Detailed DSSR results for the G-quadruplex: PDB entry 4rne
Created and maintained by Xiang-Jun Lu <xiangjun@x3dna.org>
Citation: Please cite the NAR'20 DSSR-PyMOL schematics paper and/or the NAR'15 DSSR method paper.
Summary information
- PDB id
- 4rne
- Class
- RNA
- Method
- X-ray (1.01 Å)
- Summary
- Structural variations and solvent structure of uggggu quadruplexes stabilized by sr2+ ions
- Reference
- Fyfe AC, Dunten PW, Martick MM, Scott WG (2015): "Structural Variations and Solvent Structure of r(UGGGGU) Quadruplexes Stabilized by Sr(2+) Ions." J.Mol.Biol., 427, 2205-2219. doi: 10.1016/j.jmb.2015.03.022.
- Abstract
- Guanine-rich sequences can, under appropriate conditions, adopt a distinctive, four-stranded, helical fold known as a G-quadruplex. Interest in quadruplex folds has grown in recent years as evidence of their biological relevance has accumulated from both sequence analysis and function-specific assays. The folds are unusually stable and their formation appears to require close management to maintain cell health; regulatory failure correlates with genomic instability and a number of cancer phenotypes. Biologically relevant quadruplex folds are anticipated to form transiently in mRNA and in single-stranded, unwound DNA. To elucidate factors, including bound solvent, that contribute