Detailed DSSR results for the G-quadruplex: PDB entry 5ls8
Created and maintained by Xiang-Jun Lu <xiangjun@x3dna.org>
Citation: Please cite the NAR'20 DSSR-PyMOL schematics paper and/or the NAR'15 DSSR method paper.
Summary information
- PDB id
- 5ls8
- Class
- DNA
- Method
- X-ray (1.78 Å)
- Summary
- Light-activated ruthenium complex bound to a DNA quadruplex
- Reference
- McQuaid K, Abell H, Gurung SP, Allan DR, Winter G, Sorensen T, Cardin DJ, Brazier JA, Cardin CJ, Hall JP (2019): "Structural Studies Reveal Enantiospecific Recognition of a DNA G-Quadruplex by a Ruthenium Polypyridyl Complex." Angew.Chem.Int.Ed.Engl., 58, 9881-9885. doi: 10.1002/anie.201814502.
- Abstract
- By using X-ray crystallography, we show that the complexes Λ/Δ-[Ru(TAP)2 (11-CN-dppz)]2+ (TAP=1,4,5,8-tetraazaphenanthrene, dppz=dipyridophenazine) bind DNA G-quadruplex in an enantiospecific manner that parallels the specificity of these complexes with duplex DNA. The Λ complex crystallises with the normally parallel stranded d(TAGGGTTA) tetraplex to give the first such antiparallel strand