Detailed DSSR results for the G-quadruplex: PDB entry 5ua3
Created and maintained by Xiang-Jun Lu <xiangjun@x3dna.org>
Citation: Please cite the NAR'20 DSSR-PyMOL schematics paper and/or the NAR'15 DSSR method paper.
Summary information
- PDB id
- 5ua3
- Class
- DNA
- Method
- X-ray (1.88 Å)
- Summary
- Crystal structure of a DNA g-quadruplex with a cytosine bulge
- Reference
- Meier M, Moya-Torres A, Krahn NJ, McDougall MD, Orriss GL, McRae EKS, Booy EP, McEleney K, Patel TR, McKenna SA, Stetefeld J (2018): "Structure and hydrodynamics of a DNA G-quadruplex with a cytosine bulge." Nucleic Acids Res., 46, 5319-5331. doi: 10.1093/nar/gky307.
- Abstract
- The identification of four-stranded G-quadruplexes (G4s) has highlighted the fact that DNA has additional spatial organisations at its disposal other than double-stranded helices. Recently, it became clear that the formation of G4s is not limited to the traditional G3+NL1G3+NL2G3+NL3G3+ sequence motif. Instead, the G3 triplets can be interrupted by deoxythymidylate (DNA) or uridylate (RNA) where the base forms a bulge that loops out from the G-quadruplex core. Here, we report the first high-resolution X-ray structure of a unique unimolecular DNA G4 with a cytosine bulge. The G4 forms a dimer that is stacked via its 5'-tetrads. Analytical ultracentrifugation, static light scattering and small angle X-ray scattering confirmed that the G4 adapts a predominantly dimeric structure in solution. We provide a comprehensive