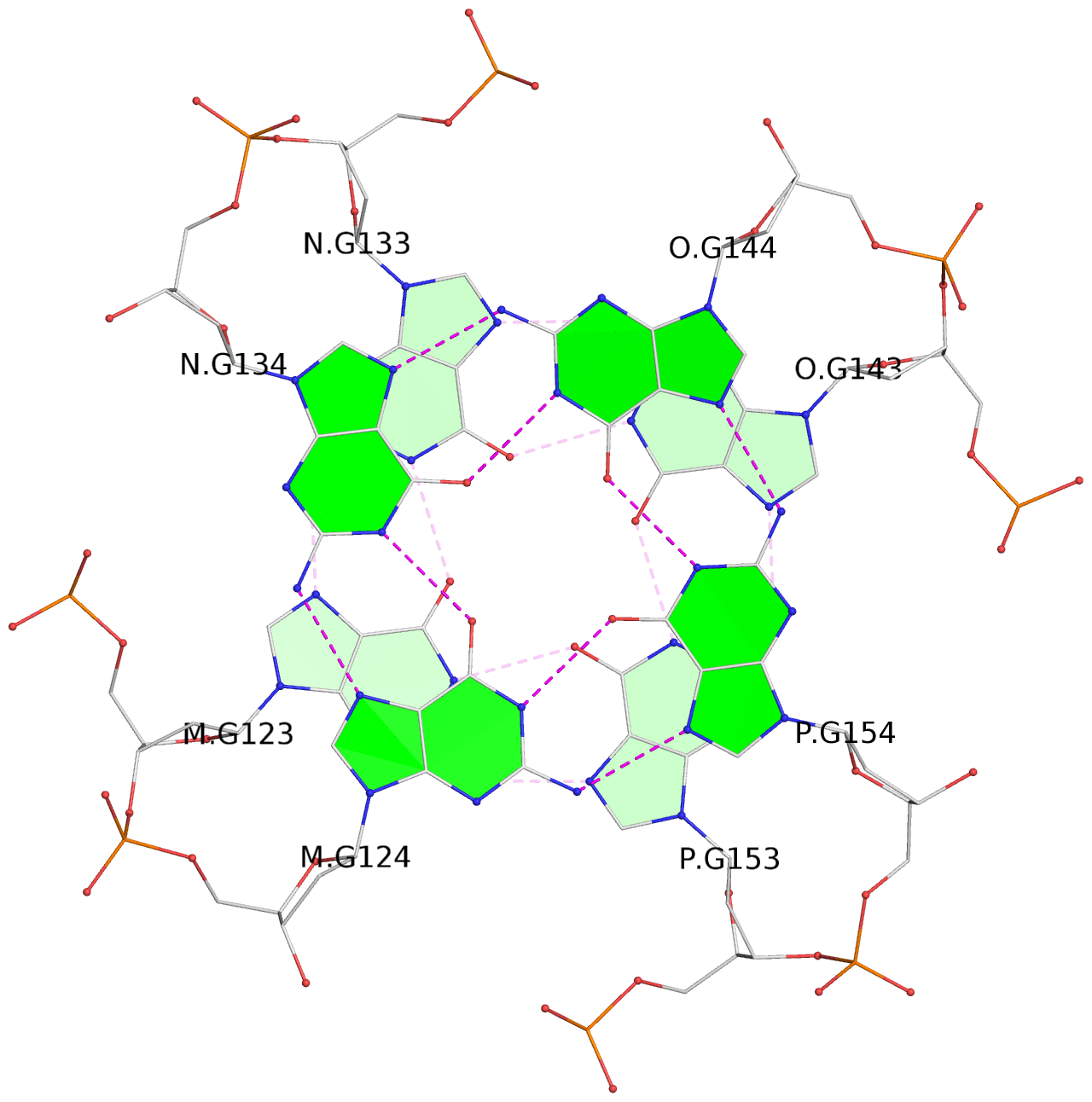
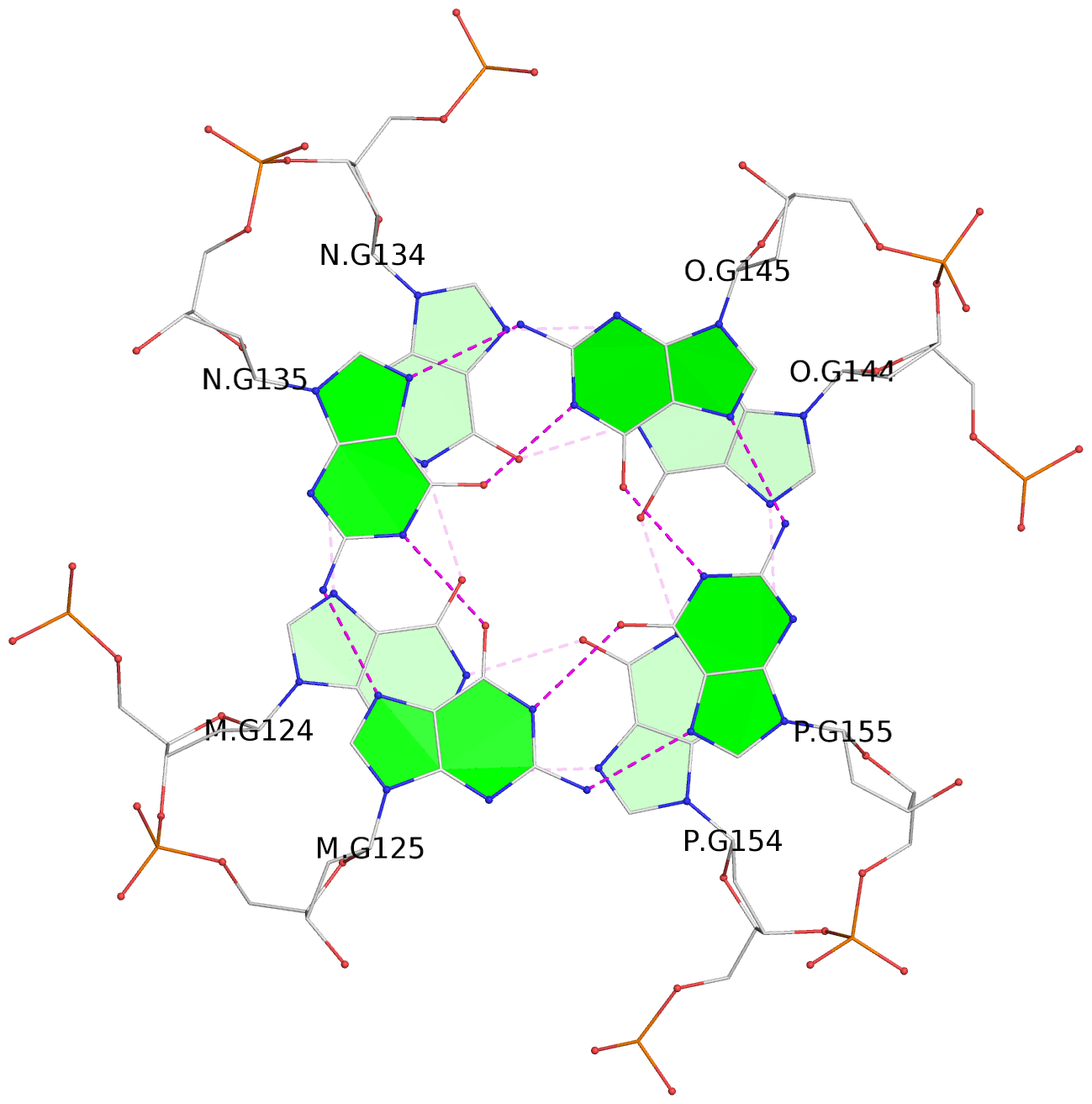
List of 16 G-tetrads
1 glyco-bond=---- sugar=---- groove=---- planarity=0.224 type=bowl nts=4 GGGG A.DG2,D.DG32,C.DG22,B.DG12 2 glyco-bond=---- sugar=---- groove=---- planarity=0.210 type=bowl nts=4 GGGG A.DG3,D.DG33,C.DG23,B.DG13 3 glyco-bond=---- sugar=---- groove=---- planarity=0.257 type=bowl nts=4 GGGG A.DG4,D.DG34,C.DG24,B.DG14 4 glyco-bond=---- sugar=---- groove=---- planarity=0.306 type=bowl nts=4 GGGG A.DG5,D.DG35,C.DG25,B.DG15 5 glyco-bond=---- sugar=3333 groove=---- planarity=0.050 type=planar nts=4 GGGG E.DG42,H.DG72,G.DG62,F.DG52 6 glyco-bond=---- sugar=---- groove=---- planarity=0.229 type=bowl nts=4 GGGG E.DG43,H.DG73,G.DG63,F.DG53 7 glyco-bond=---- sugar=---- groove=---- planarity=0.241 type=bowl nts=4 GGGG E.DG44,H.DG74,G.DG64,F.DG54 8 glyco-bond=---- sugar=-.