Detailed DSSR results for the G-quadruplex: PDB entry 3cco
Created and maintained by Xiang-Jun Lu <xiangjun@x3dna.org>
Citation: Please cite the NAR'20 DSSR-PyMOL schematics paper and/or the NAR'15 DSSR method paper.
Summary information
- PDB id
- 3cco
- Class
- DNA
- Method
- X-ray (2.2 Å)
- Summary
- Structural adaptation and conservation in quadruplex-drug recognition
- Reference
- Parkinson GN, Cuenca F, Neidle S (2008): "Topology conservation and loop flexibility in quadruplex-drug recognition: crystal structures of inter- and intramolecular telomeric DNA quadruplex-drug complexes." J.Mol.Biol., 381, 1145-1156. doi: 10.1016/j.jmb.2008.06.022.
- Abstract
- Knowledge of the biologically relevant topology is critical for the design of drugs targeting quadruplex nucleic acids. We report here crystal structures of a G-quadruplex-selective ligand complexed with two human telomeric DNA quadruplexes. The intramolecular quadruplex sequence d[TAGGG(TTAGGG)(3)] and the bimolecular quadruplex sequence d(TAGGGTTAGGGT) were co-crystallized with a tetra-substituted naphthalene diimide quadruplex-binding ligand. The structures were solved and refined to 2.10- and 2.20-A resolution, respectively, revealing that the quadruplex topology in both structures is unchanged by the addition of the ligands, retaining a parallel-stranded arrangement with external double-chain-reversal propeller loops. The parallel topology results in accessible external 5' and 3' planar G-tetrad surfaces for ligand stacking. This also enables significant ligand-induced conformational changes in several TTA propeller loops to take