to the stability of RNA quadruplexes, we examine, by X-ray crystallography and small-angle X-ray scattering, the structure of a previously reported tetramolecular quadruplex, UGGGGU stabilized by Sr(2+) ions. Crystal forms of the octameric assembly formed by this sequence exhibit unusually strong diffraction and anomalous signal enabling the construction of reliable models to a resolution of 0.88Å. The solvent structure confirms hydration patterns reported for other nucleic acid helical conformations and provides support for the greater stability of RNA quadruplexes relative to DNA. Novel features detected in the octameric RNA assembly include a new crystal form, evidence of multiple conformations and structural variations in the 3' U tetrad, including one that leads to the formation of a hydrated internal cavity.
- G4 notes
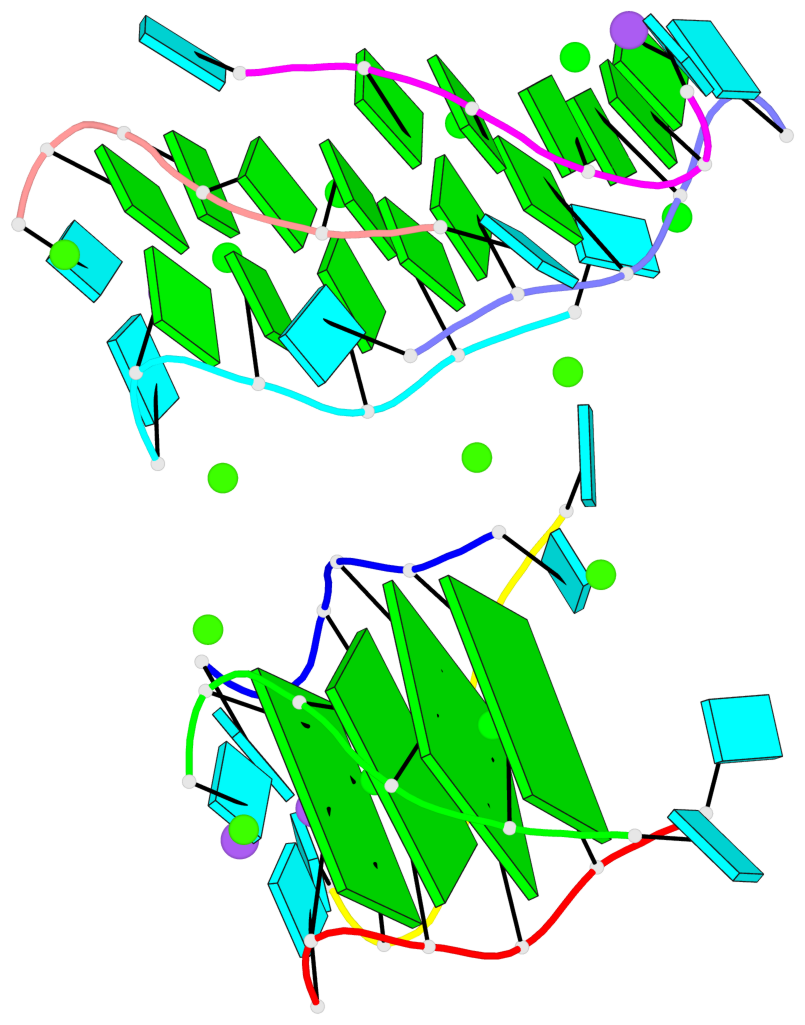