comparison of previously published G4 structures containing bulges and report a special γ torsion angle range preferentially populated by the G4 core guanylates adjacent to bulges. Since the penalty for introducing bulges appears to be negligible, it should be possible to functionalize G4s by introducing artificial or modified nucleotides at such positions. The presence of the bulge alters the surface of the DNA, providing an opportunity to develop drugs that can specifically target individual G4s.
- G4 notes
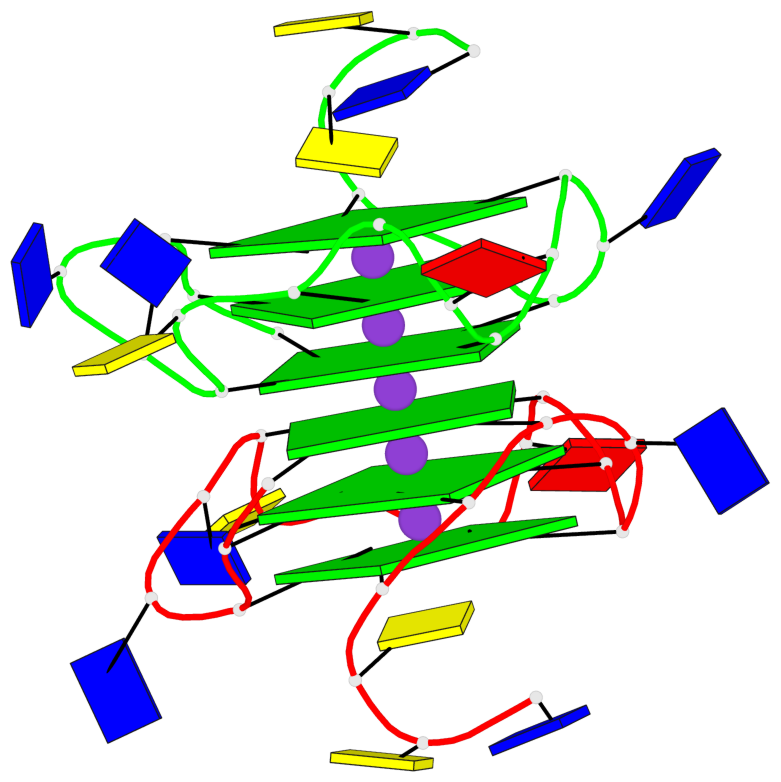
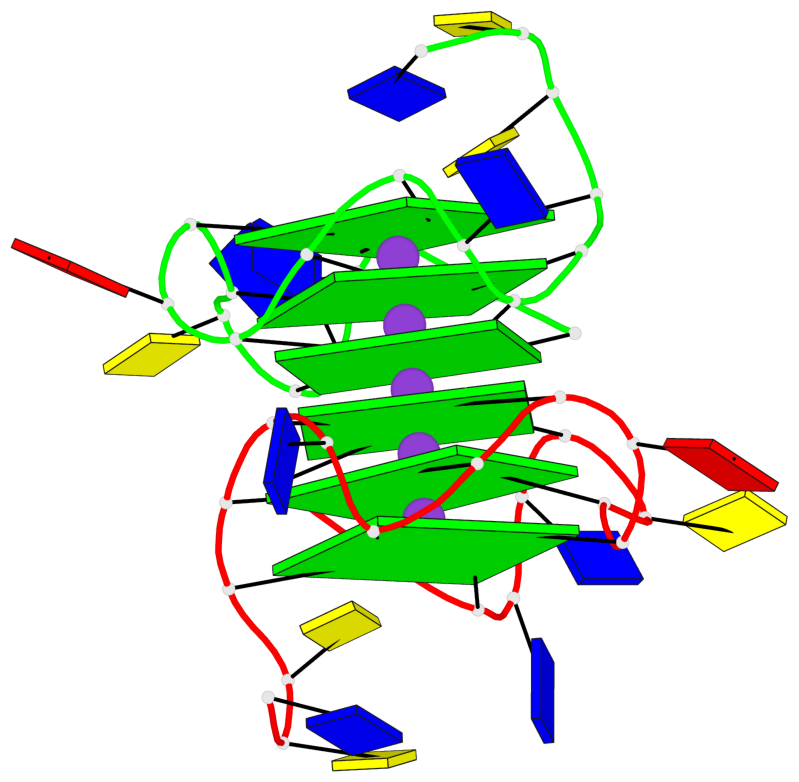
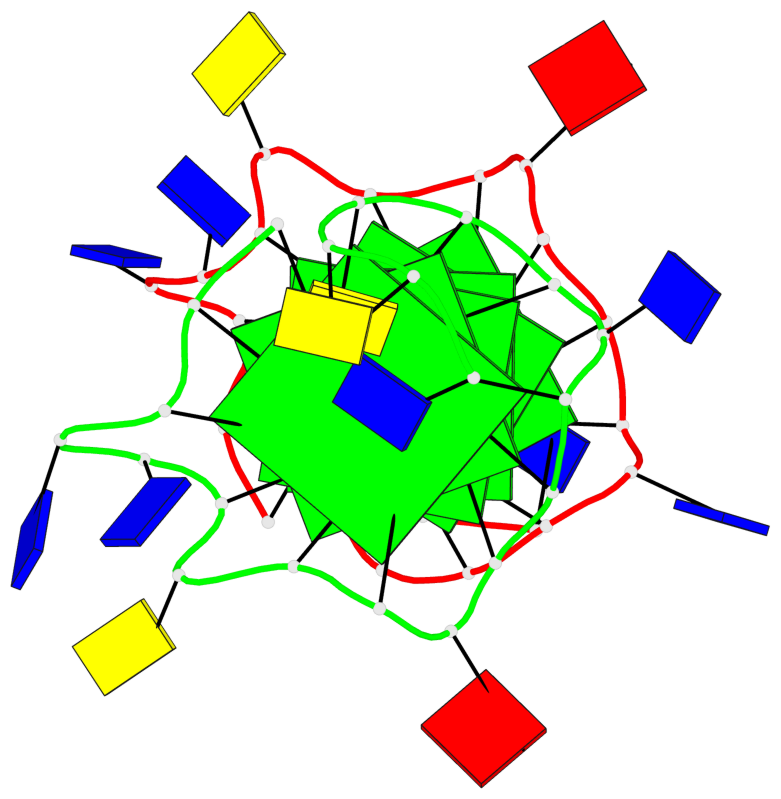
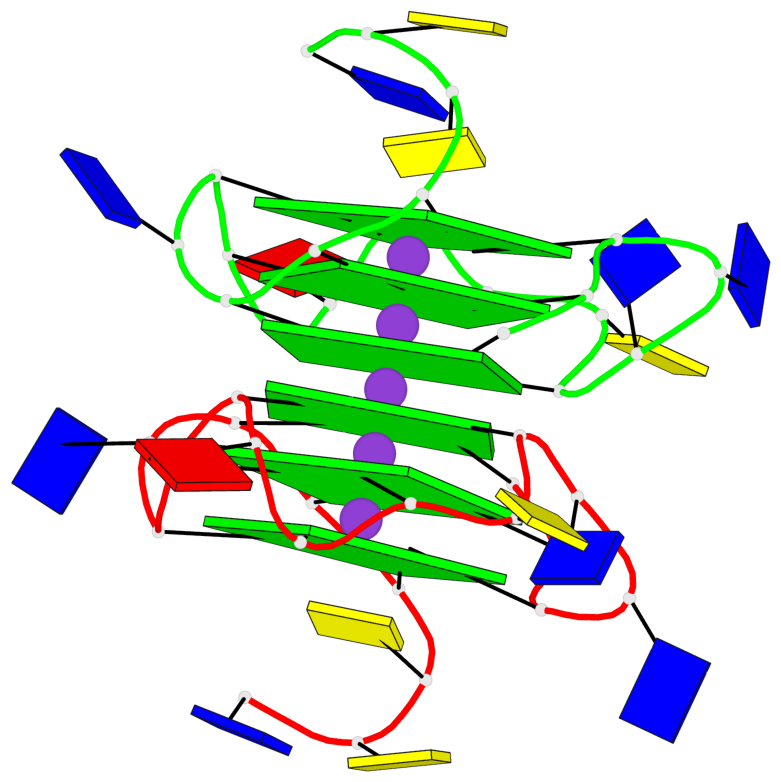
- 6 G-tetrads, 1 G4 helix, 2 G4 stems, 1 G4 coaxial stack, 3(-P-P-P), parallel(4+0), UUUU, coaxial interfaces: 5'/5'
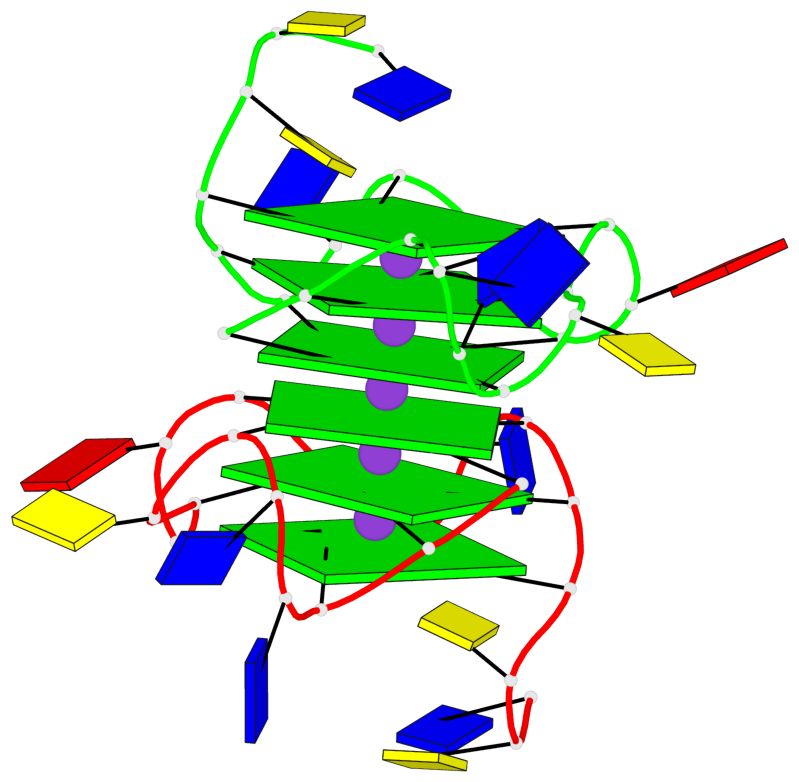
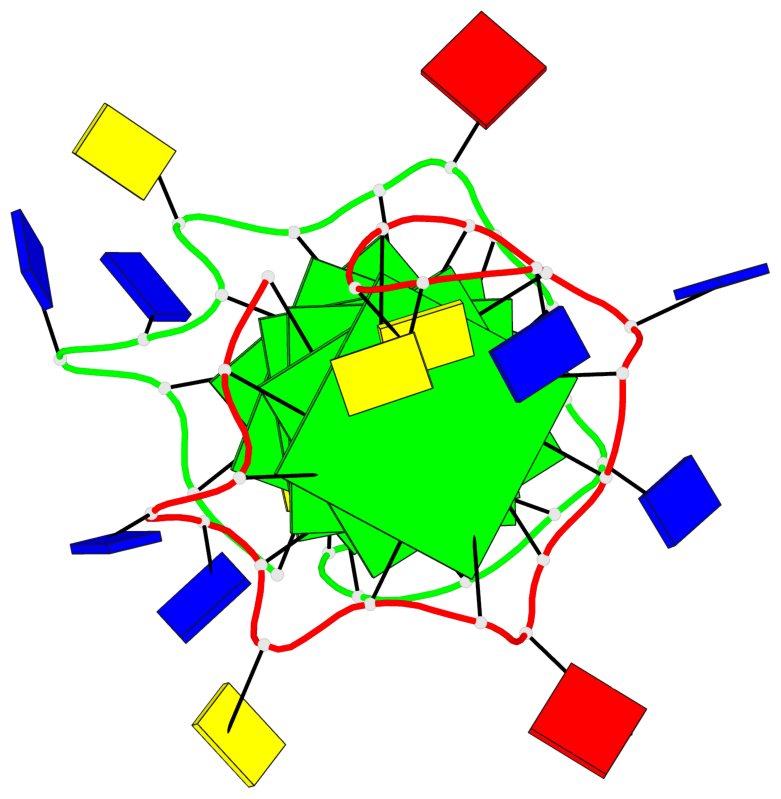
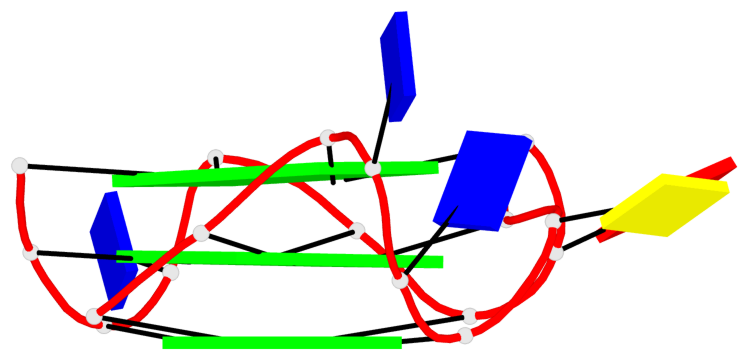
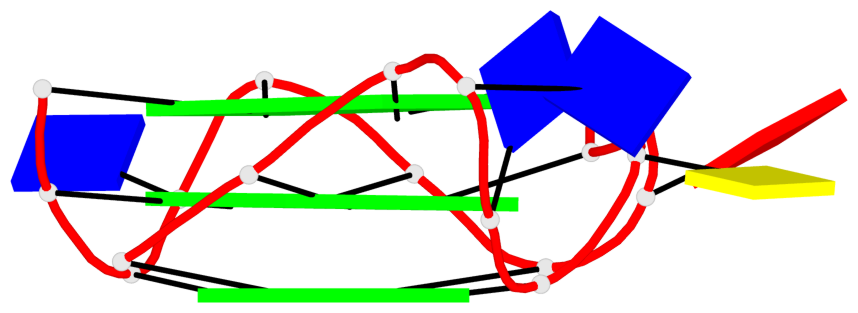
Base-block schematics in six views