place such that the loops themselves are able to accommodate bound drug molecules without affecting the parallel quadruplex topology, by stacking on the external TTA connecting loop nucleotides. Ligands are bound into the external TTA loop nucleotides and stack onto G-tetrad surfaces. These crystal structures provide a framework for further ligand development of the naphthalene diimide series to enhance selectivity and affinity.
- G4 notes
- 3 G-tetrads, 1 G4 helix, 1 G4 stem, parallel(4+0), UUUU
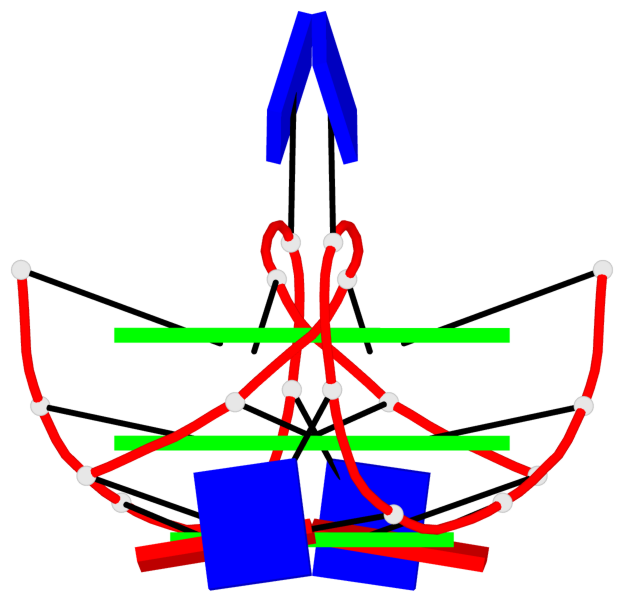