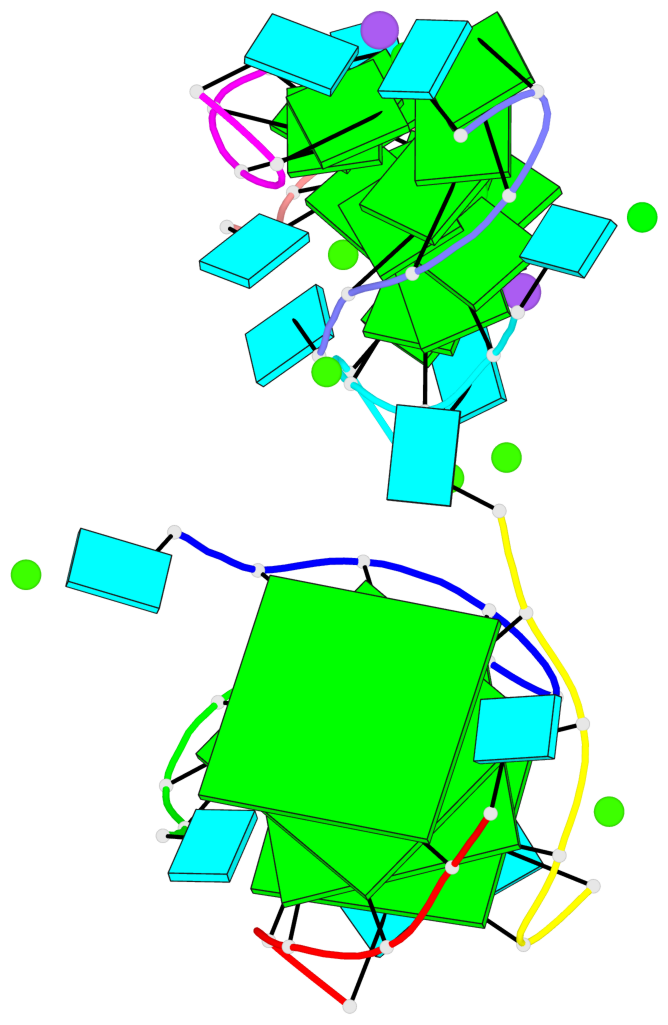
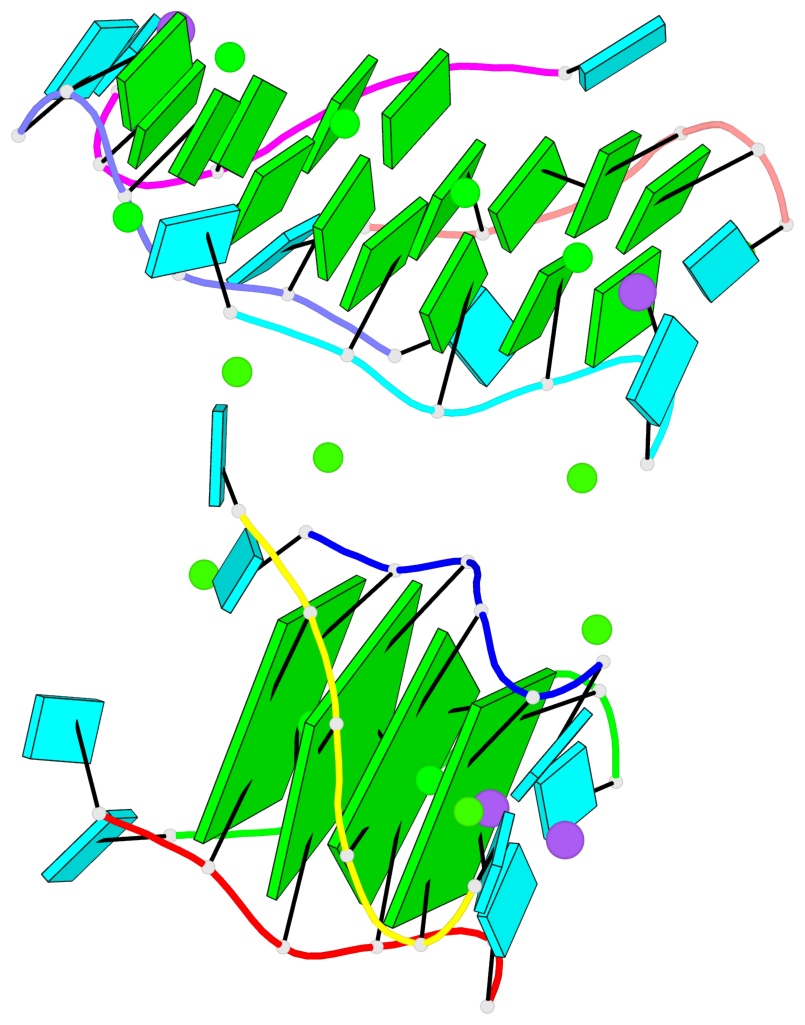
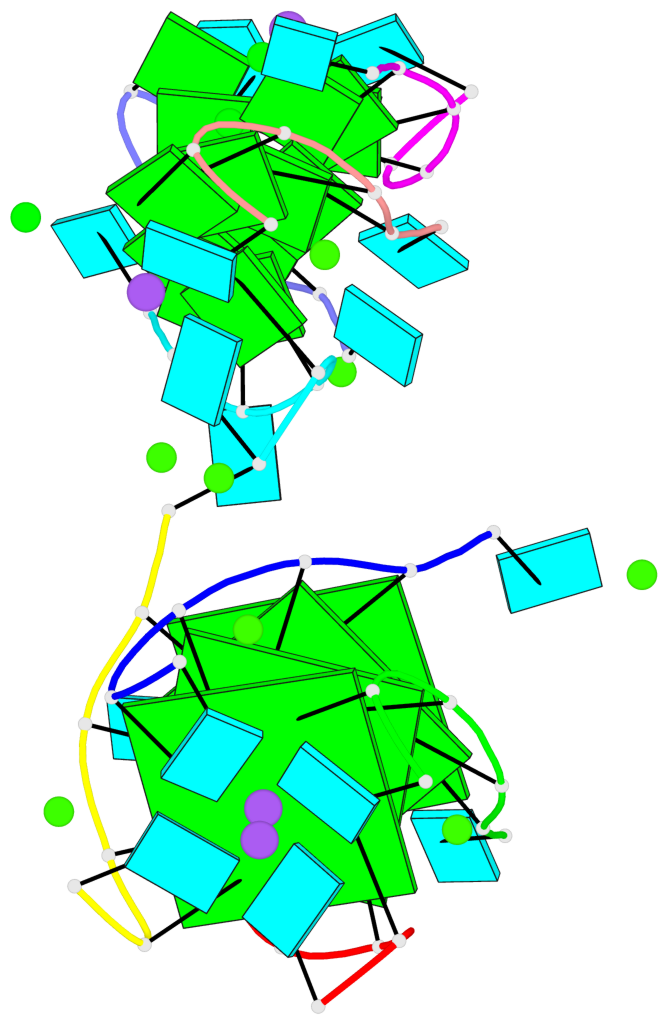
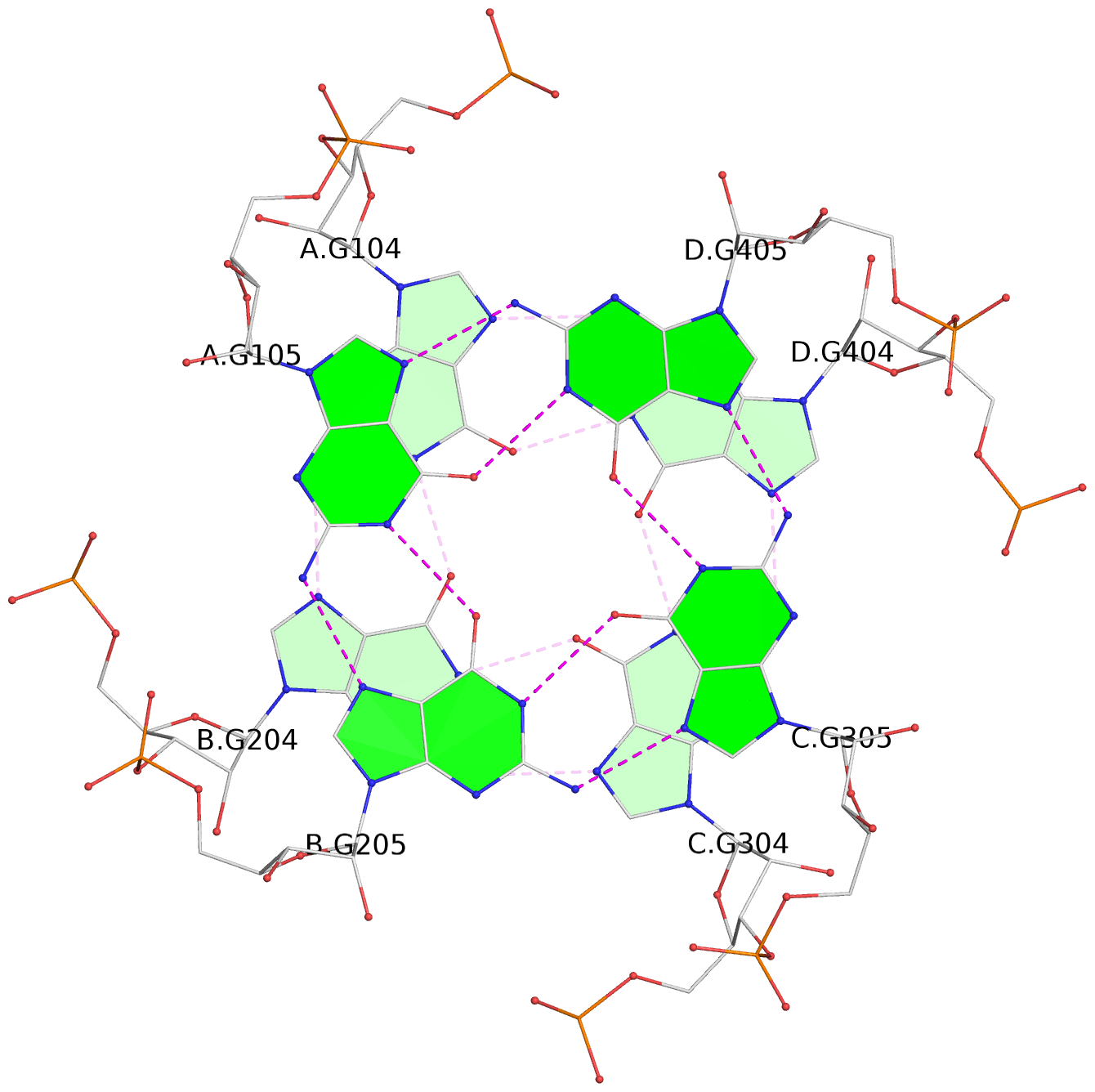
- 4 G-tetrads, 1 G4 helix, 1 G4 stem, parallel(4+0), UUUU
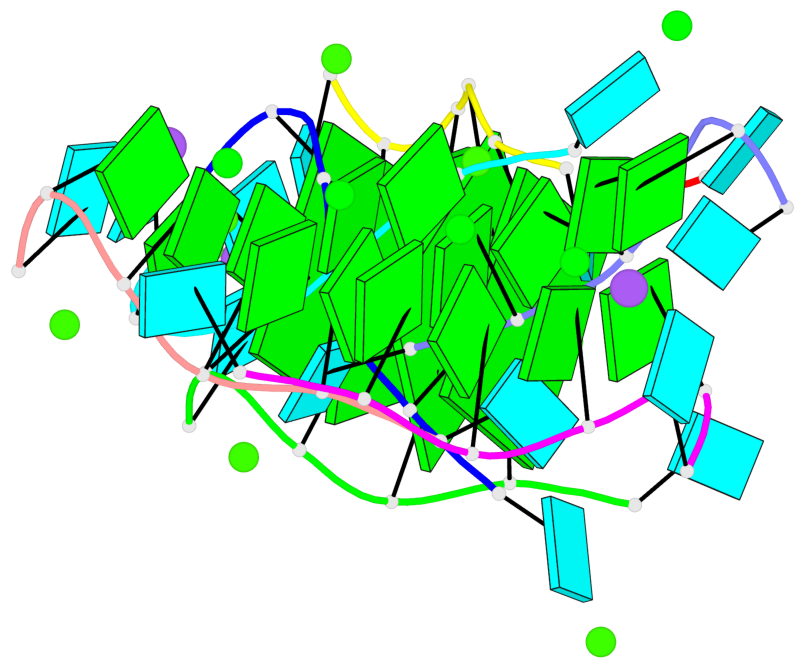
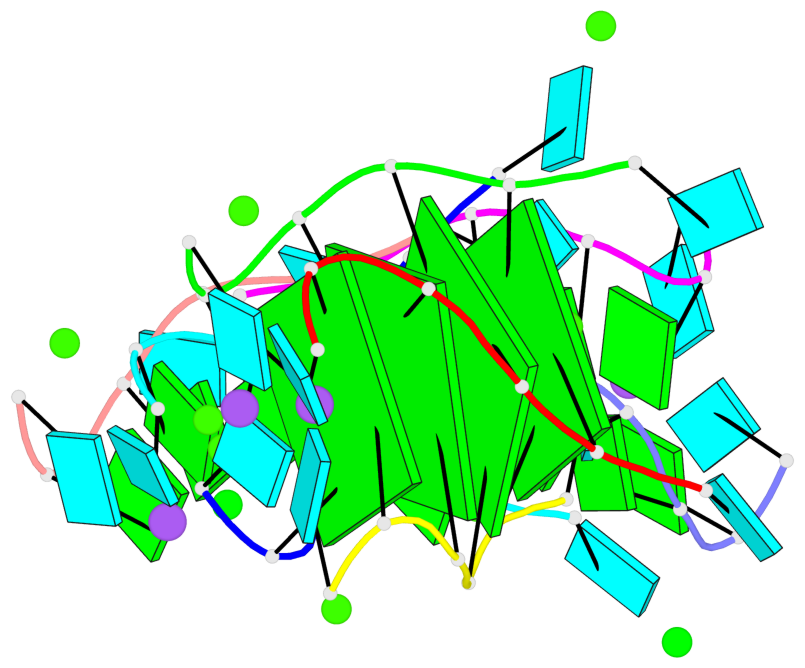
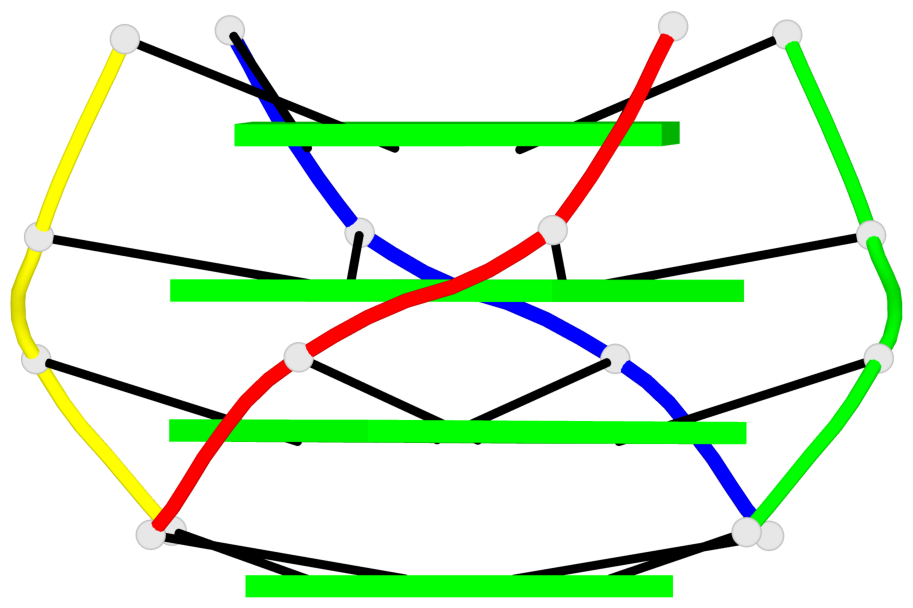
Base-block schematics in six views
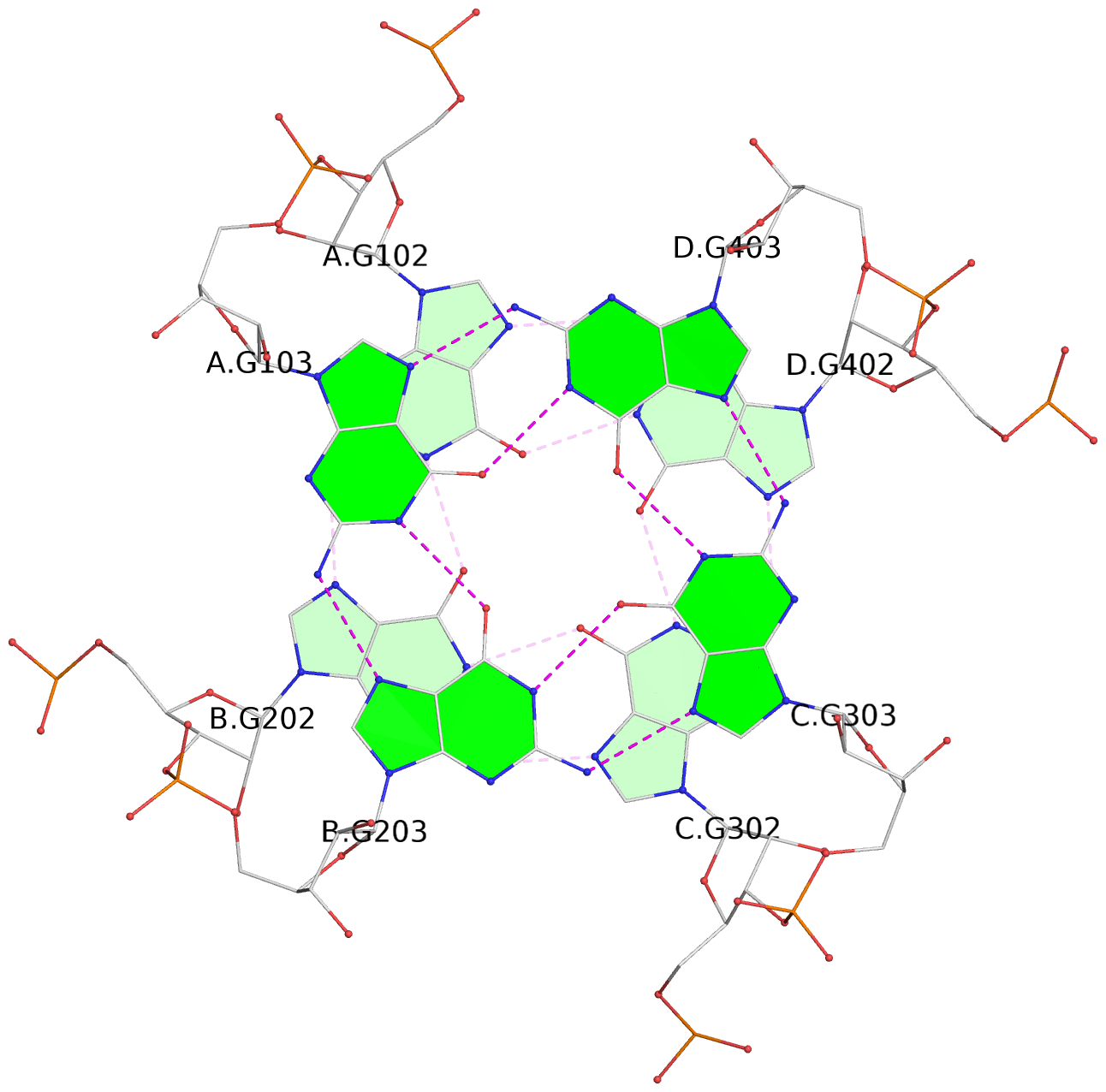