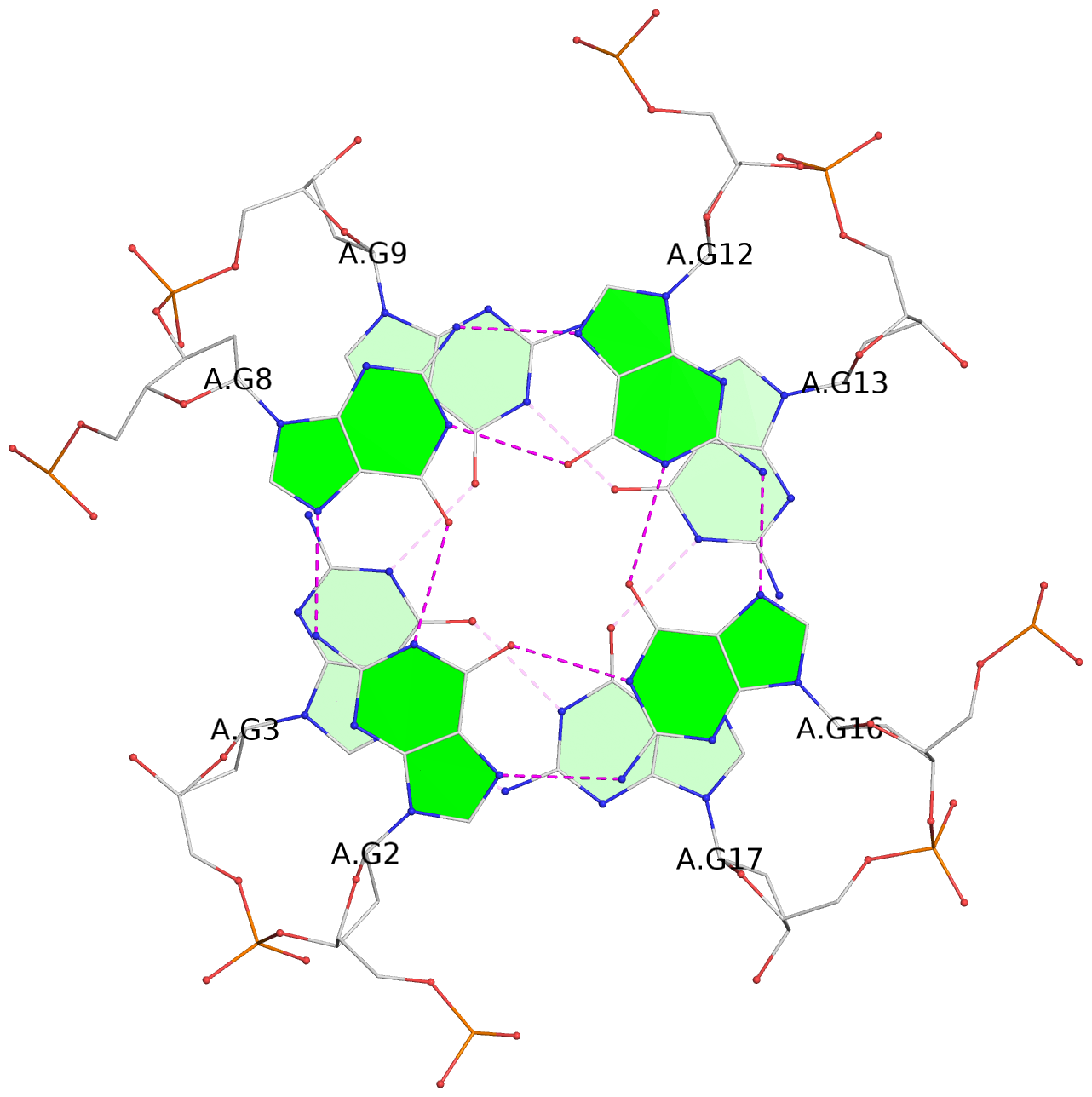
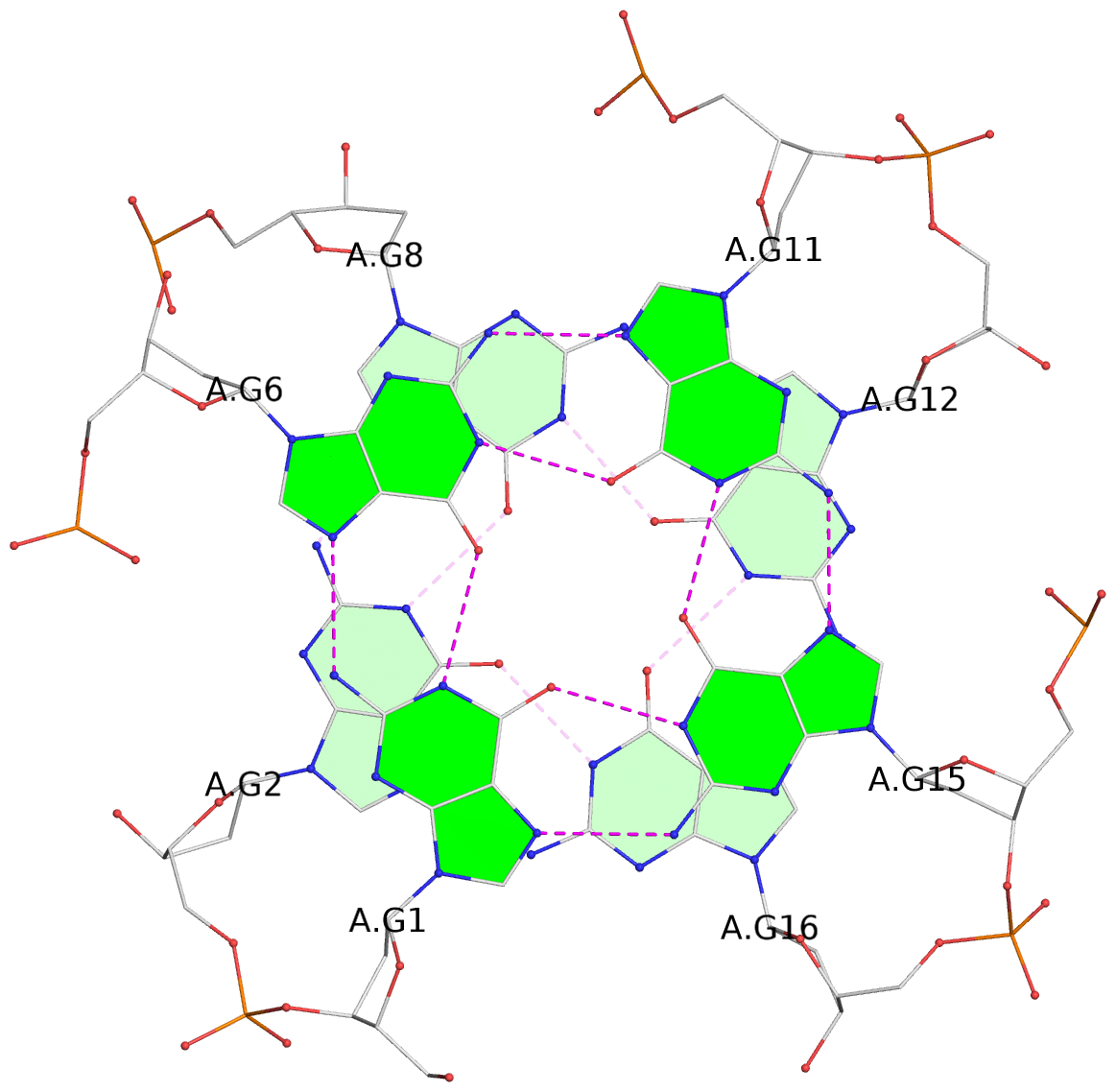
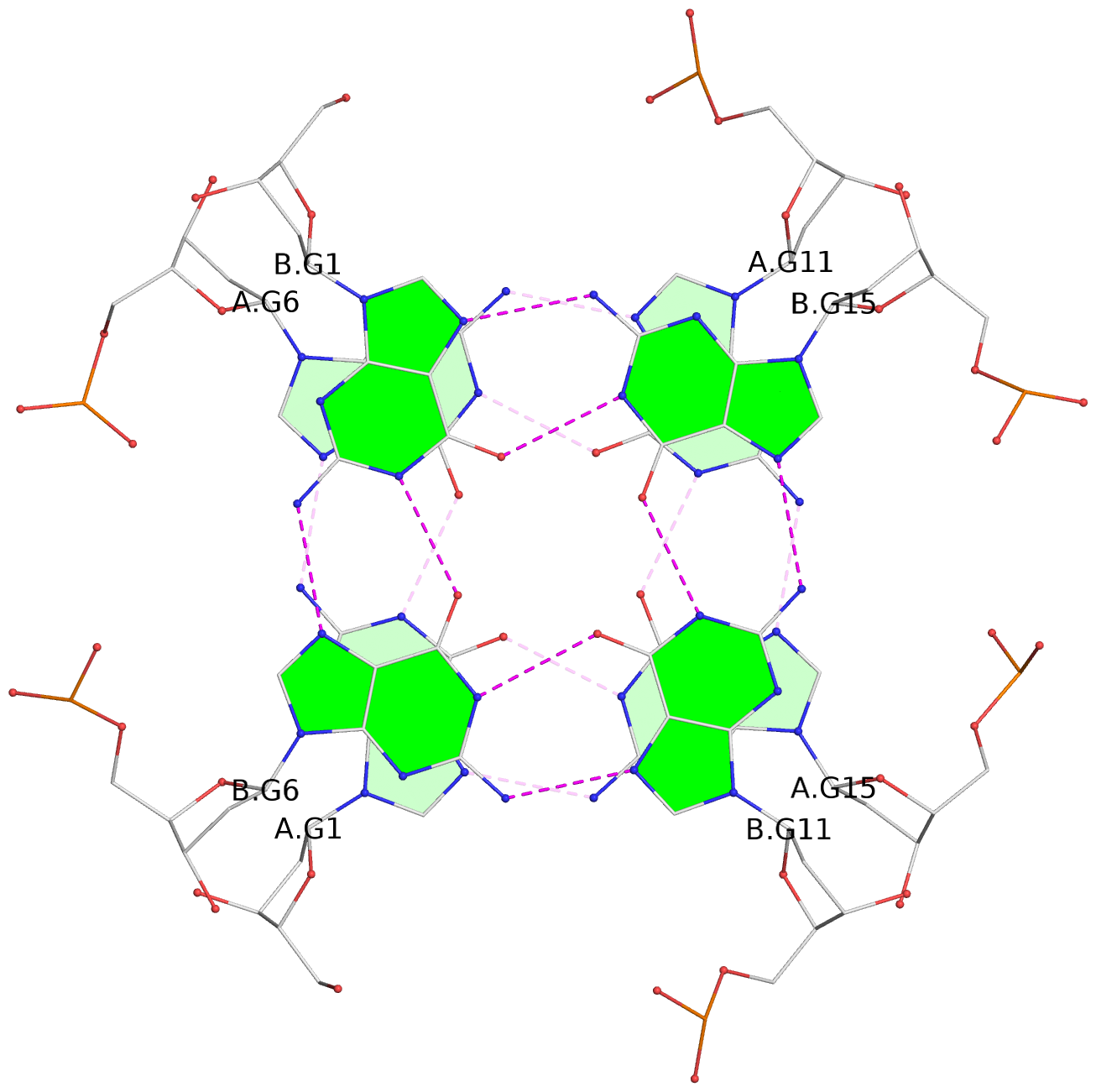
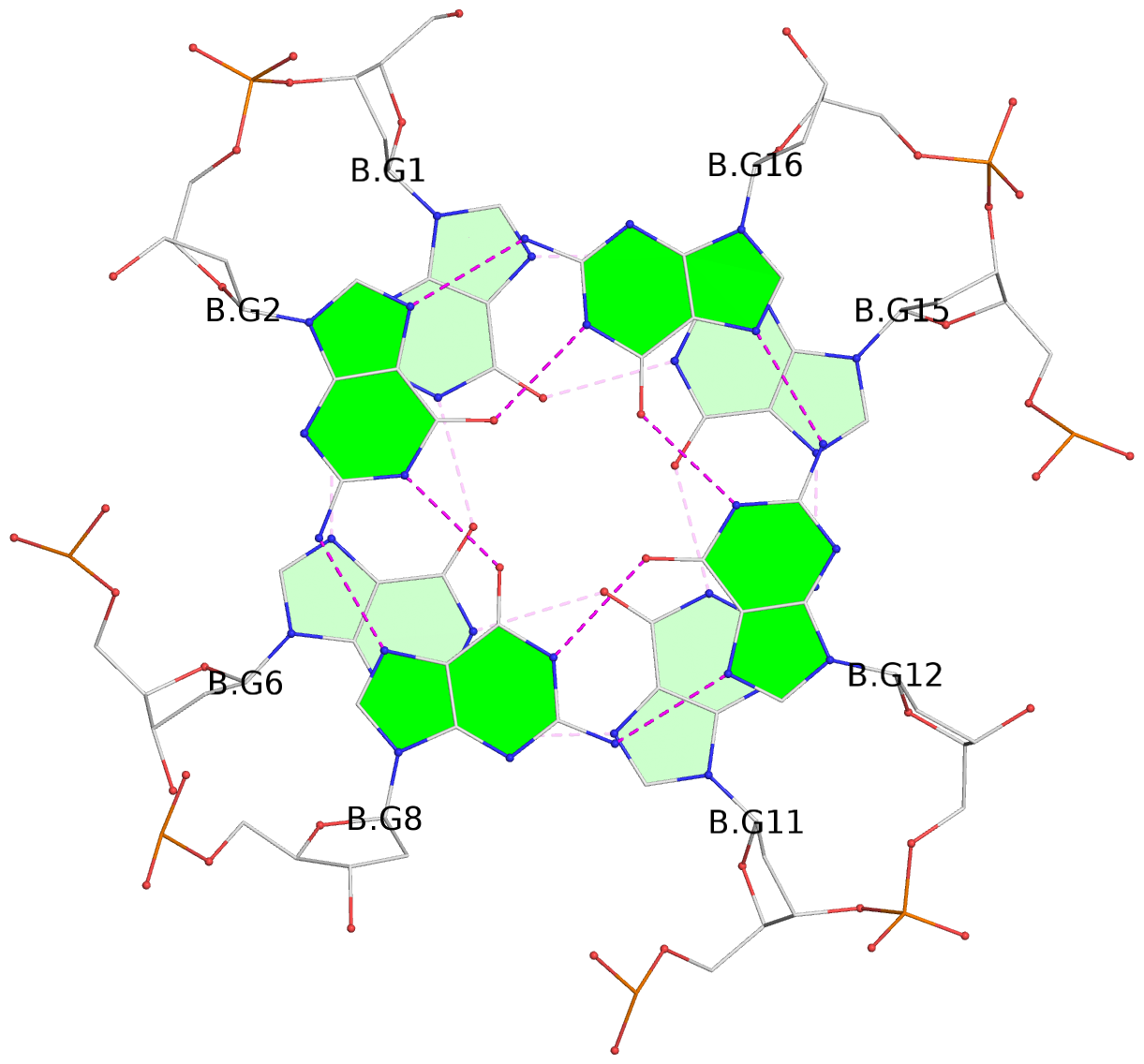
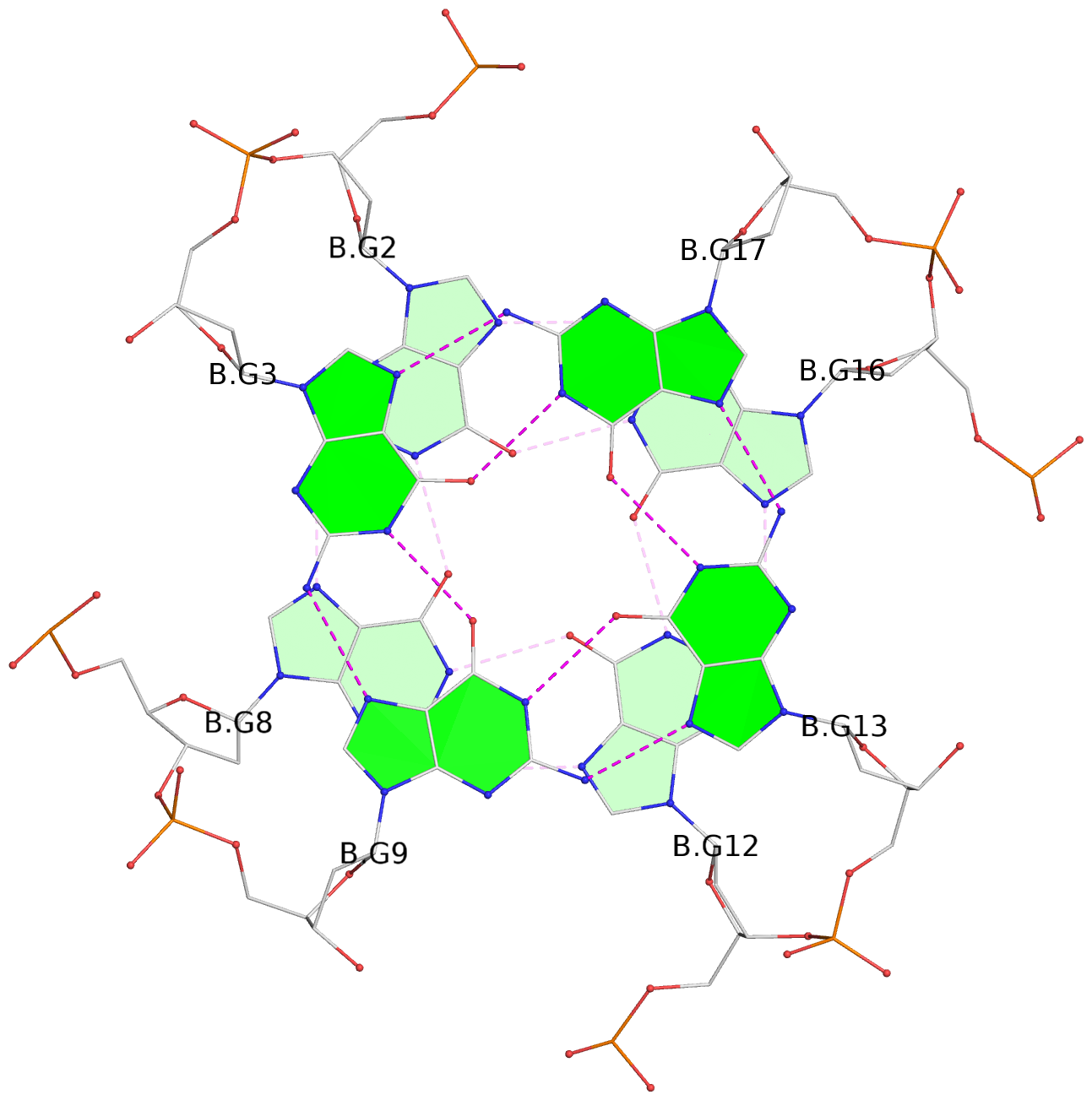
List of 6 G-tetrads
1 glyco-bond=---- sugar=---- groove=---- planarity=0.097 type=planar nts=4 GGGG A.DG1,A.DG6,A.DG11,A.DG15 2 glyco-bond=---- sugar=-3-- groove=---- planarity=0.195 type=bowl-2 nts=4 GGGG A.DG2,A.DG8,A.DG12,A.DG16 3 glyco-bond=---- sugar=---- groove=---- planarity=0.229 