Detailed DSSR results for the G-quadruplex: PDB entry 4lz1
Created and maintained by Xiang-Jun Lu <xiangjun@x3dna.org>
Citation: Please cite the NAR'20 DSSR-PyMOL schematics paper and/or the NAR'15 DSSR method paper.
Summary information
- PDB id
- 4lz1
- Class
- hydrolase-hydrolase inhibitor-DNA
- Method
- X-ray (1.65 Å)
- Summary
- X-ray structure of the complex between human thrombin and the tba deletion mutant lacking thymine 12 nucleobase
- Reference
- Pica A, Russo Krauss I, Merlino A, Nagatoishi S, Sugimoto N, Sica F (2013): "Dissecting the contribution of thrombin exosite I in the recognition of thrombin binding aptamer." Febs J., 280, 6581-6588. doi: 10.1111/febs.12561.
- Abstract
- Thrombin plays a pivotal role in the coagulation cascade; therefore, it represents a primary target in the treatment of several blood diseases. The 15-mer DNA oligonucleotide 5'-GGTTGGTGTGGTTGG-3', known as thrombin binding aptamer (TBA), is a highly potent inhibitor of the enzyme. TBA folds as an antiparallel chair-like G-quadruplex structure, with two G-tetrads surrounded by two TT loops on one side and a TGT loop on