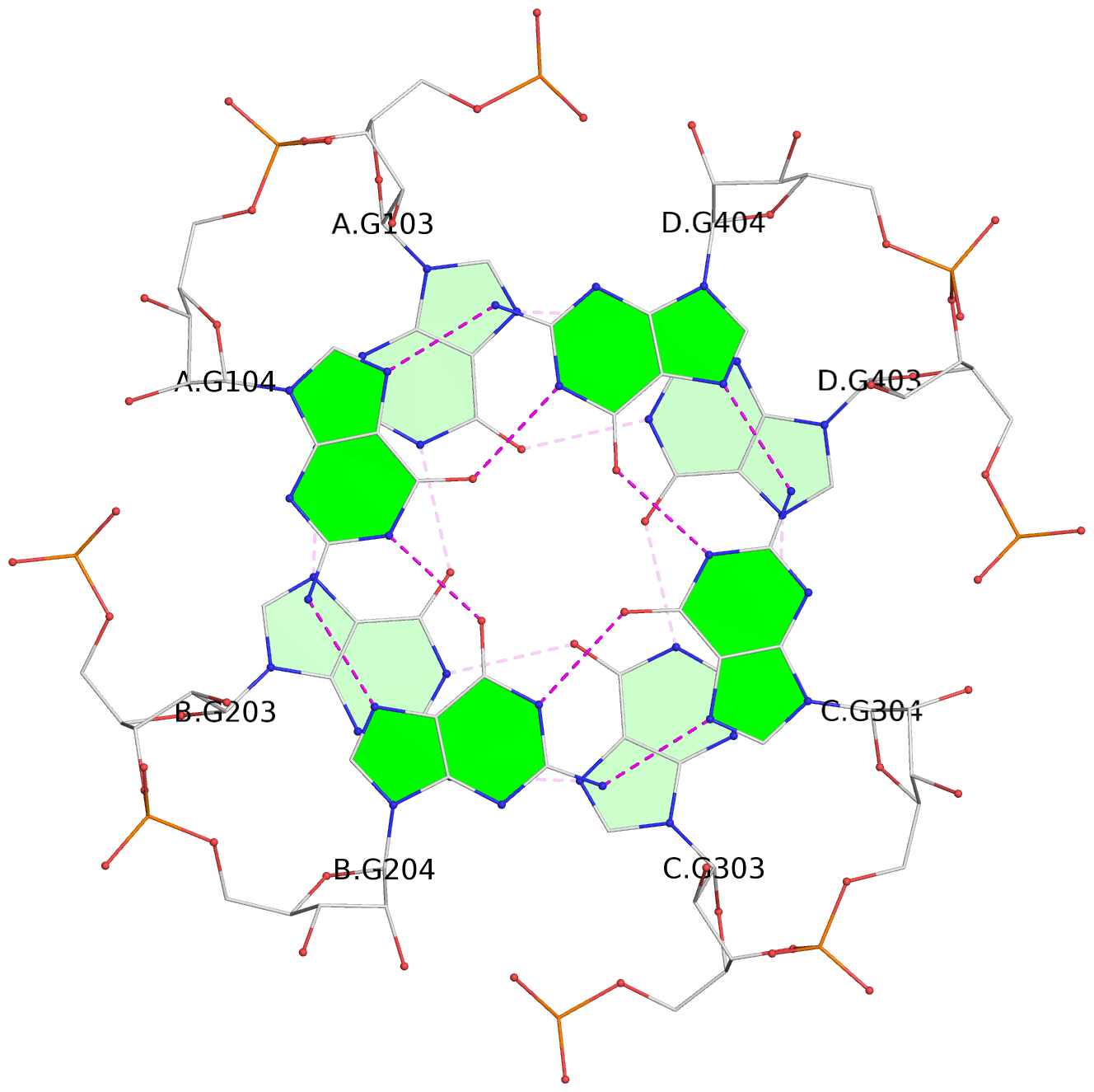
List of 4 G-tetrads
1 glyco-bond=---- sugar=3333 groove=---- planarity=0.231 type=other nts=4 GGGG A.G102,B.G202,C.G302,D.G402 2 glyco-bond=---- sugar=---- groove=---- planarity=0.231 type=other nts=4 GGGG A.G103,B.G203,C.G303,D.G403 3 glyco-bond=---- sugar=3333 groove=---- planarity=0.121 type=planar nts=4 GGGG A.G104,B.G204,C.G304,D.G404 4 glyco-bond=---- sugar=3333 groove=---- planarity=0.283 type=bowl nts=4 GGGG A.G105,B.G205,C.G305,D.G405