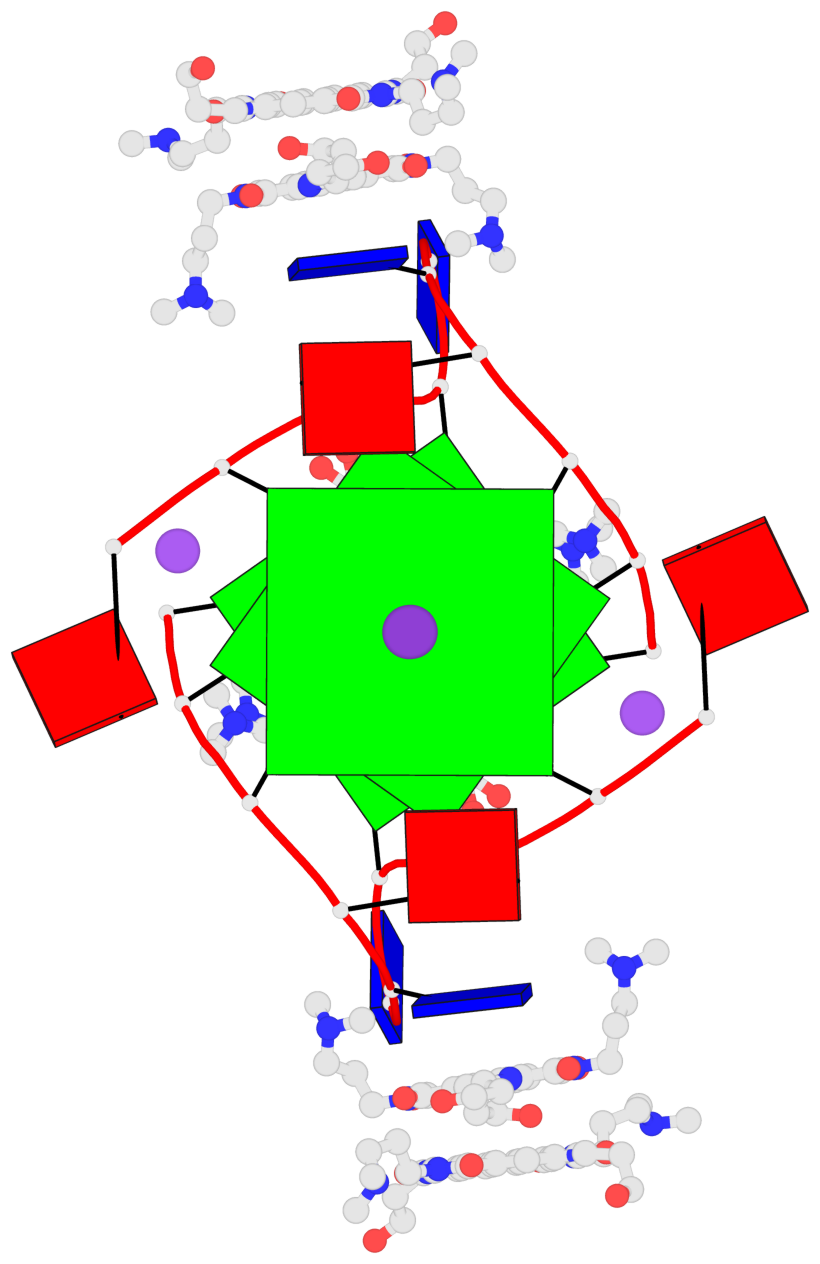
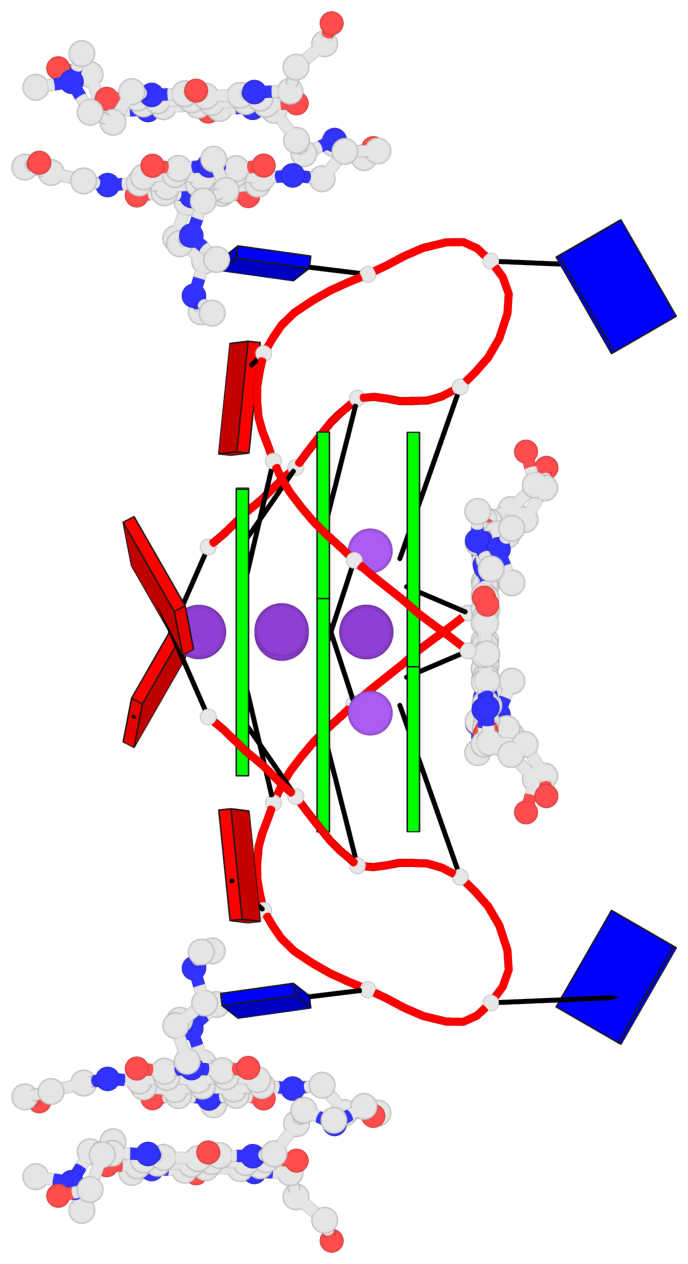
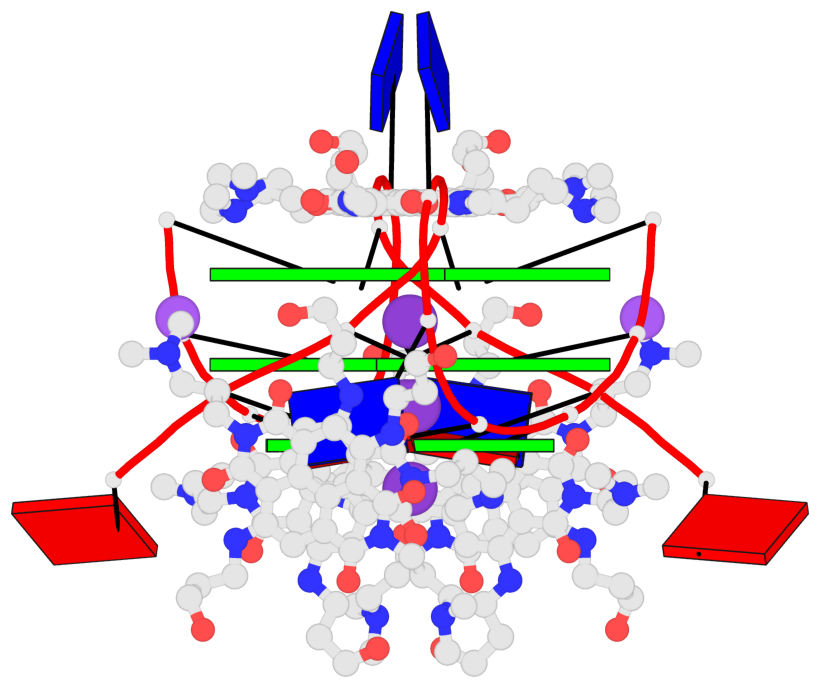
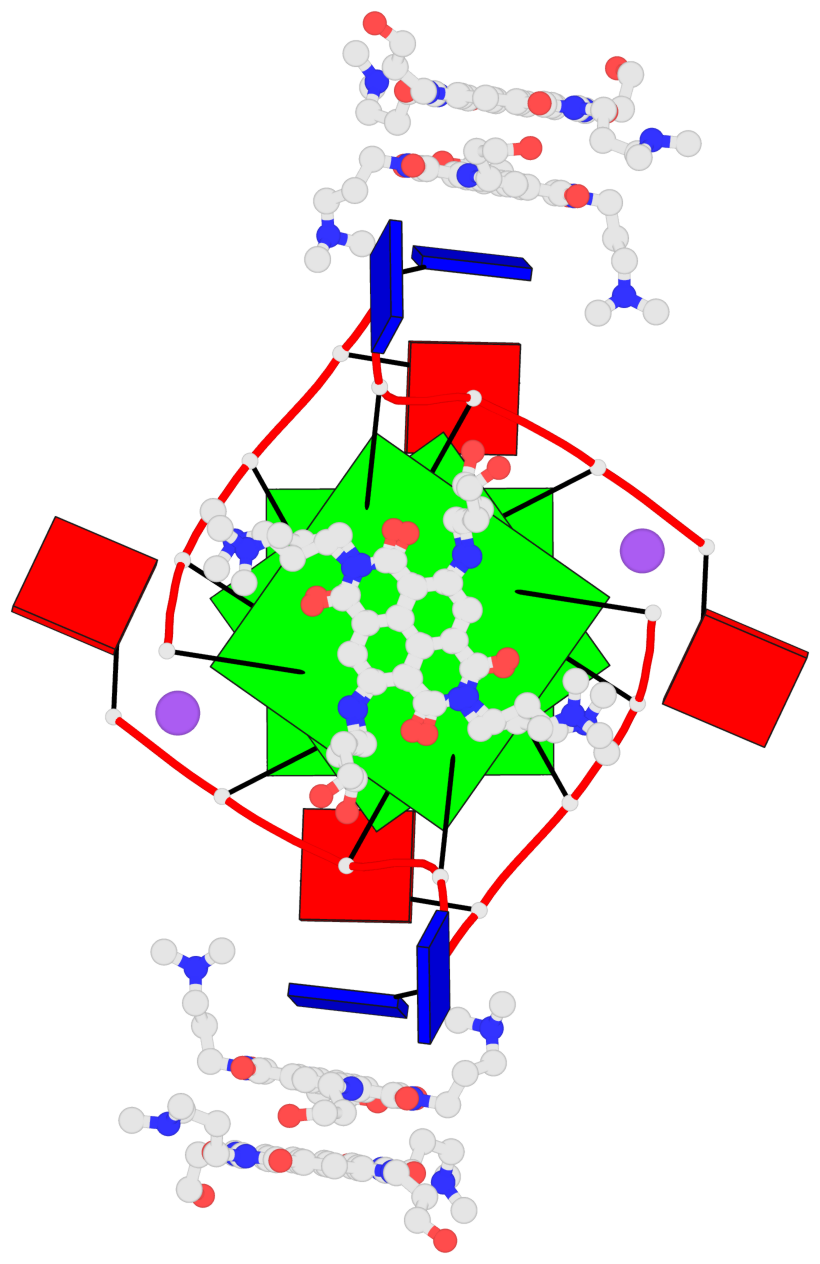
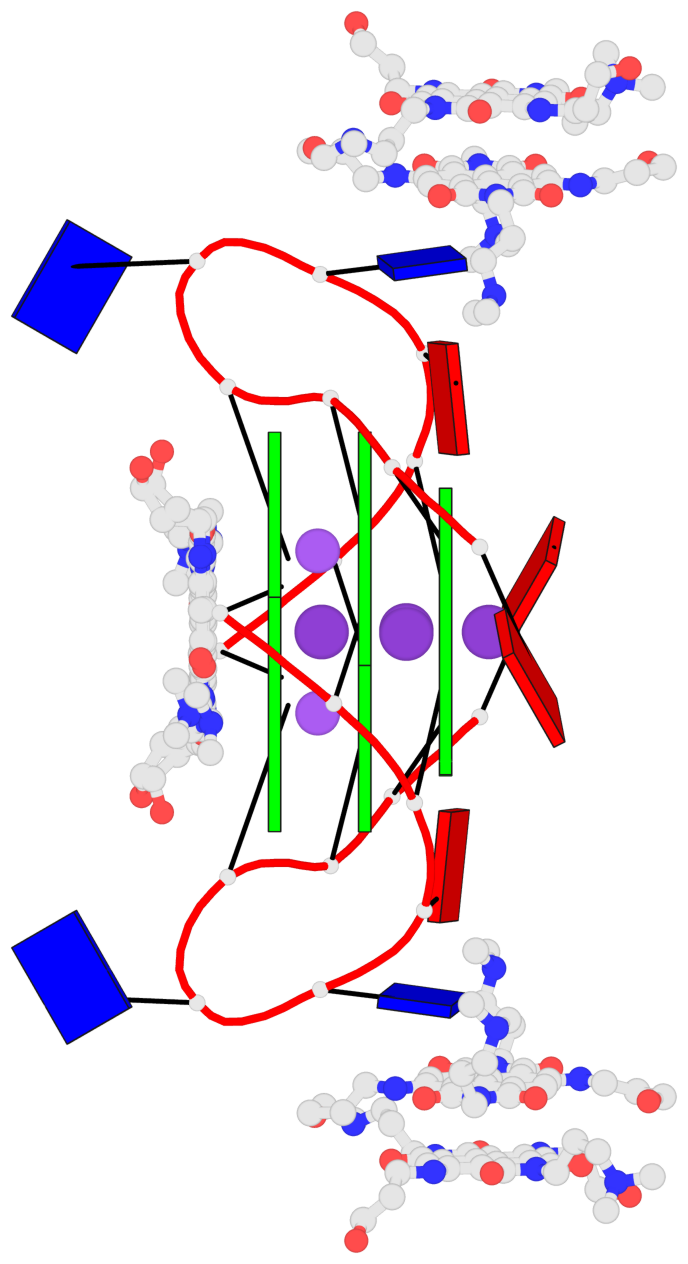
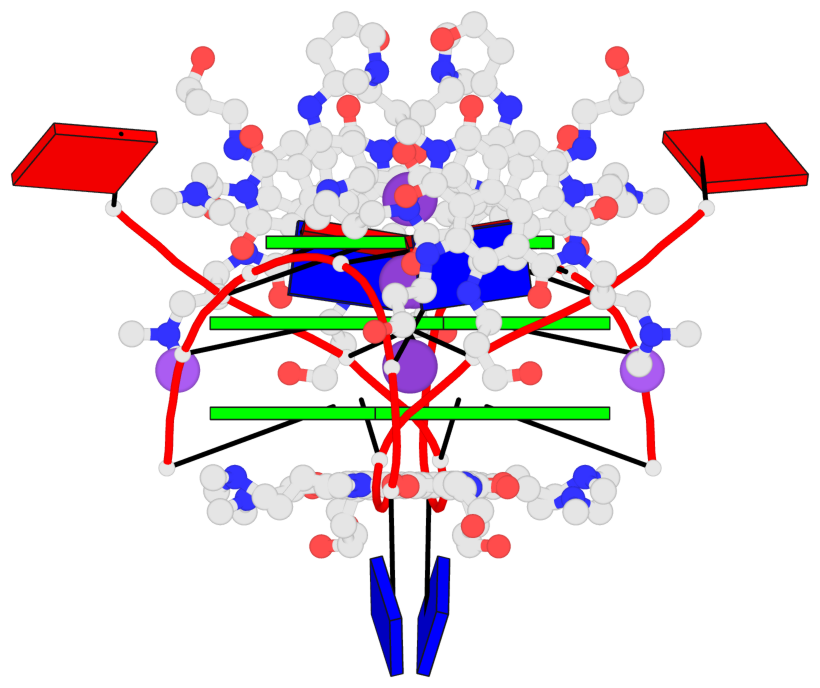
Base-block schematics in six views
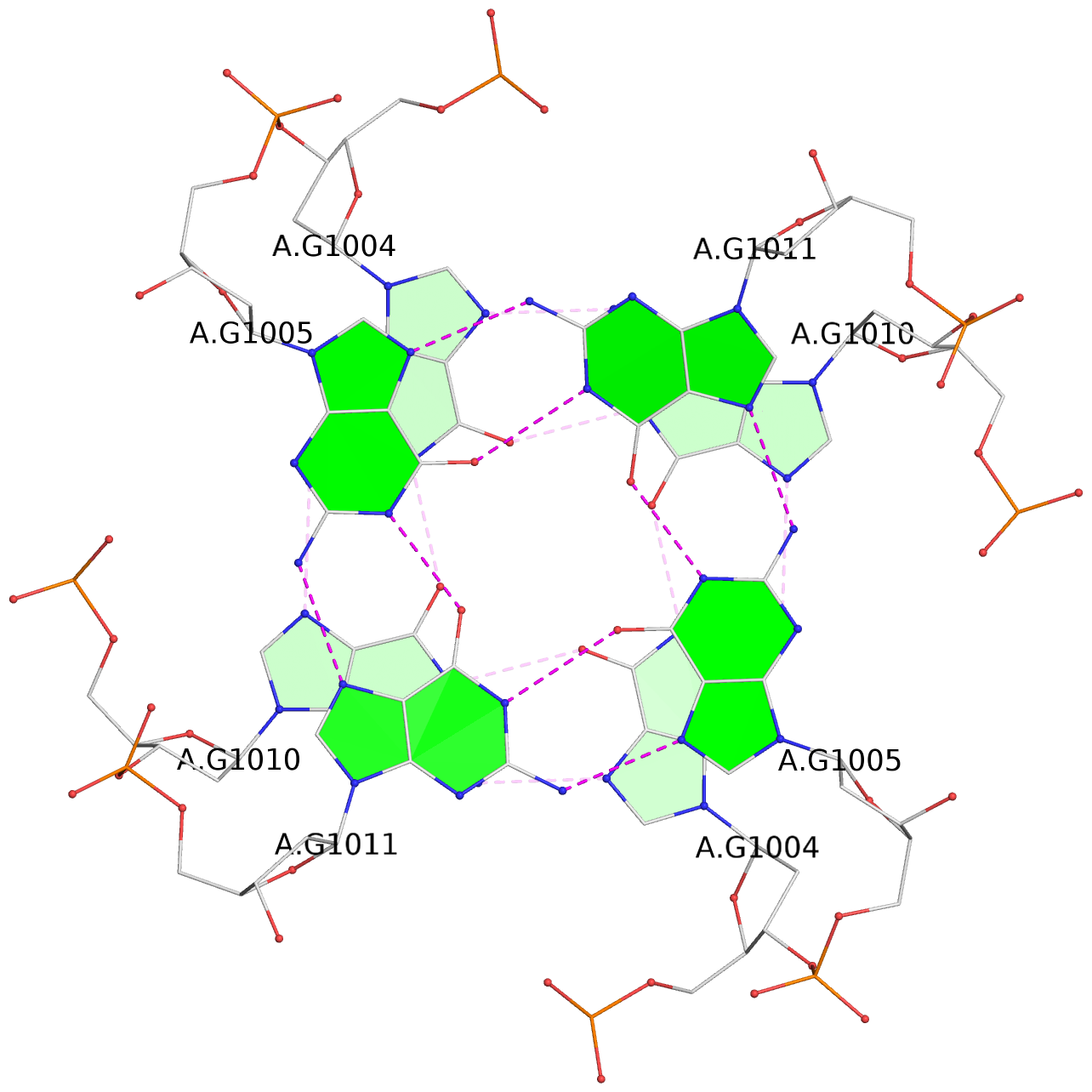
List of 3 G-tetrads
1 glyco-bond=---- sugar=-3-3 groove=---- planarity=0.409 type=saddle nts=4 GGGG 1:A.DG1003,1:A.DG1009,2:A.DG1003,2:A.DG1009 2 glyco-bond=---- sugar=3333 groove=---- planarity=0.192 type=saddle nts=4 GGGG 1:A.DG1004,1:A.DG1010,2:A.DG1004,2:A.DG1010 3 glyco-bond=---- sugar=---- groove=---- planarity=0.387 type=bowl nts=4 GGGG 1:A.DG1005,1:A.DG1011,2:A.DG1005,2:A.DG1011
List of 1 G4-helix
In DSSR, a G4-helix is defined by stacking interactions of G-tetrads, regardless of backbone connectivity, and may contain more than one G4-stem.
Helix#1, 3 G-tetrad layers, inter-molecular, with 1 stem
List of 1 G4-stem
In DSSR, a G4-stem is defined as a G4-helix with backbone connectivity. Bulges are also allowed along each of the four strands.