Detailed DSSR results for the G-quadruplex: PDB entry 5ob3
Created and maintained by Xiang-Jun Lu <xiangjun@x3dna.org>
Citation: Please cite the NAR'20 DSSR-PyMOL schematics paper and/or the NAR'15 DSSR method paper.
Summary information
- PDB id
- 5ob3
- Class
- RNA
- Method
- X-ray (2.004 Å)
- Summary
- Ispinach aptamer
- Reference
- Fernandez-Millan P, Autour A, Ennifar E, Westhof E, Ryckelynck M (2017): "Crystal structure and fluorescence properties of the iSpinach aptamer in complex with DFHBI." RNA, 23, 1788-1795. doi: 10.1261/rna.063008.117.
- Abstract
- Fluorogenic RNA aptamers are short nucleic acids able to specifically interact with small molecules and strongly enhance their fluorescence upon complex formation. Among the different systems recently introduced, Spinach, an aptamer forming a fluorescent complex with the 3,5-difluoro-4-hydroxybenzylidene imidazolinone (DFHBI), is one of the most promising. Using random mutagenesis and ultrahigh-throughput screening, we recently developed iSpinach, an improved version of the aptamer, endowed with an increased folding efficiency and thermal stability. iSpinach is a shorter version of Spinach, comprising five mutations for which the exact role has not yet been deciphered. In this work, we cocrystallized a reengineered version of iSpinach in complex with the DFHBI and solved the X-ray structure of the complex at 2 Å resolution. Only a few mutations were required to optimize iSpinach production and crystallization, underlying the good folding capacity of the molecule. The measured fluorescence half-lives in the crystal were 60% higher than in solution. Comparisons with structures previously reported for Spinach sheds some light on the possible function of the different beneficial mutations carried by iSpinach.
