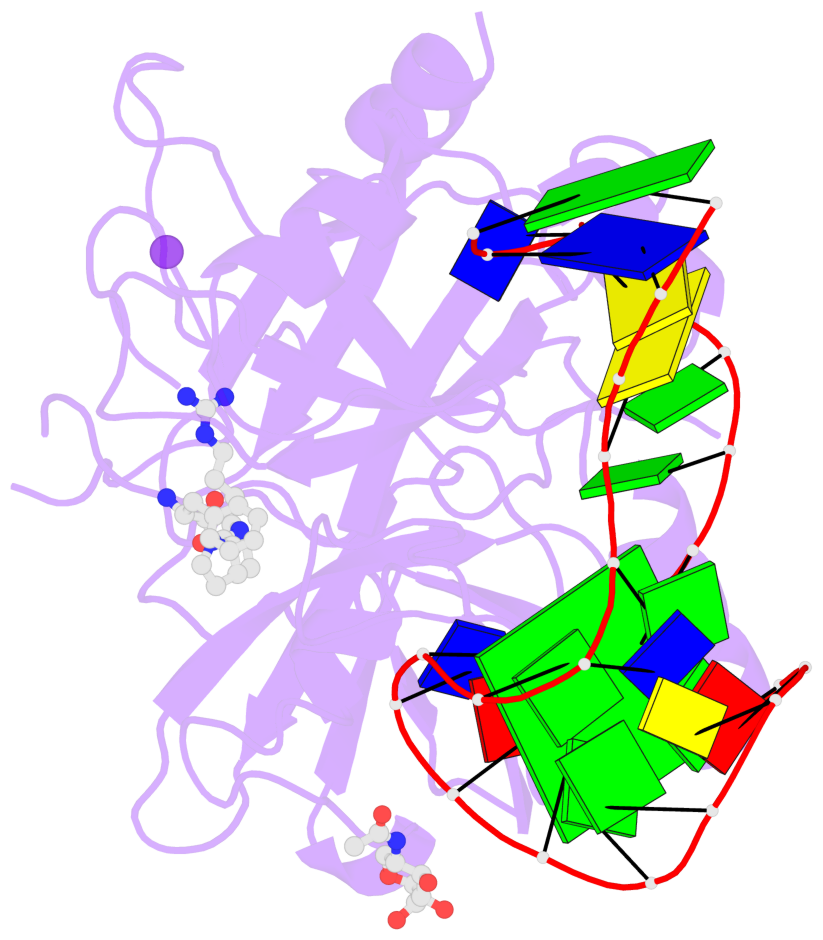
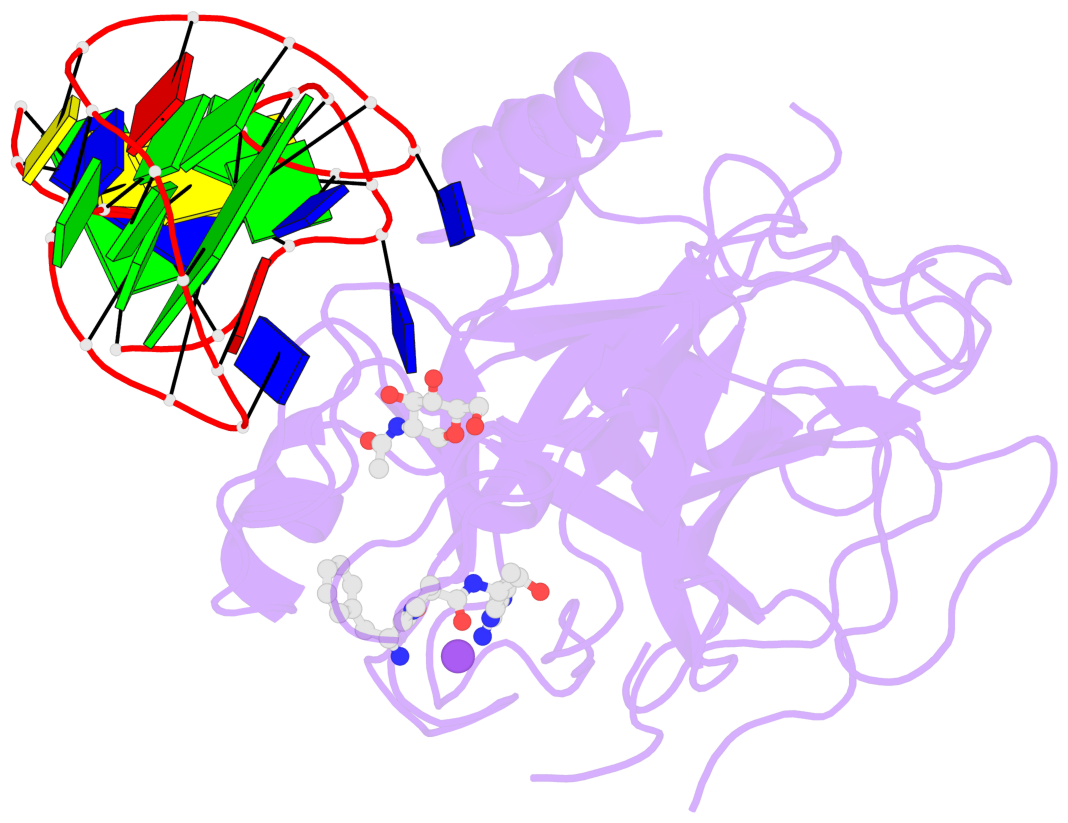
Detailed DSSR results for the G-quadruplex: PDB entry 4i7y
Created and maintained by Xiang-Jun Lu <xiangjun@x3dna.org>
Citation: Please cite the NAR'20 DSSR-PyMOL schematics paper and/or the NAR'15 DSSR method paper.
Summary information
- PDB id
- 4i7y
- Class
- hydrolase-hydrolase inhibitor-DNA
- Method
- X-ray (2.4 Å)
- Summary
- Crystal structure of human alpha thrombin in complex with a 27-mer aptamer bound to exosite ii
- Reference
- Russo Krauss I, Pica A, Merlino A, Mazzarella L, Sica F (2013): "Duplex-quadruplex motifs in a peculiar structural organization cooperatively contribute to thrombin binding of a DNA aptamer." Acta Crystallogr.,Sect.D, 69, 2403-2411. doi: 10.1107/S0907444913022269.
- Abstract
- Potent second-generation thrombin aptamers adopt a duplex-quadruplex bimodular folding and recognize thrombin exosite II with very high affinity and specificity. A sound model of these oligonucleotides, either free or in complex with thrombin, is not yet available. Here, a structural study of one of these aptamers, HD22-27mer, is presented. The crystal structure of this aptamer in complex with thrombin displays a novel architecture in which the helical stem is enchained to a pseudo-G-quadruplex. The results also underline the role of the residues that join the duplex and quadruplex motifs and control their recruitment in thrombin binding.