the opposite side. Previous crystallographic studies have shown that TBA binds thrombin exosite I by its TT loops, T3T4 and T12T13. In order to get a better understanding of the thrombin-TBA interaction, we have undertaken a crystallographic characterization of the complexes between thrombin and two TBA mutants, TBAΔT3 and TBAΔT12, which lack the nucleobase of T3 and T12, respectively. The structural details of the two complexes show that exosite I is actually split into two regions, which contribute differently to TBA recognition. These results provide the basis for a more rational design of new aptamers with improved therapeutic action.
- G4 notes
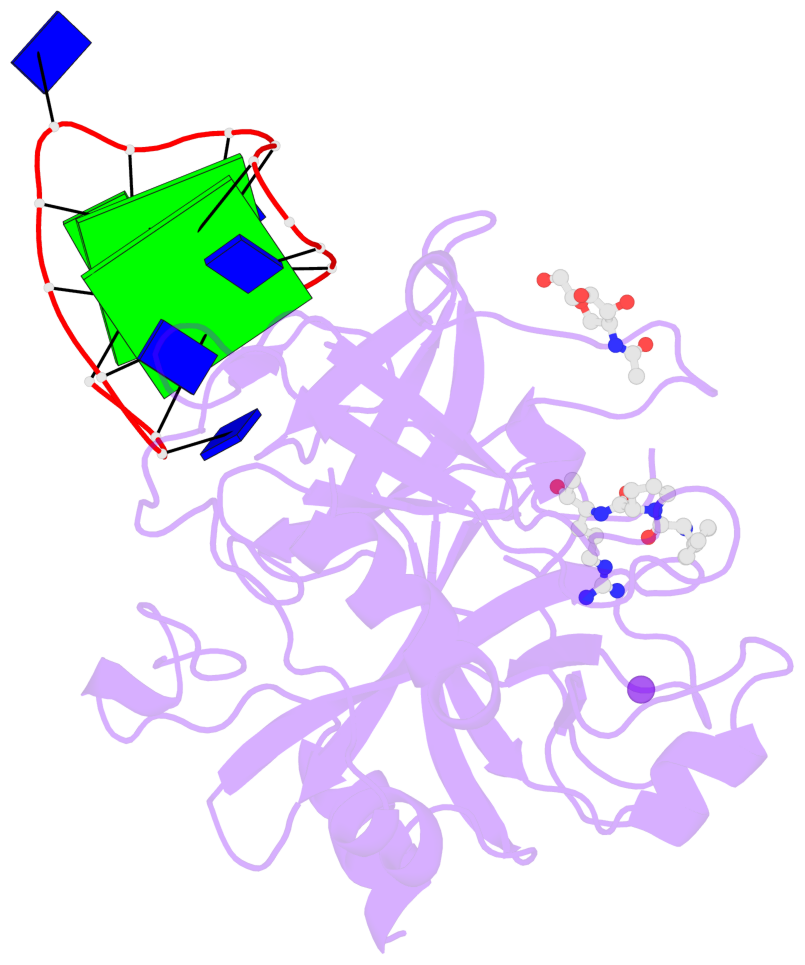
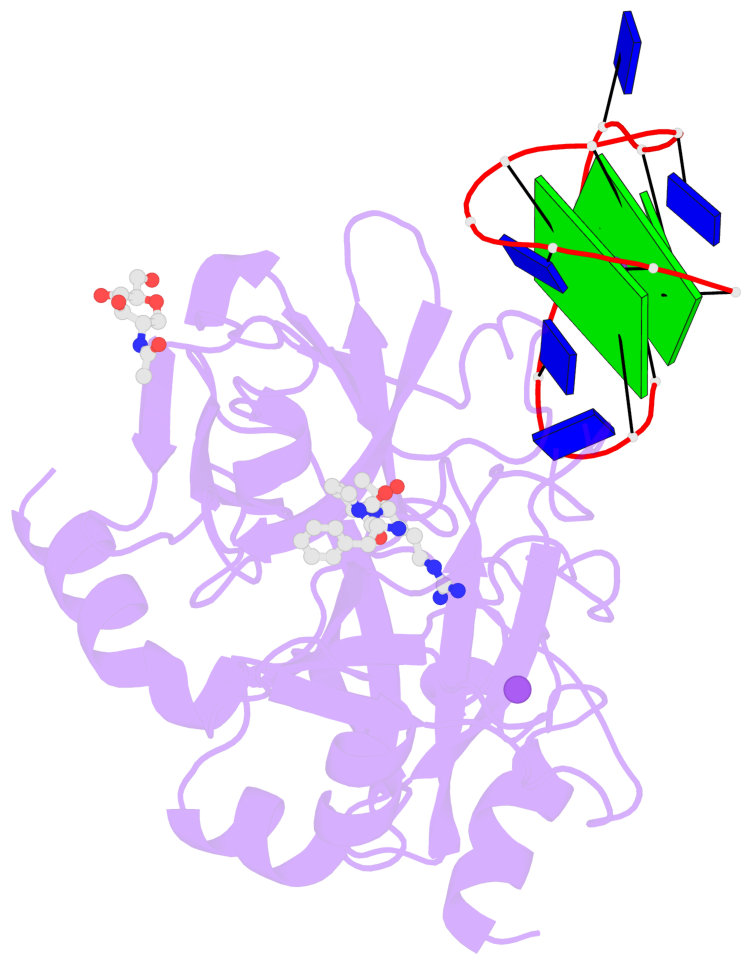
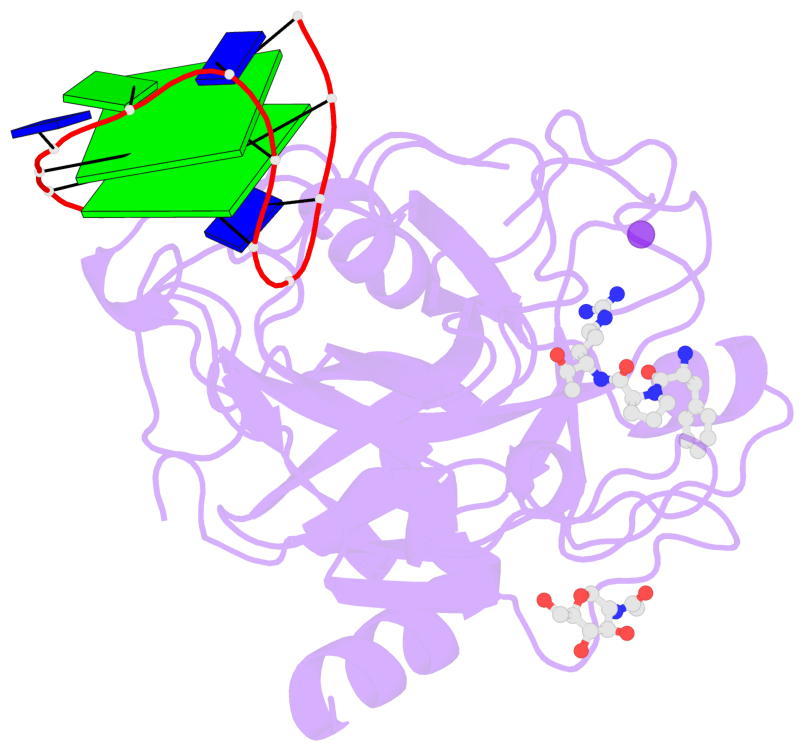
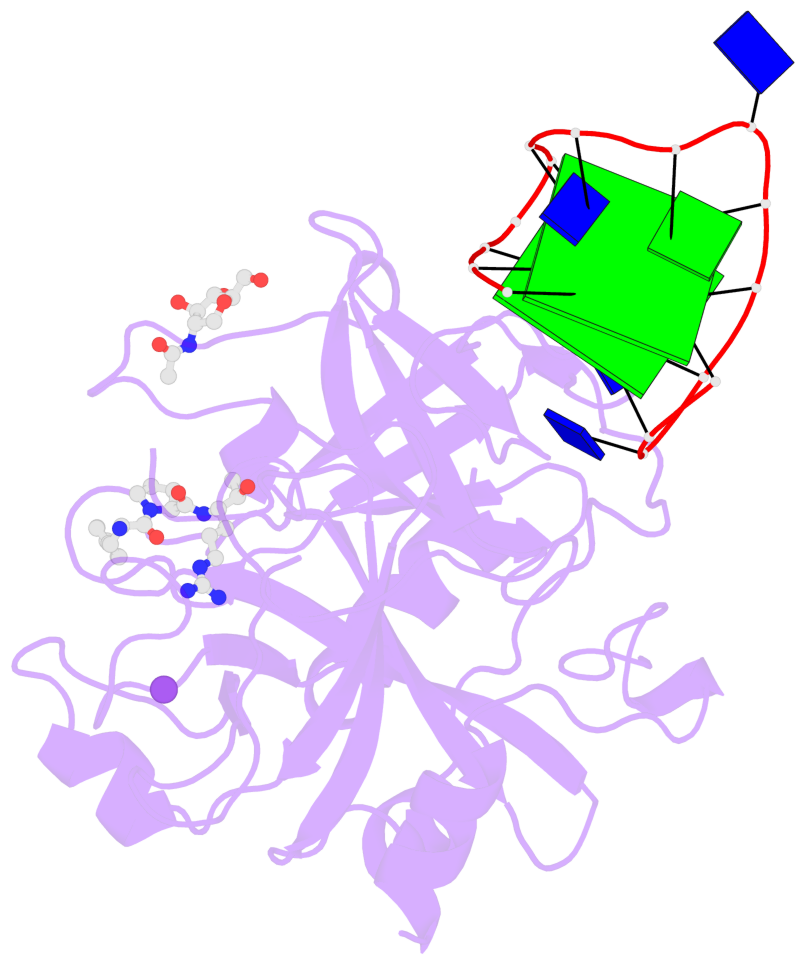
- 2 G-tetrads, 1 G4 helix, 1 G4 stem, 2(+Ln+Lw+Ln), chair(2+2), UDUD
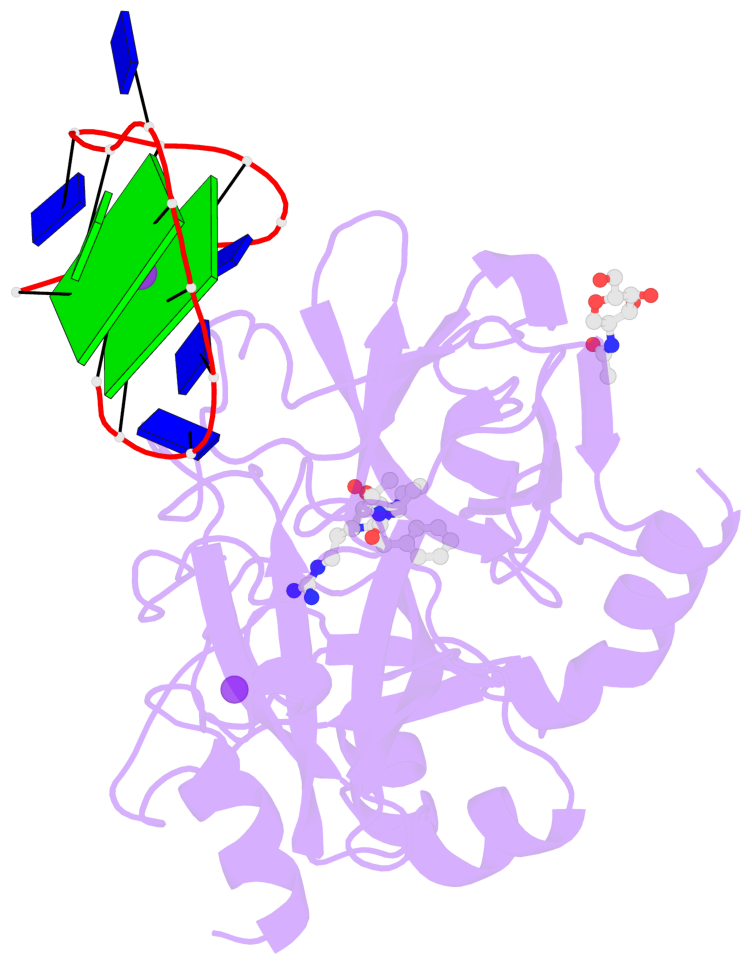
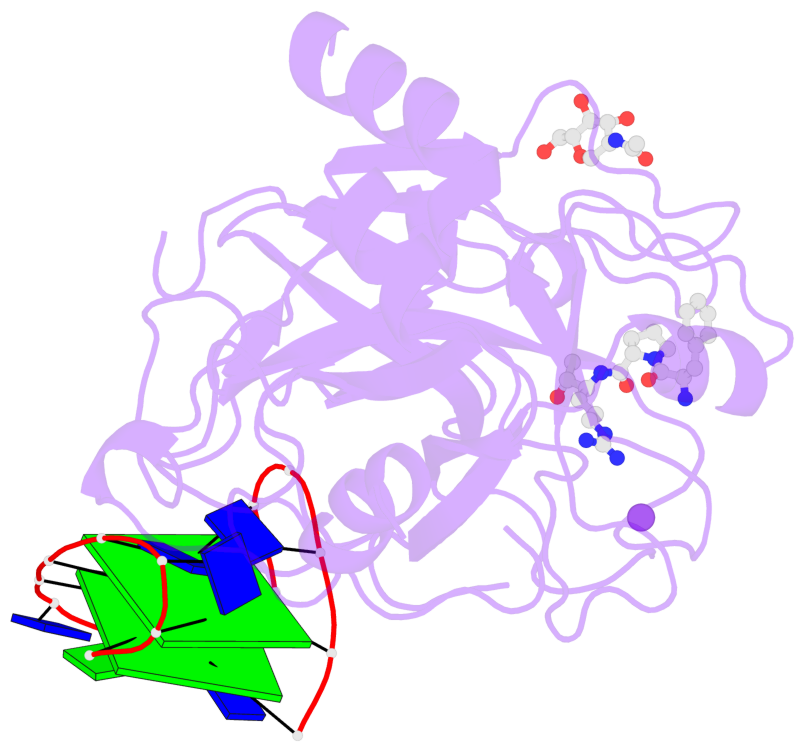
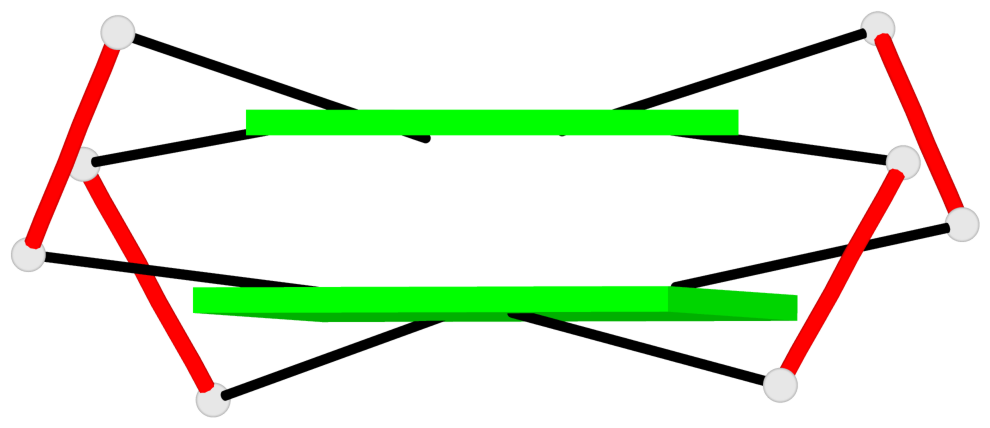
Base-block schematics in six views
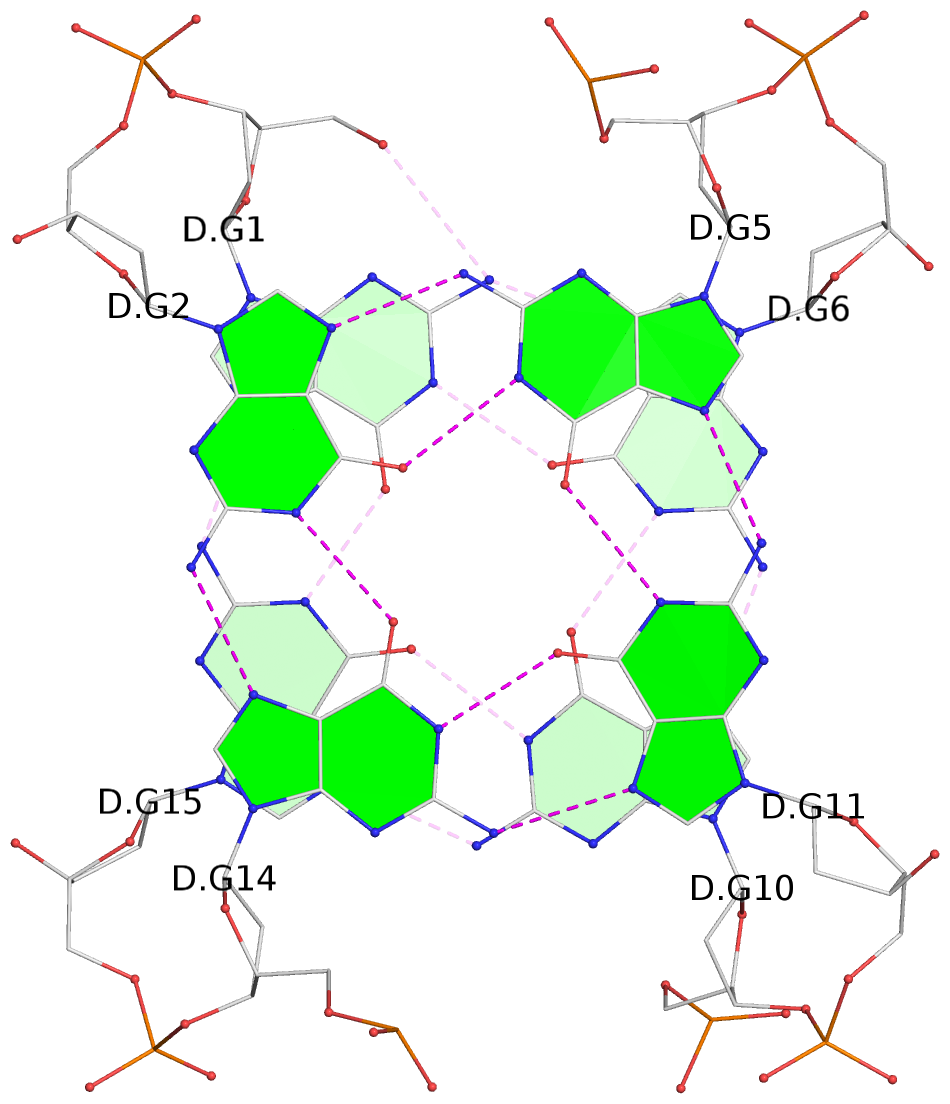
List of 2 G-tetrads
1 glyco-bond=s-s- sugar=---- groove=wnwn planarity=0.456 type=saddle nts=4 GGGG D.DG1,D.DG15,D.DG10,D.DG6 2 glyco-bond=-s-s sugar=---- groove=wnwn planarity=0.506 type=other nts=4 GGGG D.DG2,D.DG14,D.DG11,D.DG5
List of 1 G4-helix
In DSSR, a G4-helix is defined by stacking interactions of G-tetrads, regardless of backbone connectivity, and may contain more than one G4-stem.
Helix#1, 2 G-tetrad layers, INTRA-molecular, with 1 stem
List of 1 G4-stem
In DSSR, a G4-stem is defined as a G4-helix with backbone connectivity. Bulges are also allowed along each of the four strands.