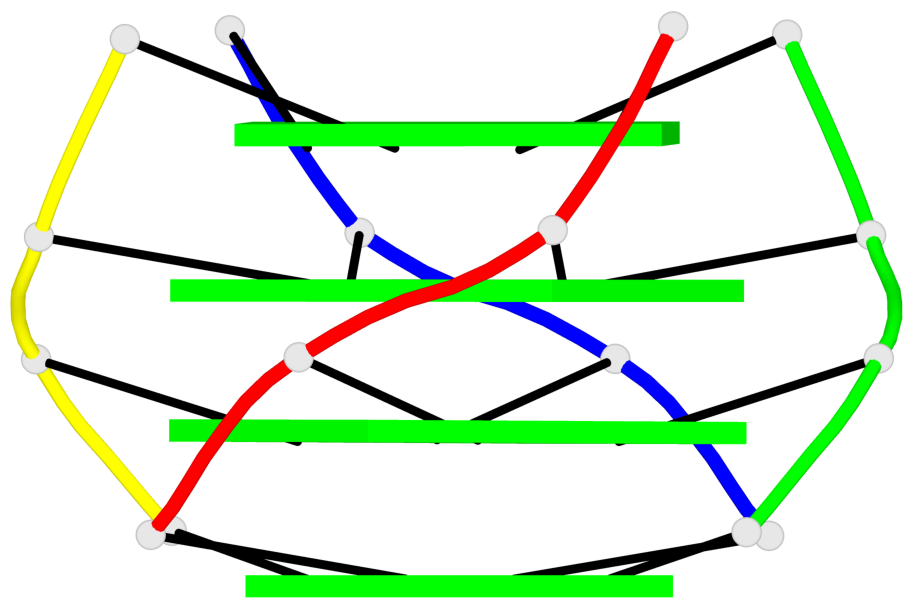
List of 1 G4-helix
In DSSR, a G4-helix is defined by stacking interactions of G-tetrads, regardless of backbone connectivity, and may contain more than one G4-stem.
Helix#1, 4 G-tetrad layers, inter-molecular, with 1 stem
List of 1 G4-stem
In DSSR, a G4-stem is defined as a G4-helix with backbone connectivity. Bulges are also allowed along each of the four strands.