assembly in which syn-guanosine is adjacent to the complex at the 5' end of the quadruplex core. SRCD measurements confirm that the same conformational switch occurs in solution. The Δ enantiomer, by contrast, is present in the structure but stacked at the ends of the assembly. In addition, we report the structure of Λ-[Ru(phen)2 (11-CN-dppz)]2+ bound to d(TCGGCGCCGA), a duplex-forming sequence, and use both structural models to provide insight into the motif-specific luminescence response of the isostructural phen analogue enantiomers.
- G4 notes
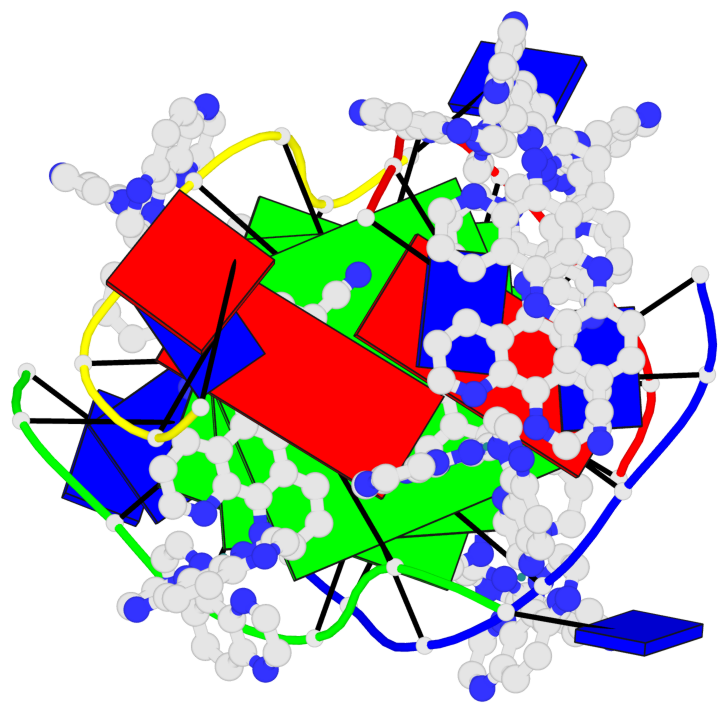
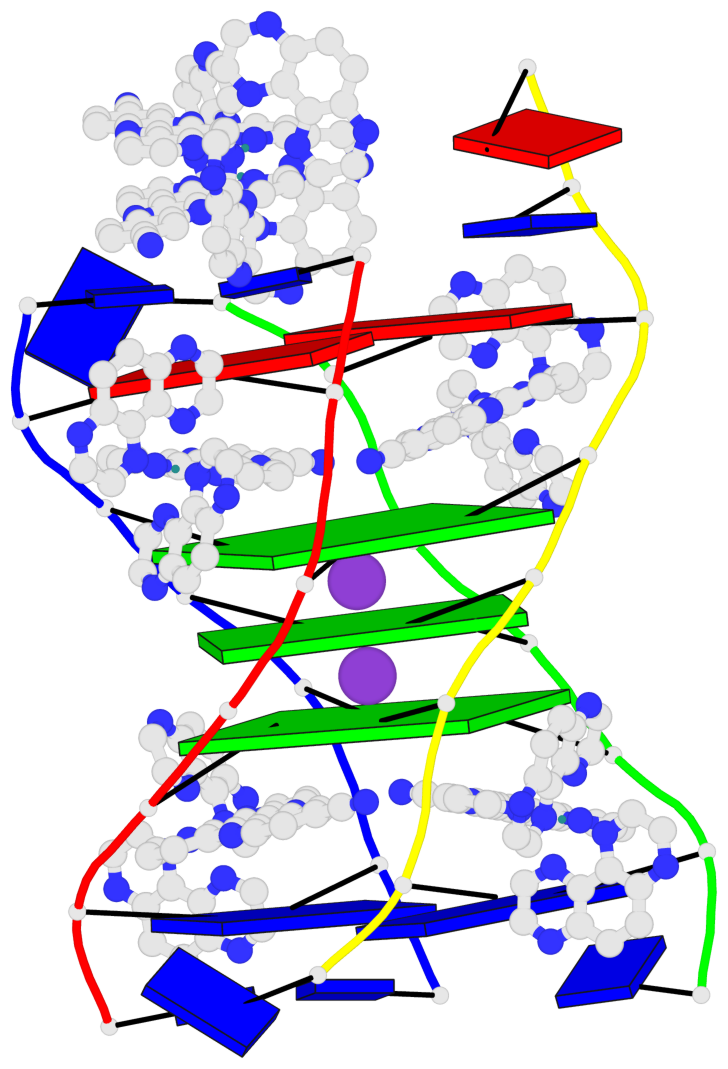
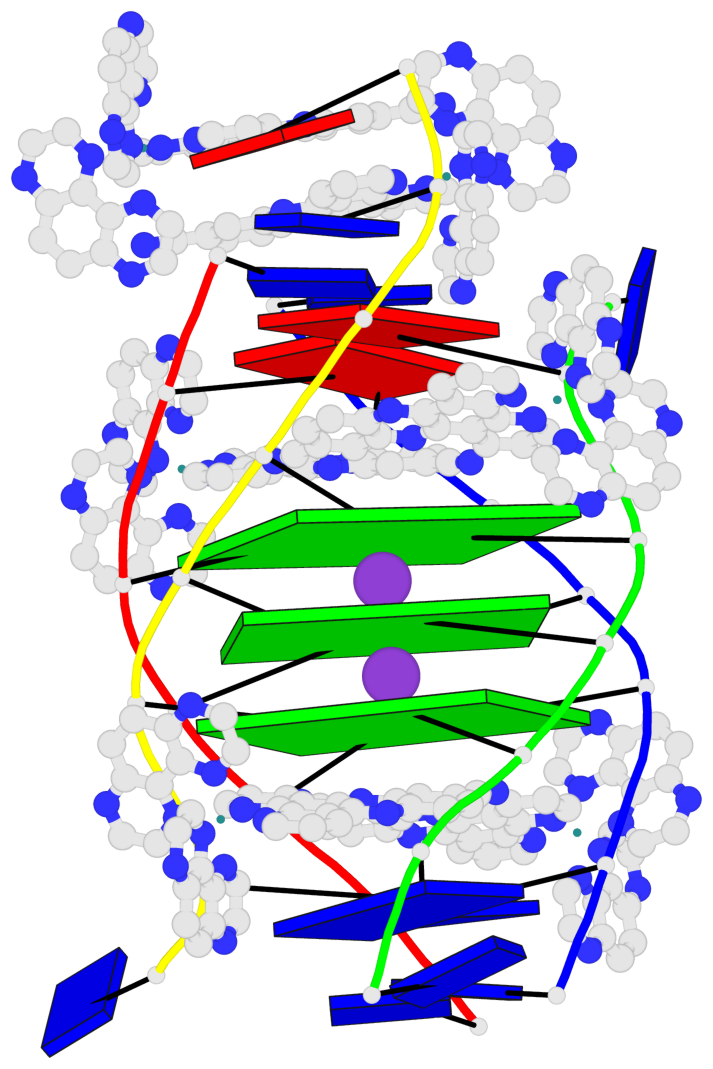
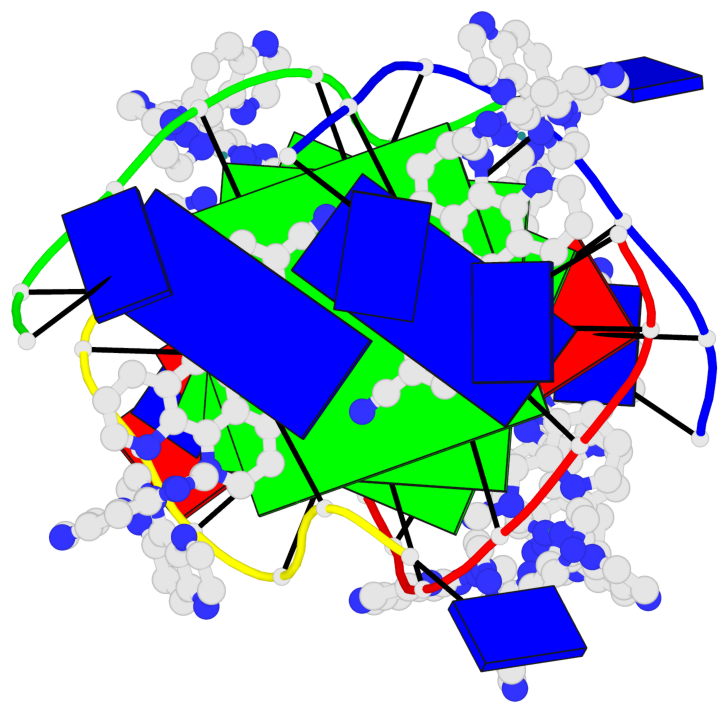
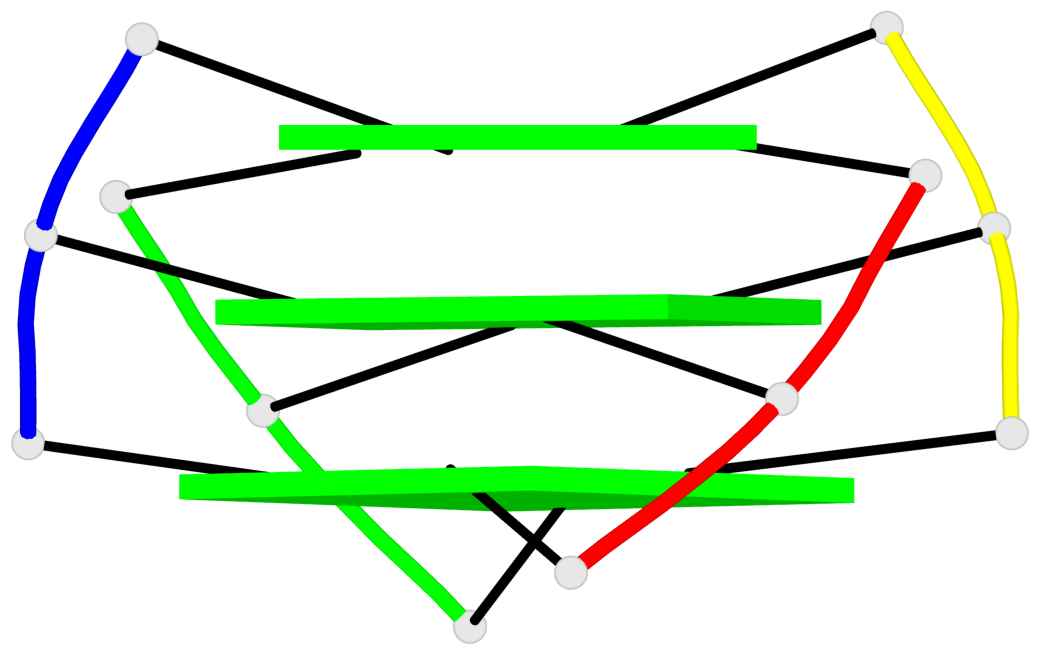
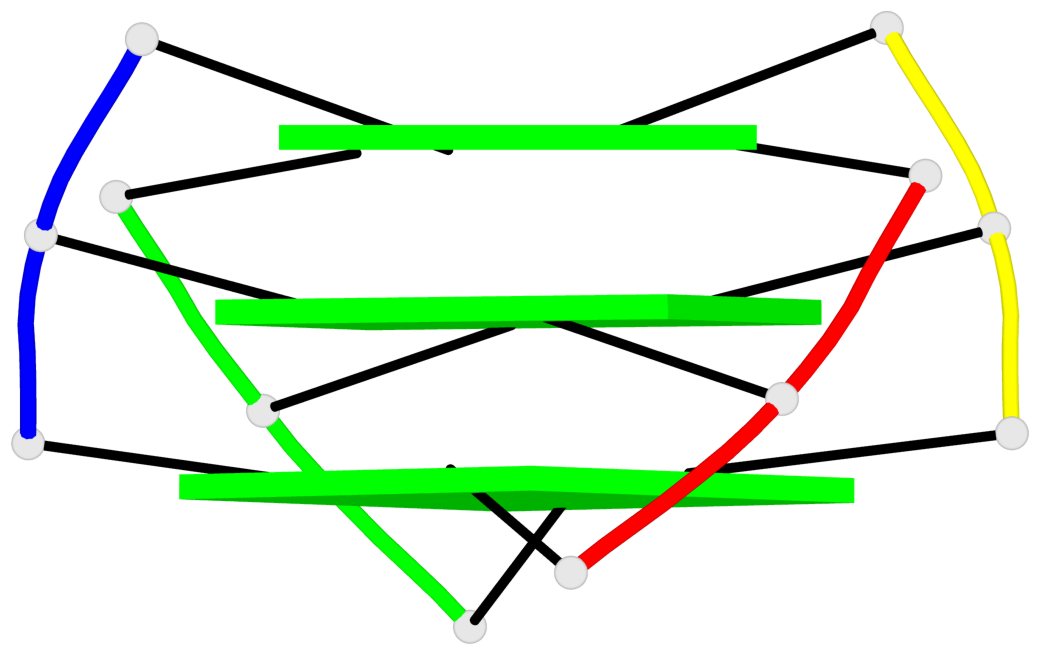
- 3 G-tetrads, 1 G4 helix, 1 G4 stem, (2+2), UDUD
Base-block schematics in six views
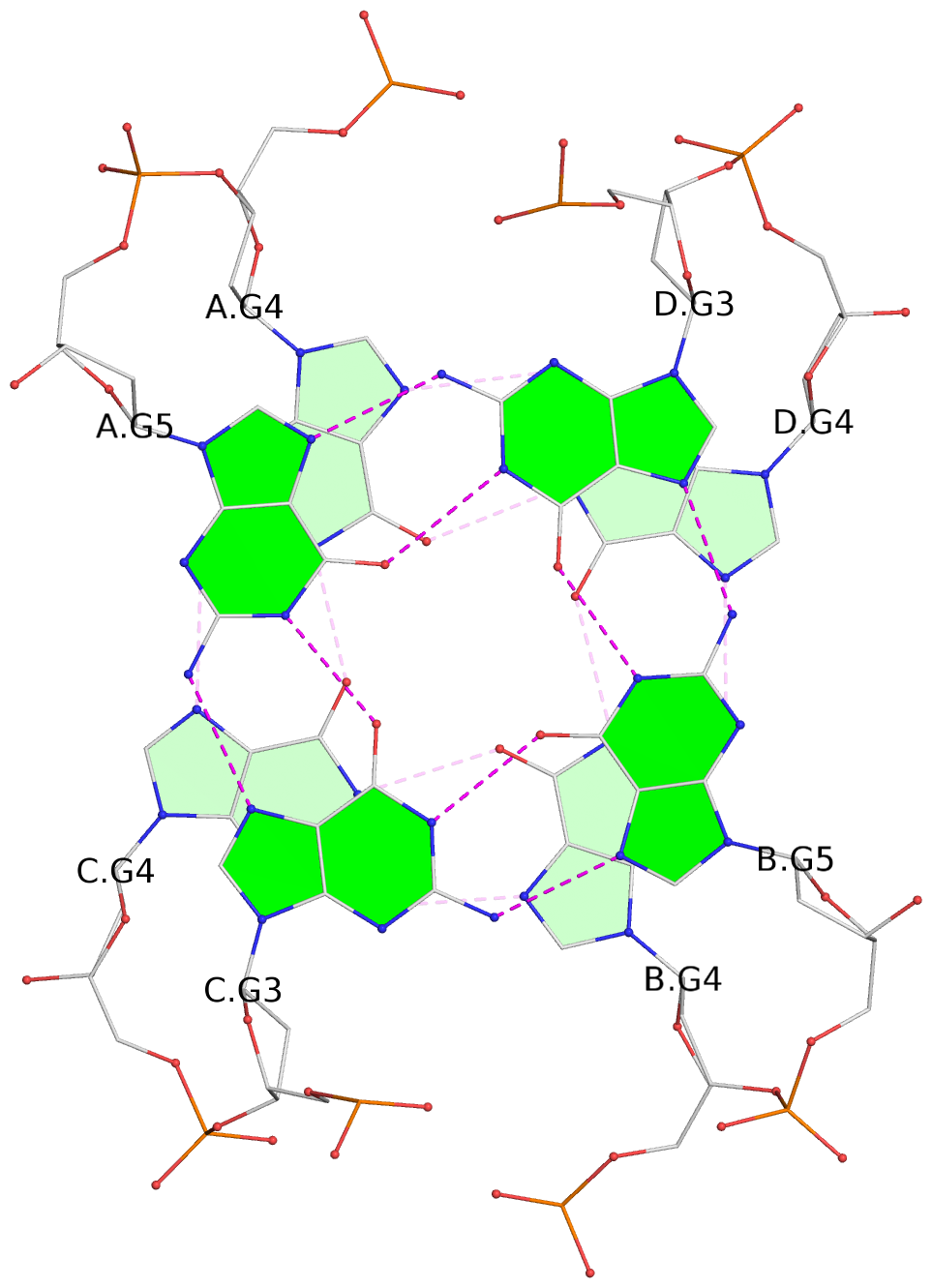
List of 3 G-tetrads
1 glyco-bond=s-s- sugar=---- groove=wnwn planarity=0.456 type=bowl-2 nts=4 GGGG A.DG3,C.DG5,B.DG3,D.DG5 2 glyco-bond=-s-s sugar=3--- groove=wnwn planarity=0.358 type=other nts=4 GGGG A.DG4,C.DG4,B.DG4,D.DG4 3 glyco-bond=-s-s sugar=---- groove=wnwn planarity=0.500 type=bowl-2 nts=4 GGGG A.DG5,C.DG3,B.DG5,D.DG3
List of 1 G4-helix
In DSSR, a G4-helix is defined by stacking interactions of G-tetrads, regardless of backbone connectivity, and may contain more than one G4-stem.
Helix#1, 3 G-tetrad layers, inter-molecular, with 1 stem
List of 1 G4-stem
In DSSR, a G4-stem is defined as a G4-helix with backbone connectivity. Bulges are also allowed along each of the four strands.