Detailed DSSR results for the G-quadruplex: PDB entry 5ew2
Created and maintained by Xiang-Jun Lu <xiangjun@x3dna.org>
Citation: Please cite the NAR'20 DSSR-PyMOL schematics paper and/or the NAR'15 DSSR method paper.
Summary information
- PDB id
- 5ew2
- Class
- protein-DNA
- Method
- X-ray (3.59 Å)
- Summary
- Human thrombin sandwiched between two DNA aptamers: hd22 and hd1-deltat12
- Reference
- Pica A, Russo Krauss I, Parente V, Tateishi-Karimata H, Nagatoishi S, Tsumoto K, Sugimoto N, Sica F (2017): "Through-bond effects in the ternary complexes of thrombin sandwiched by two DNA aptamers." Nucleic Acids Res., 45, 461-469. doi: 10.1093/nar/gkw1113.
- Abstract
- Aptamers directed against human thrombin can selectively bind to two different exosites on the protein surface. The simultaneous use of two DNA aptamers, HD1 and HD22, directed to exosite I and exosite II respectively, is a very powerful approach to exploit their combined affinity. Indeed, strategies to link HD1 and HD22 together have been proposed in order to create a single bivalent molecule with an enhanced ability to control thrombin activity. In this work, the crystal structures of two ternary complexes, in which thrombin is sandwiched between two DNA aptamers, are