type=bowl nts=4 GGGG A.DG3,A.DG9,A.DG13,A.DG17 4 glyco-bond=---- sugar=---- groove=---- planarity=0.088 type=planar nts=4 GGGG B.DG1,B.DG6,B.DG11,B.DG15 5 glyco-bond=---- sugar=-3-- groove=---- planarity=0.171 type=bowl-2 nts=4 GGGG B.DG2,B.DG8,B.DG12,B.DG16 6 glyco-bond=---- sugar=---- groove=---- planarity=0.275 type=bowl nts=4 GGGG B.DG3,B.DG9,B.DG13,B.DG17
List of 1 G4-helix
In DSSR, a G4-helix is defined by stacking interactions of G-tetrads, regardless of backbone connectivity, and may contain more than one G4-stem.
Helix#1, 6 G-tetrad layers, inter-molecular, with 2 stems
List of 2 G4-stems
In DSSR, a G4-stem is defined as a G4-helix with backbone connectivity. Bulges are also allowed along each of the four strands.
Stem#1, 3 G-tetrad layers, 3 loops, INTRA-molecular, UUUU, parallel, 3(-P-P-P), parallel(4+0)
Stem#2, 3 G-tetrad layers, 3 loops, INTRA-molecular, UUUU, parallel, 3(-P-P-P), parallel(4+0)
List of 1 G4 coaxial stack
1 G4 helix#1 contains 2 G4 stems: [#1,#2] [5'/5']