-- groove=---- planarity=0.261 type=bowl nts=4 GGGG E.DG45,H.DG75,G.DG65,F.DG55 9 glyco-bond=---- sugar=3333 groove=---- planarity=0.050 type=planar nts=4 GGGG I.DG82,L.DG112,K.DG102,J.DG92 10 glyco-bond=---- sugar=---- groove=---- planarity=0.232 type=bowl nts=4 GGGG I.DG83,L.DG113,K.DG103,J.DG93 11 glyco-bond=---- sugar=---- groove=---- planarity=0.240 type=bowl nts=4 GGGG I.DG84,L.DG114,K.DG104,J.DG94 12 glyco-bond=---- sugar=---- groove=---- planarity=0.231 type=bowl nts=4 GGGG I.DG85,L.DG115,K.DG105,J.DG95 13 glyco-bond=---- sugar=---- groove=---- planarity=0.219 type=bowl nts=4 GGGG M.DG122,P.DG152,O.DG142,N.DG132 14 glyco-bond=---- sugar=---- groove=---- planarity=0.210 type=bowl nts=4 GGGG M.DG123,P.DG153,O.DG143,N.DG133 15 glyco-bond=---- sugar=---- groove=---- planarity=0.226 type=bowl nts=4 GGGG M.DG124,P.DG154,O.DG144,N.DG134 16 glyco-bond=---- sugar=---- groove=---- planarity=0.306 type=bowl nts=4 GGGG M.DG125,P.DG155,O.DG145,N.DG135
List of 2 G4-helices
In DSSR, a G4-helix is defined by stacking interactions of G-tetrads, regardless of backbone connectivity, and may contain more than one G4-stem.
Helix#1, 8 G-tetrad layers, inter-molecular, with 2 stems
Helix#2, 8 G-tetrad layers, inter-molecular, with 2 stems
List of 4 G4-stems
In DSSR, a G4-stem is defined as a G4-helix with backbone connectivity. Bulges are also allowed along each of the four strands.
Stem#1, 4 G-tetrad layers, 0 loops, inter-molecular, UUUU, parallel, parallel(4+0)
Stem#2, 4 G-tetrad layers, 0 loops, inter-molecular, UUUU, parallel, parallel(4+0)
Stem#3, 4 G-tetrad layers, 0 loops, inter-molecular, UUUU, parallel, parallel(4+0)
Stem#4, 4 G-tetrad layers, 0 loops, inter-molecular, UUUU, parallel, parallel(4+0)
List of 2 G4 coaxial stacks
1 G4 helix#1 contains 2 G4 stems: [#1,#2] [5'/5'] 2 G4 helix#2 contains 2 G4 stems: [#3,#4] [5'/5']