presented and discussed. The structures shed light on the cross talk between the two exosites. The through-bond effects are particularly evident at exosite II, with net consequences on the HD22 structure. Moreover, thermodynamic data on the binding of the two aptamers are also reported and analyzed.
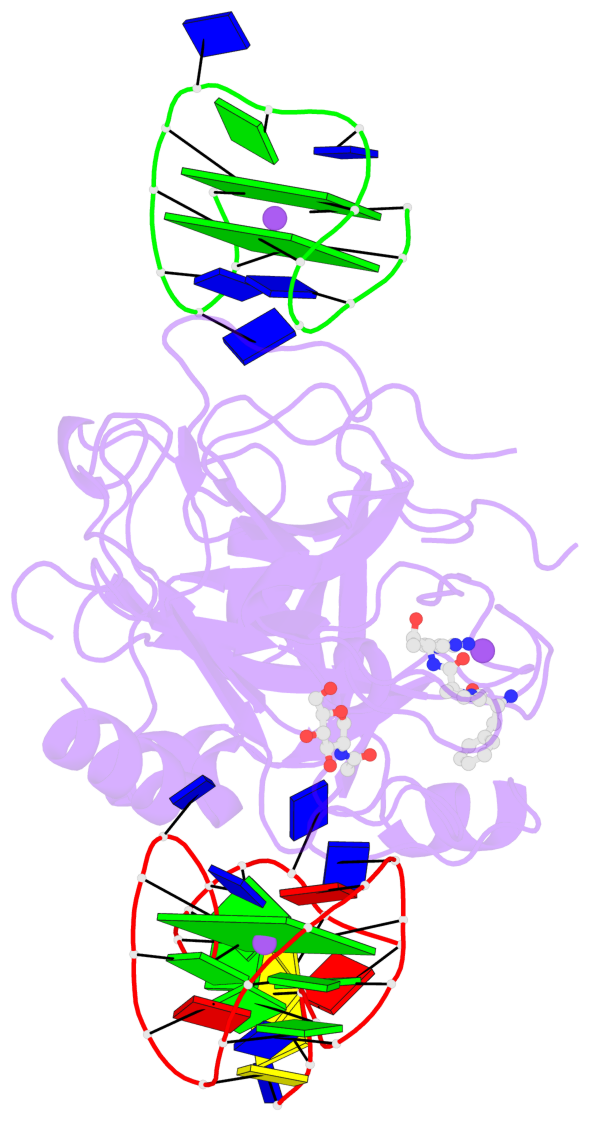
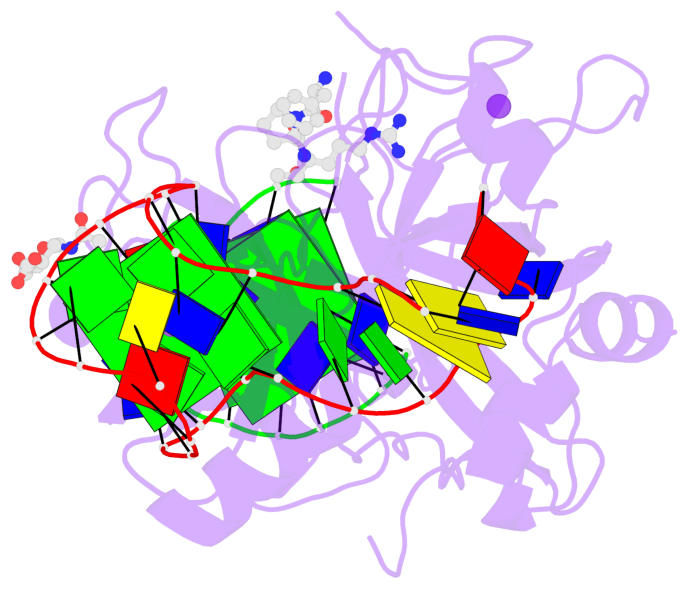
- G4 notes
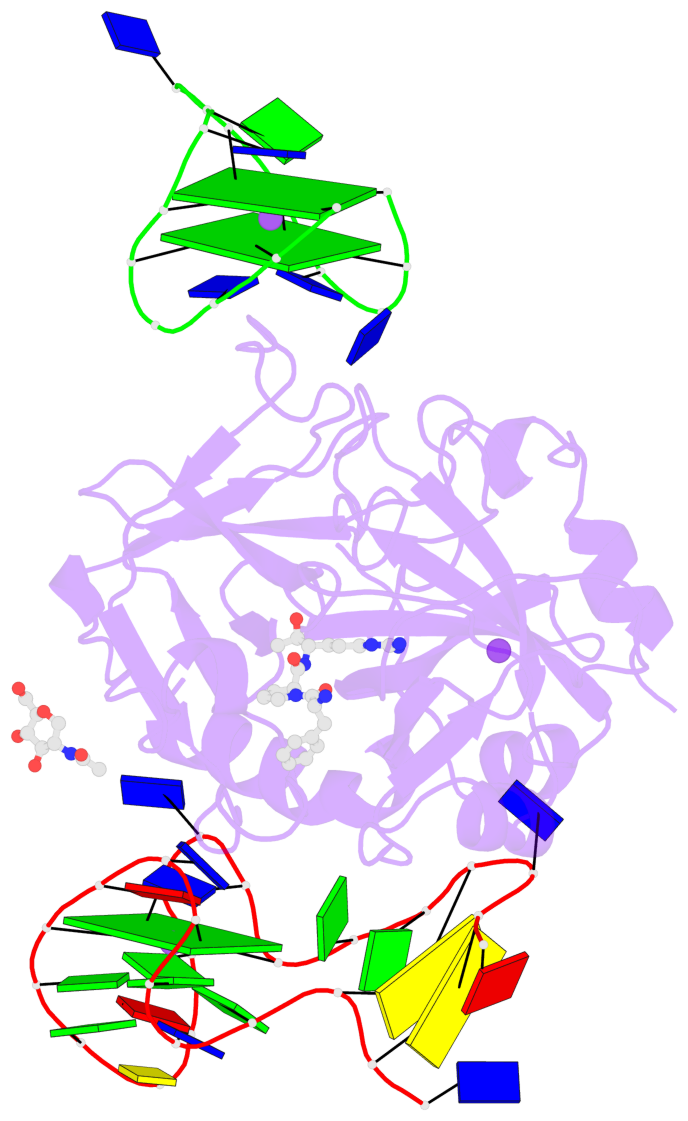
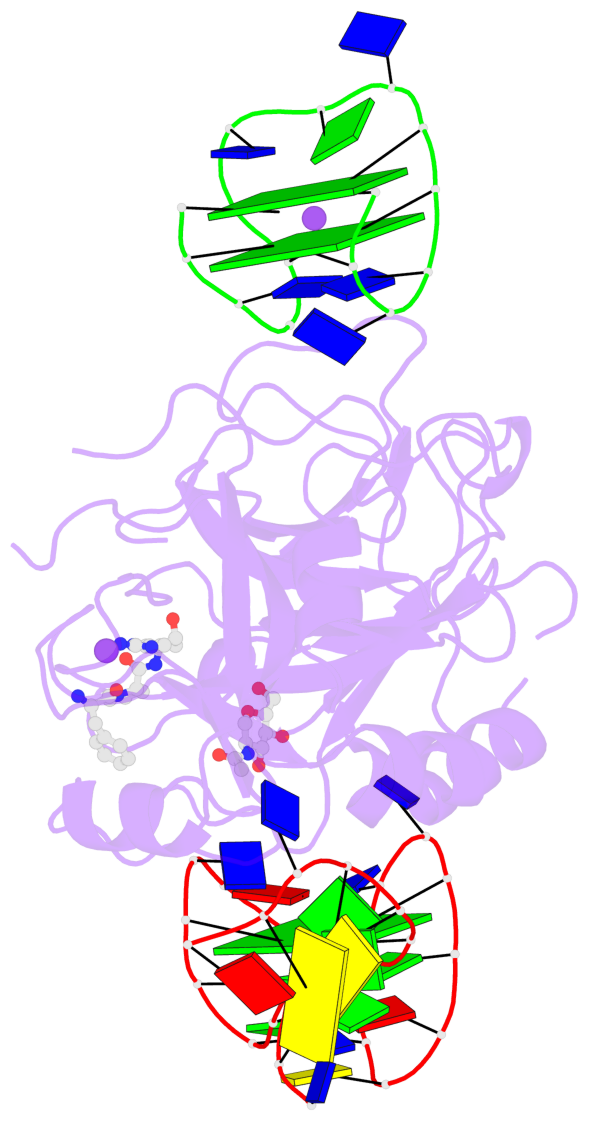
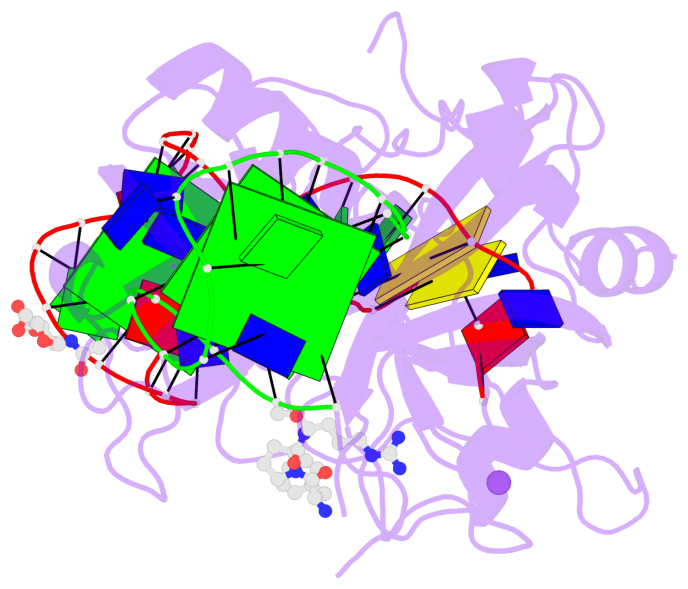
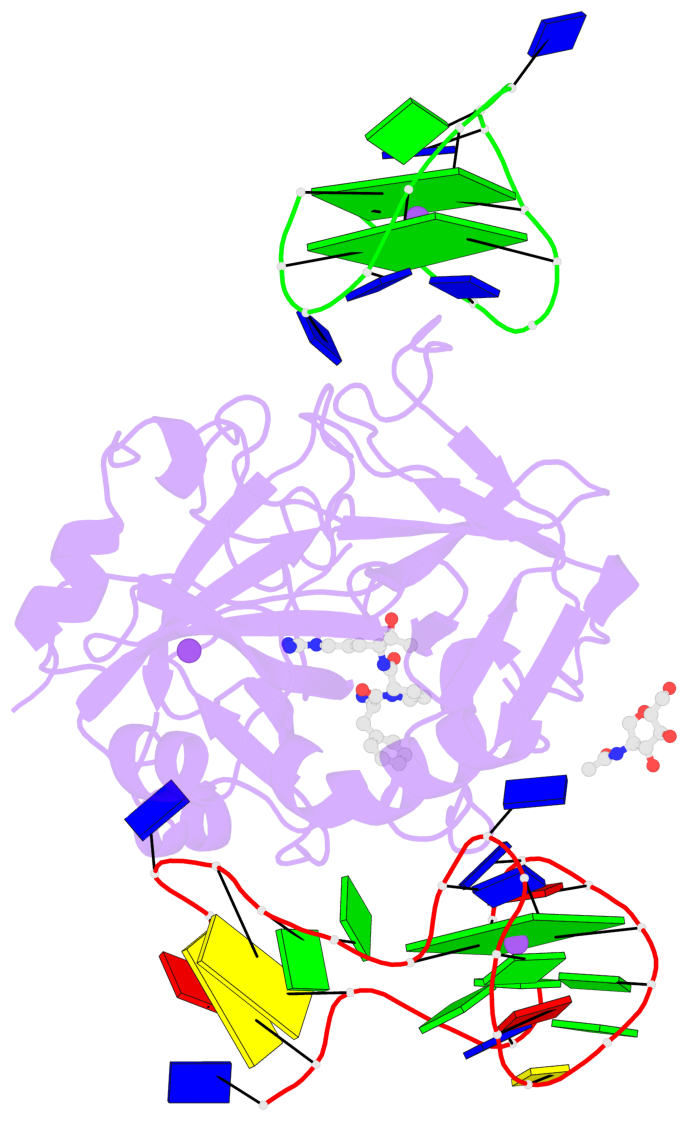
- 3 G-tetrads, 1 G4 helix, 1 G4 stem, 2(+Ln+Lw+Ln), chair(2+2), UDUD