- G4 notes
- 1 G-tetrad
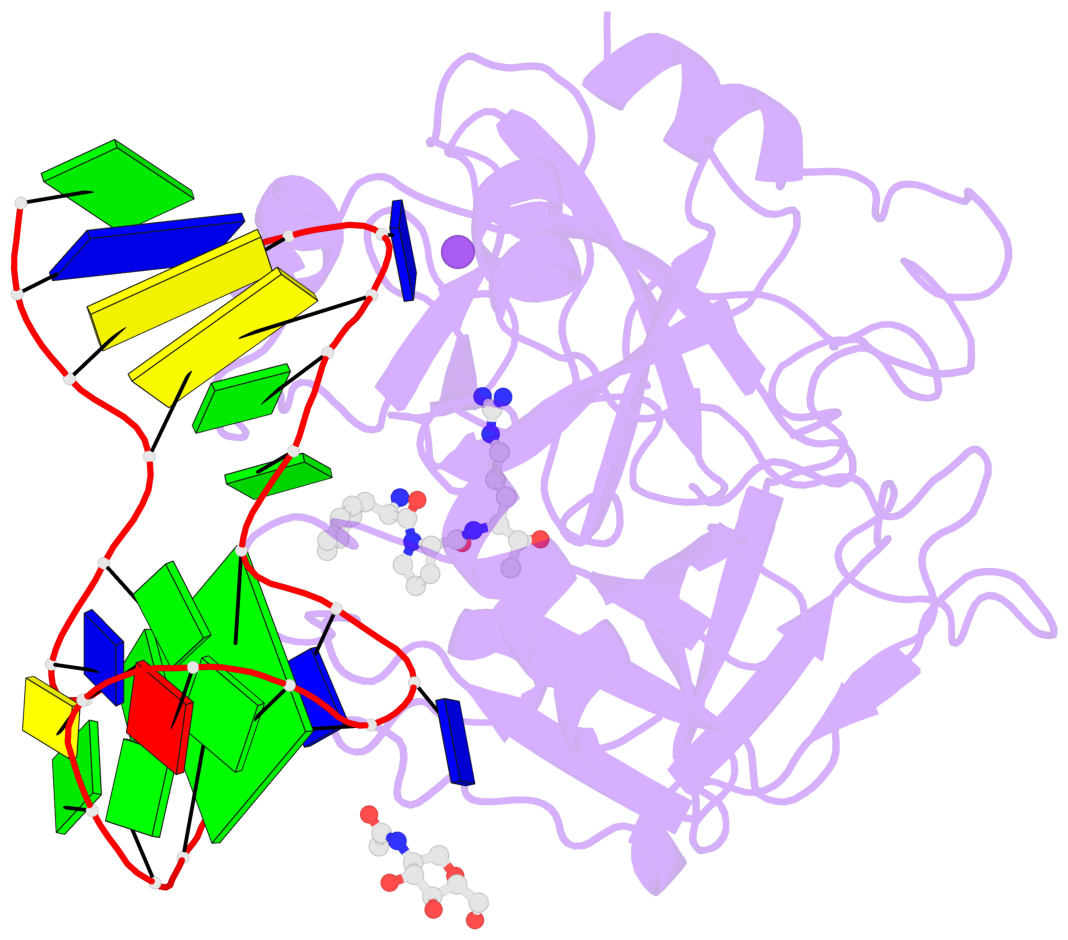
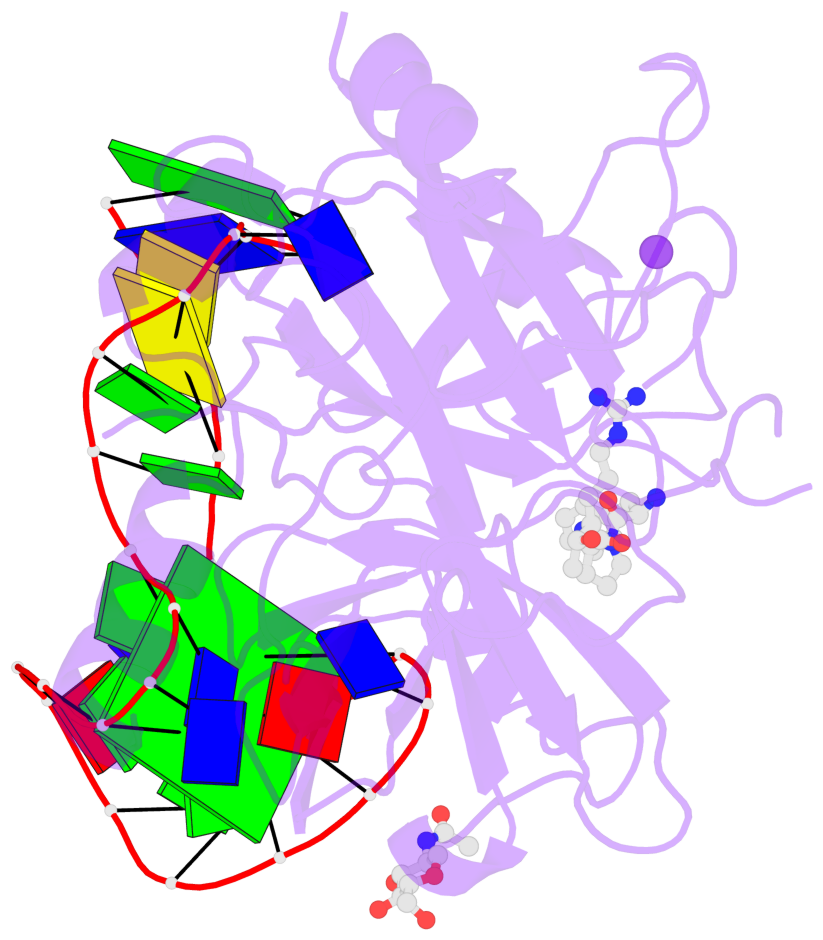
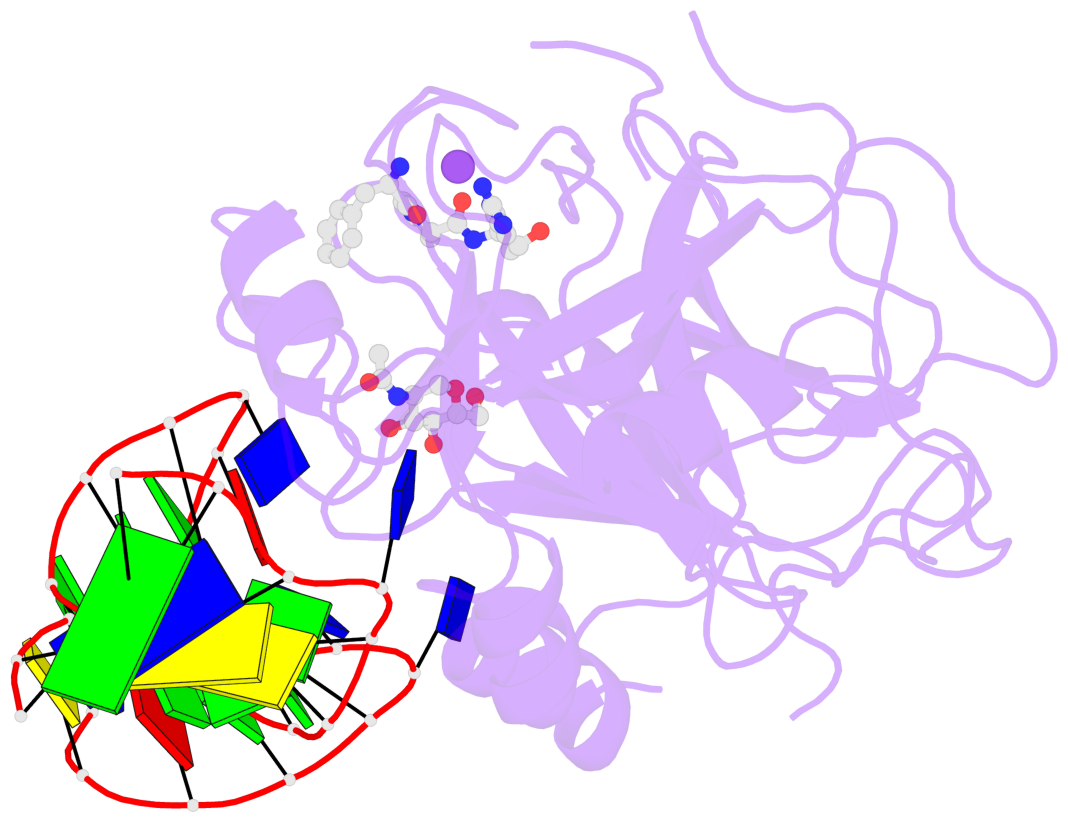
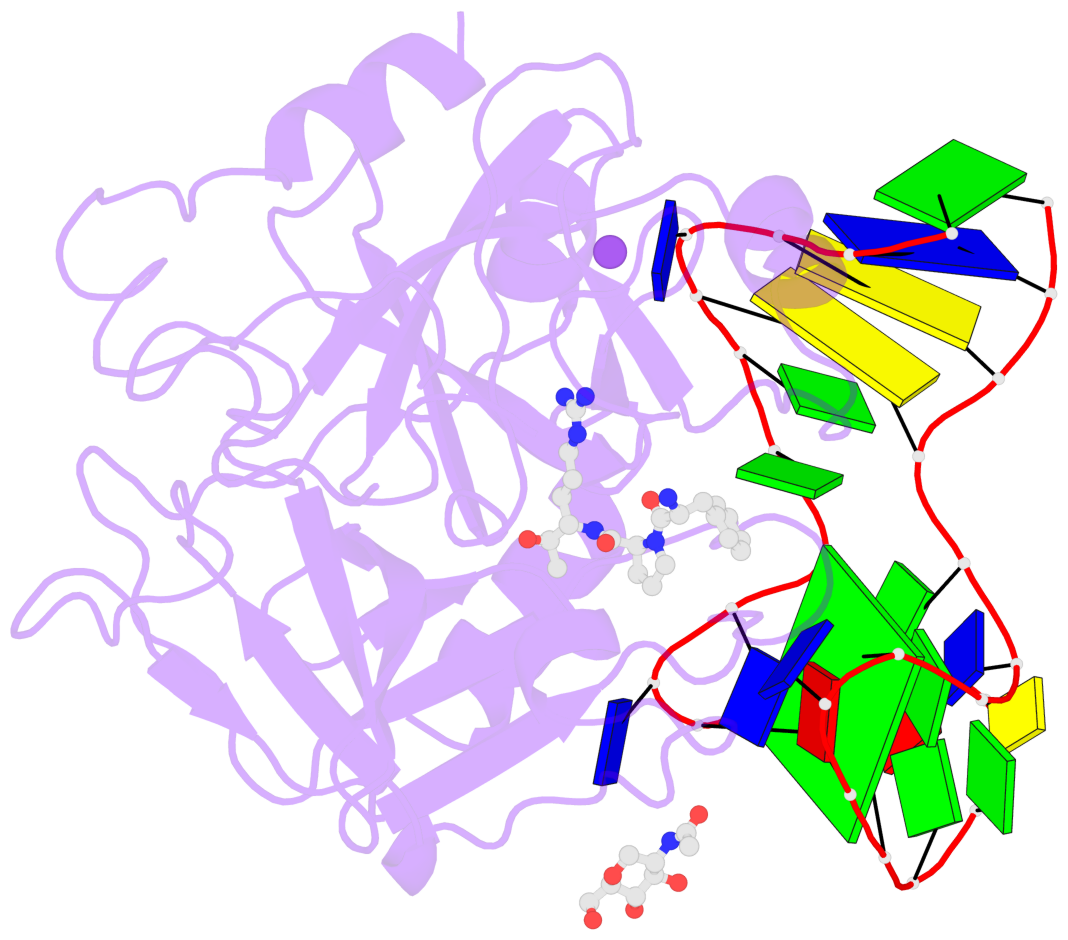
Base-block schematics in six views
List of 1 G-tetrad
1 glyco-bond=-s-s sugar=-3.- groove=wnwn planarity=0.842 type=other nts=4 GGGG D.DG8,D.DG20,D.DG17,D.DG11
List of 3 non-stem G4-loops (including the two closing Gs)
1 type=lateral helix=#-1 nts=4 GTAG D.DG8,D.DT9,D.DA10,D.DG11 2 type=lateral helix=#-1 nts=7 GGGCAGG D.DG11,D.DG12,D.DG13,D.DC14,D.DA15,D.DG16,D.DG17 3 type=lateral helix=#-1 nts=4 GTTG D.DG17,D.DT18,D.DT19,D.DG20