- G4 notes
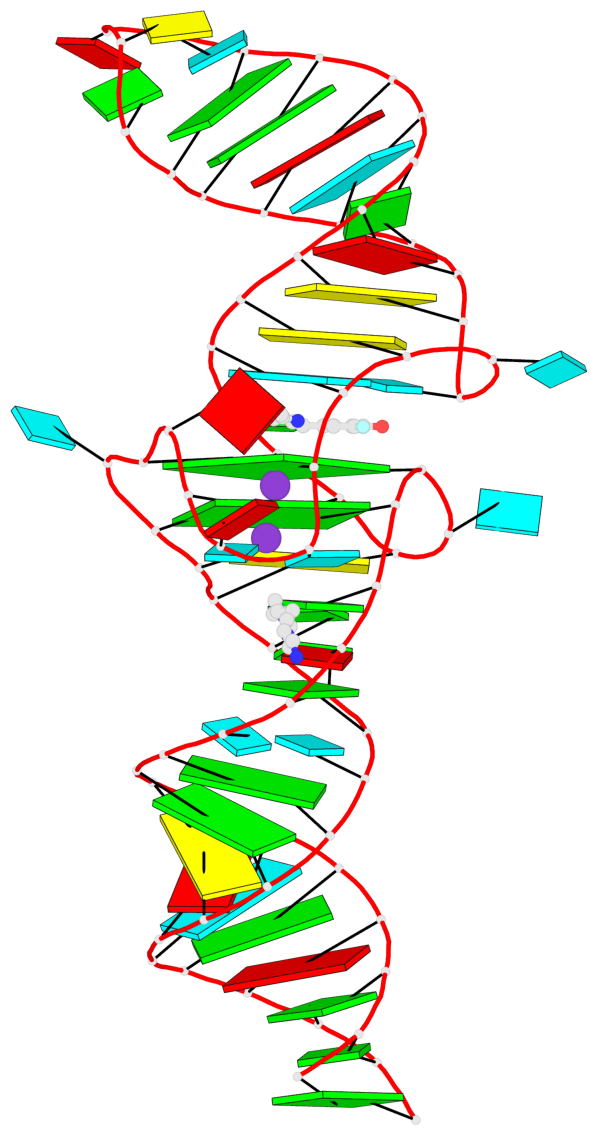
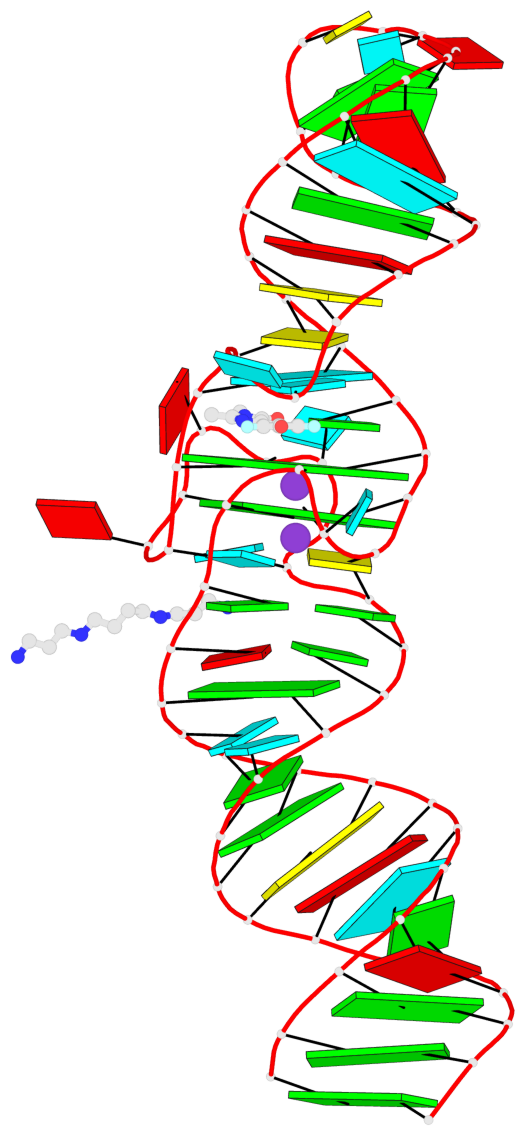
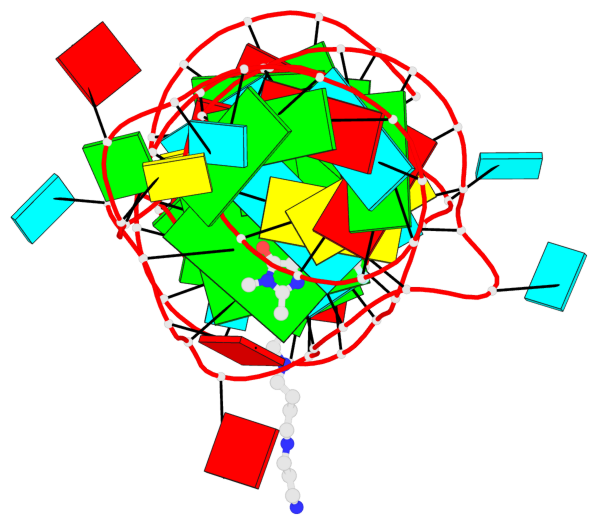
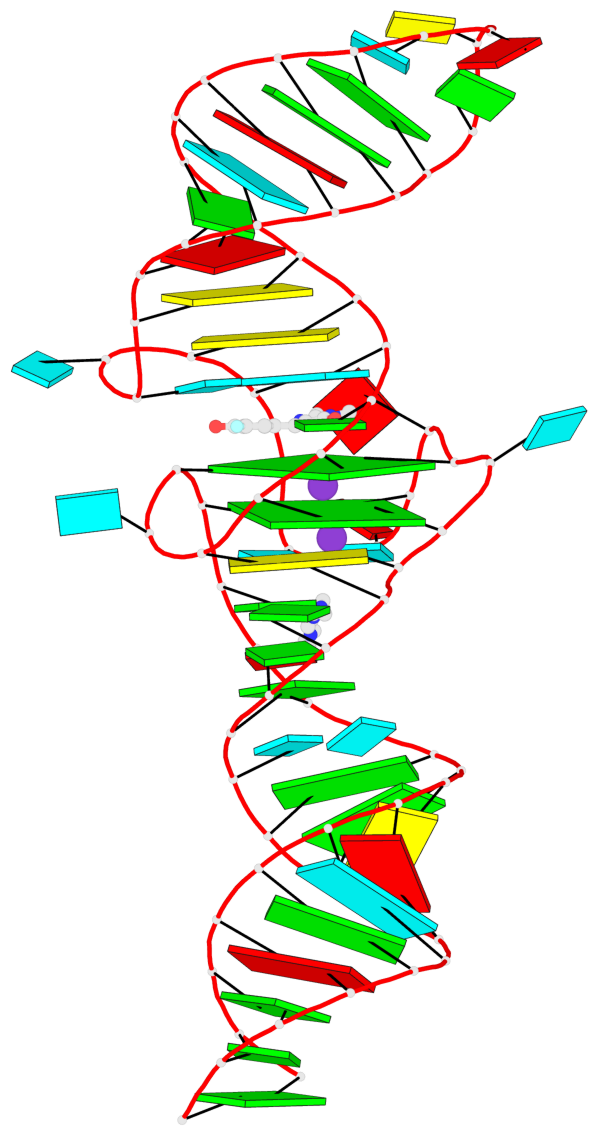
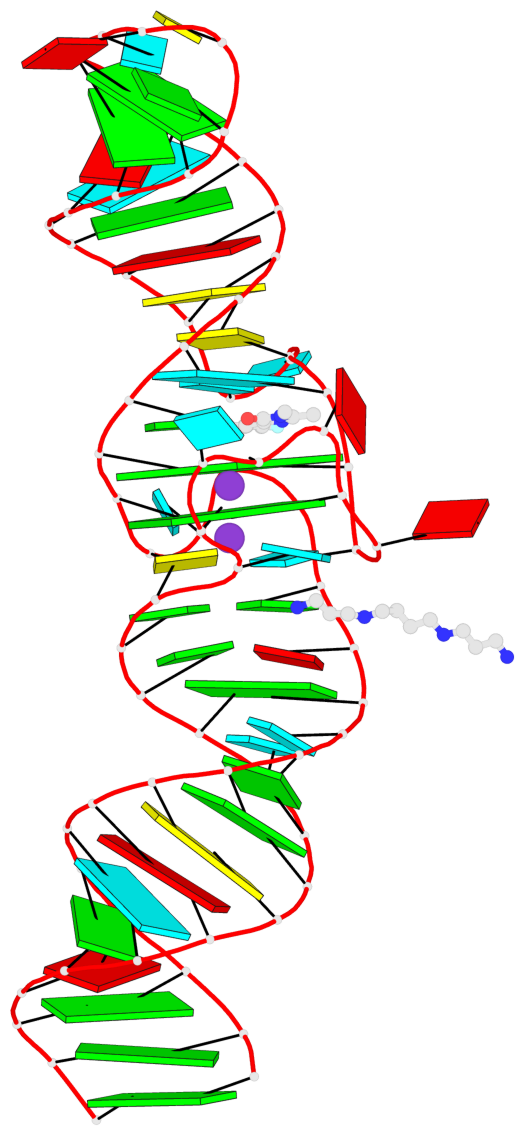
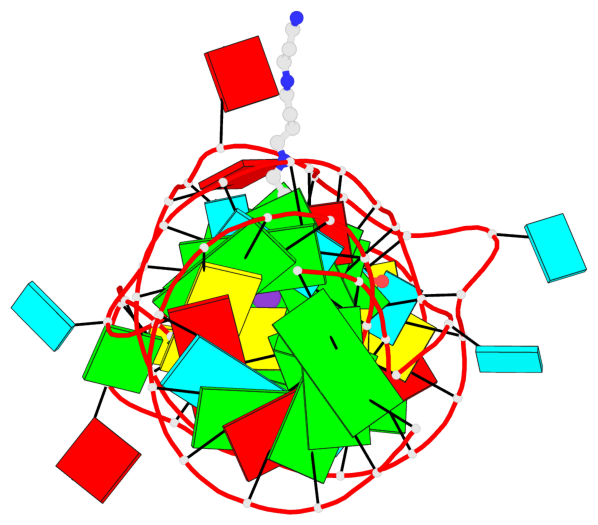
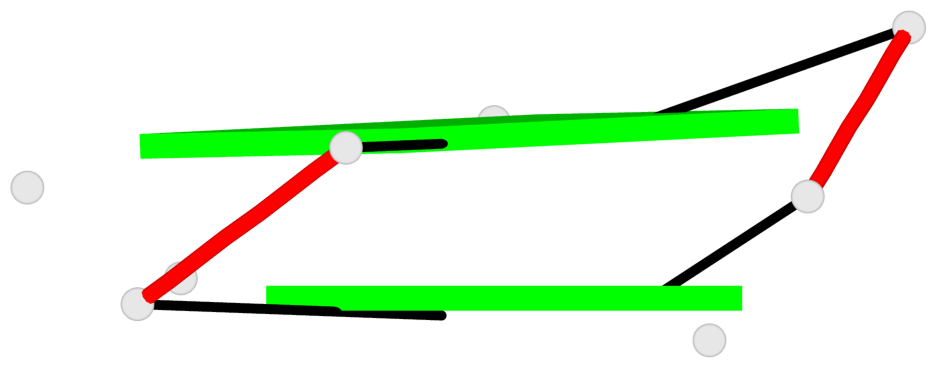
- 2 G-tetrads, 1 G4 helix, 1 G4 stem, 2(-PD+P), (2+2), UUDD
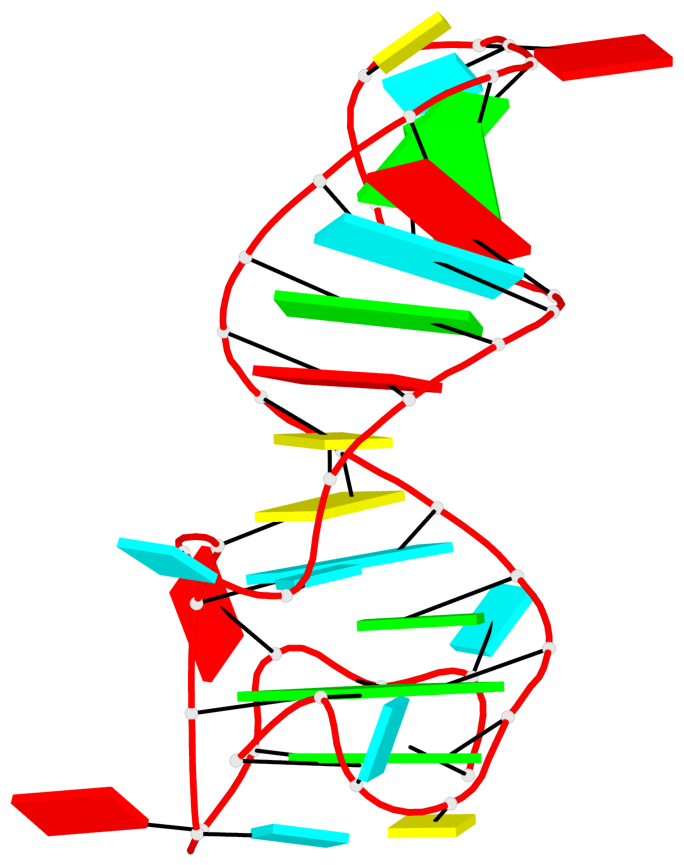
Base-block schematics in six views
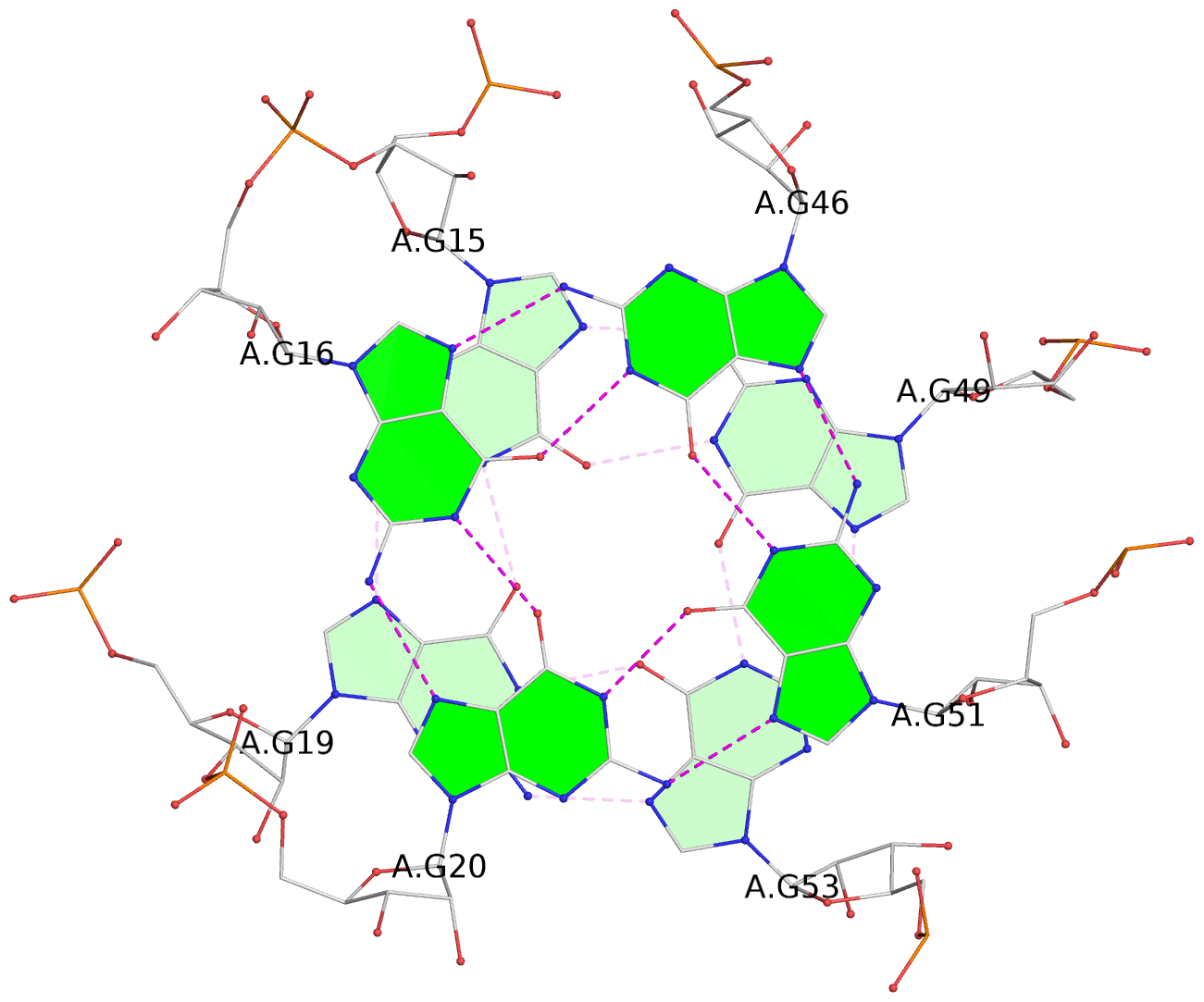
List of 2 G-tetrads
1 glyco-bond=--s- sugar=-33. groove=-wn- planarity=0.297 type=other nts=4 GGGG A.G15,A.G19,A.G53,A.G49 2 glyco-bond=--ss sugar=-3-3 groove=-w-n planarity=0.194 type=other nts=4 GGGG A.G16,A.G20,A.G51,A.G46
List of 1 G4-helix
In DSSR, a G4-helix is defined by stacking interactions of G-tetrads, regardless of backbone connectivity, and may contain more than one G4-stem.
Helix#1, 2 G-tetrad layers, INTRA-molecular, with 1 stem
List of 1 G4-stem
In DSSR, a G4-stem is defined as a G4-helix with backbone connectivity. Bulges are also allowed along each of the four strands.