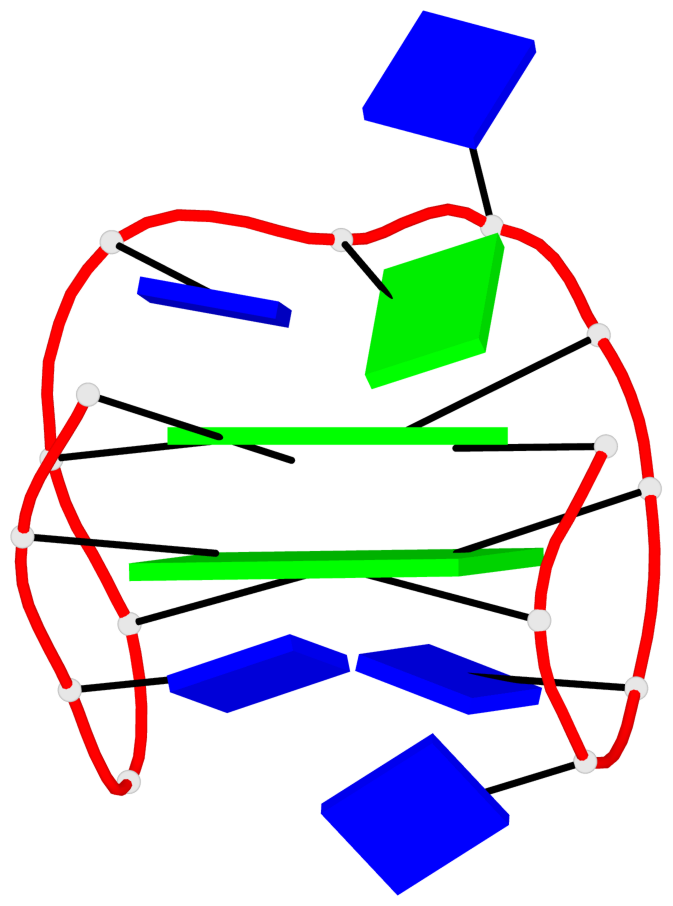
Base-block schematics in six views
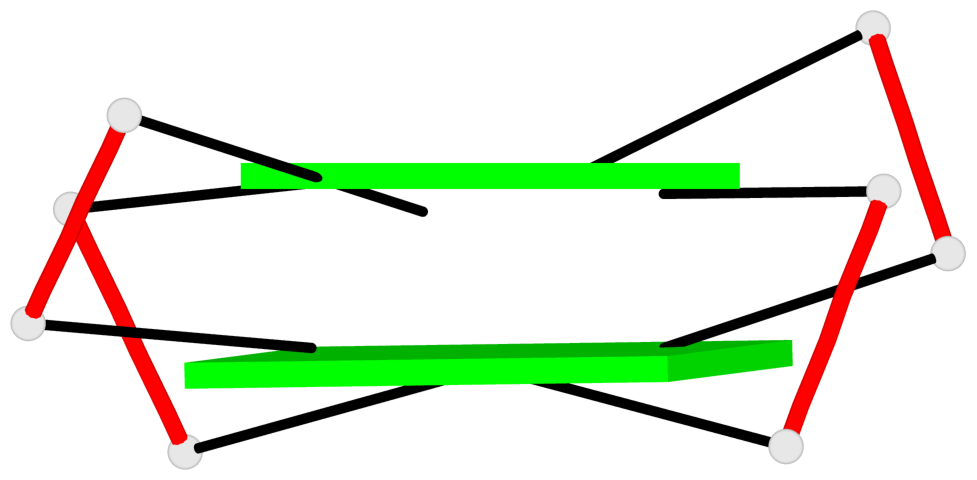
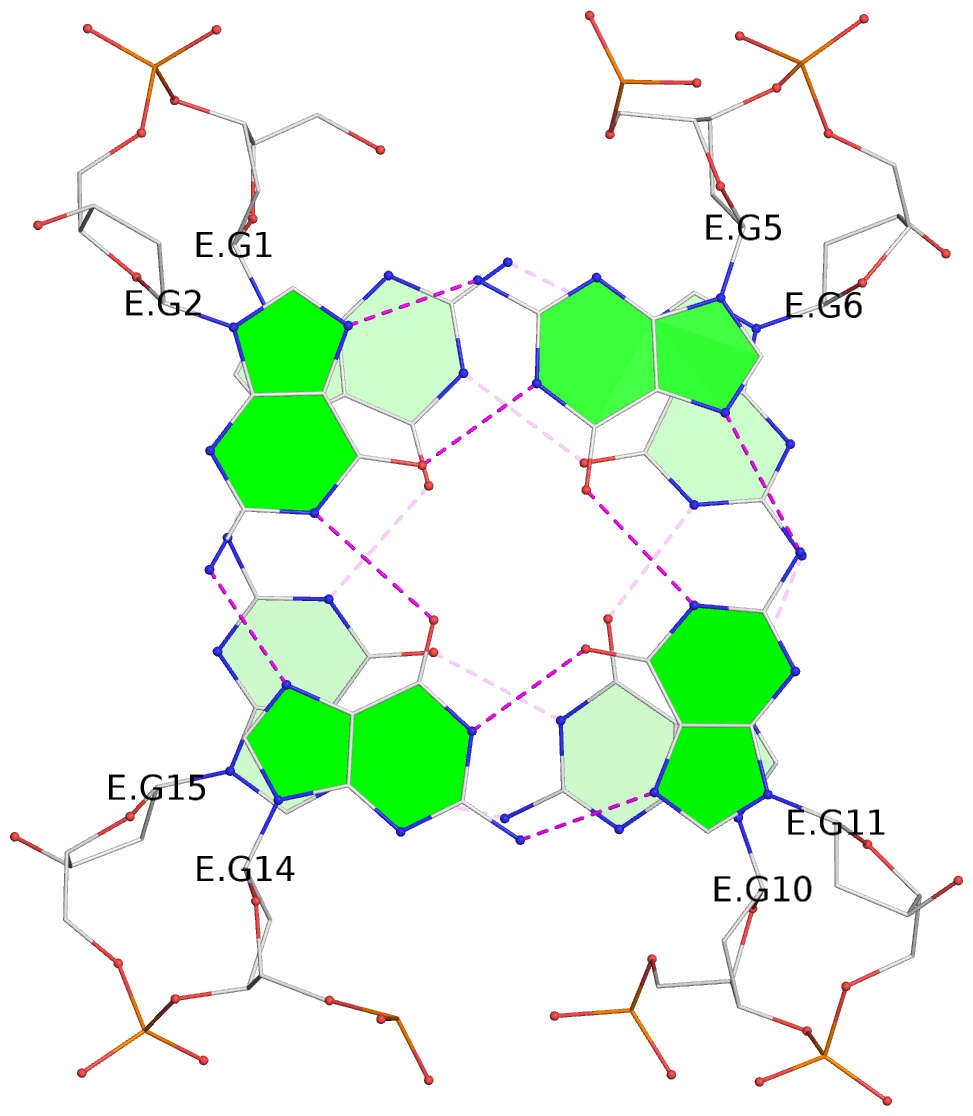
List of 3 G-tetrads
1 glyco-bond=-s-s sugar=-3-- groove=wnwn planarity=0.756 type=bowl nts=4 GGGG D.DG8,D.DG20,D.DG17,D.DG11 2 glyco-bond=s-s- sugar=---- groove=wnwn planarity=0.637 type=bowl nts=4 GGGG E.DG1,E.DG15,E.DG10,E.DG6 3 glyco-bond=-s-s sugar=---- groove=wnwn planarity=0.543 type=other nts=4 GGGG E.DG2,E.DG14,E.DG11,E.DG5
List of 1 G4-helix
In DSSR, a G4-helix is defined by stacking interactions of G-tetrads, regardless of backbone connectivity, and may contain more than one G4-stem.
Helix#1, 2 G-tetrad layers, INTRA-molecular, with 1 stem
List of 1 G4-stem
In DSSR, a G4-stem is defined as a G4-helix with backbone connectivity. Bulges are also allowed along each of the four strands.
Stem#1, 2 G-tetrad layers, 3 loops, INTRA-molecular, UDUD, anti-parallel, 2(+Ln+Lw+Ln), chair(2+2)
List of 3 non-stem G4-loops (including the two closing Gs)
1 type=lateral helix=#-1 nts=4 GTAG D.DG8,D.DT9,D.DA10,D.DG11 2 type=lateral helix=#-1 nts=7 GGGCAGG D.DG11,D.DG12,D.DG13,D.DC14,D.DA15,D.DG16,D.DG17 3 type=lateral helix=#-1 nts=4 GTTG D.DG17,D.DT18,D.DT19,D.DG20