Detailed DSSR results for the G-quadruplex: PDB entry 6ac7
Created and maintained by Xiang-Jun Lu <xiangjun@x3dna.org>
Citation: Please cite the NAR'20 DSSR-PyMOL schematics paper and/or the NAR'15 DSSR method paper.
Summary information
- PDB id
- 6ac7
- Class
- DNA
- Method
- NMR
- Summary
- Structure of a (3+1) hybrid g-quadruplex in the parp1 promoter
- Reference
- Sengar A, Vandana JJ, Chambers VS, Di Antonio M, Winnerdy FR, Balasubramanian S, Phan AT (2019): "Structure of a (3+1) hybrid G-quadruplex in the PARP1 promoter." Nucleic Acids Res., 47, 1564-1572. doi: 10.1093/nar/gky1179.
- Abstract
- Poly (ADP-ribose) polymerase 1 (PARP1) has emerged as an attractive target for cancer therapy due to its key role in DNA repair processes. Inhibition of PARP1 in BRCA-mutated cancers has been observed to be clinically beneficial. Recent genome-mapping experiments have identified a non-canonical G-quadruplex-forming sequence containing bulges within the PARP1 promoter. Structural features, like bulges, provide opportunities for selective chemical targeting of the non-canonical G-quadruplex structure within the PARP1 promoter, which could serve as an alternative therapeutic approach for the regulation of PARP1 expression. Here we report the G-quadruplex structure formed by a 23-nucleotide G-rich sequence in the PARP1 promoter. Our study revealed a three-layered intramolecular (3+1) hybrid G-quadruplex scaffold, in which three strands are oriented in one direction and the fourth in the opposite direction. This structure exhibits unique structural features such as an adenine bulge and a G·G·T base triple capping structure formed between the central edgewise loop, propeller loop and 5' flanking terminal. Given the highly important role of PARP1 in DNA repair and cancer intervention, this structure presents an attractive opportunity to explore the therapeutic potential of PARP1 inhibition via G-quadruplex DNA targeting.
- G4 notes
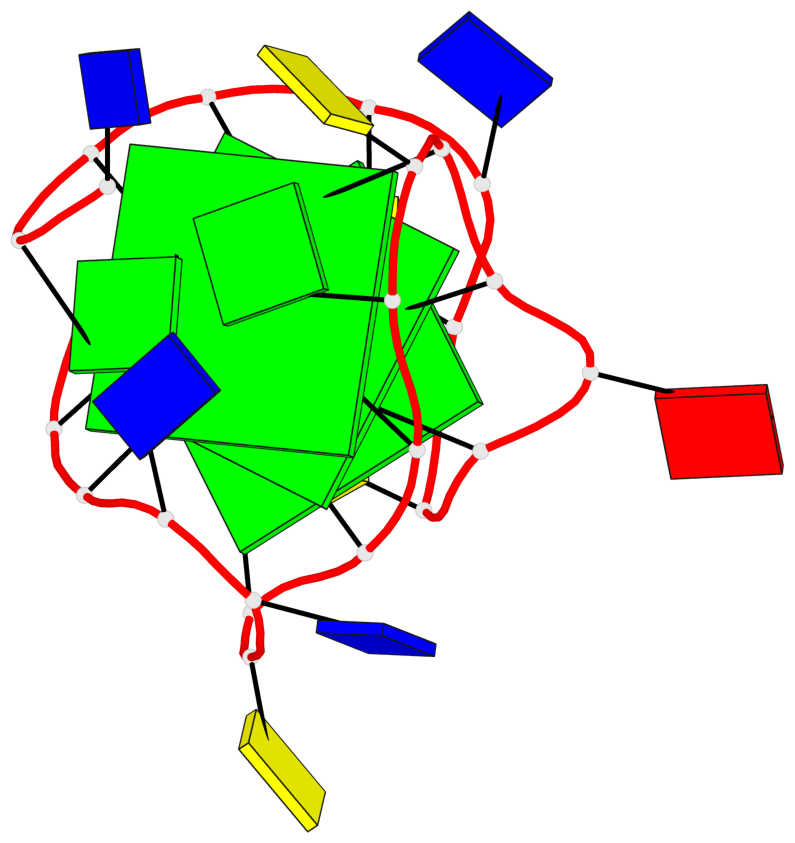
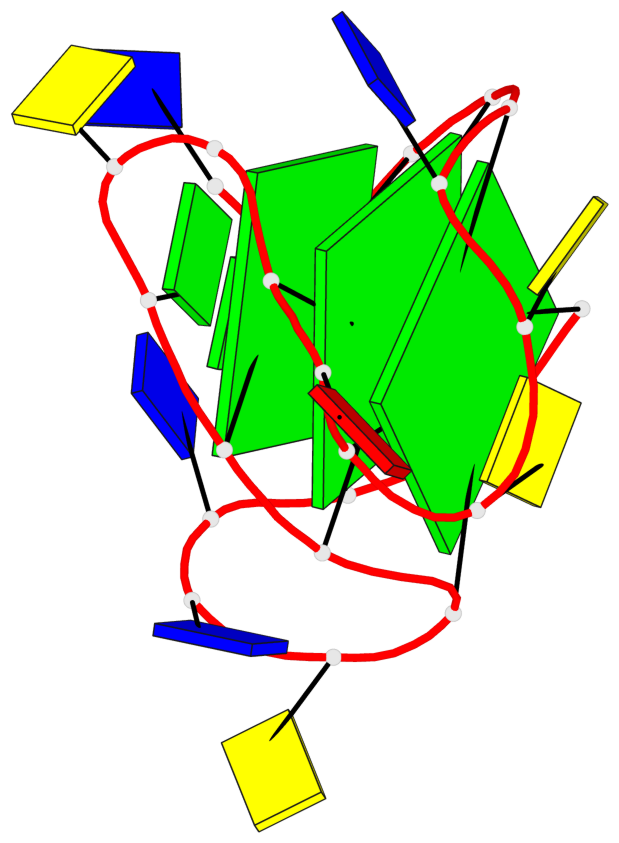
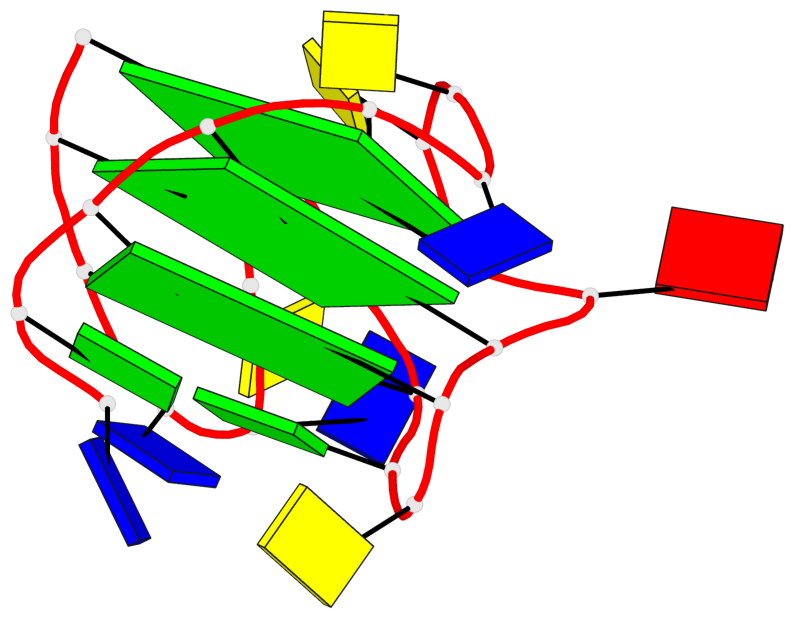
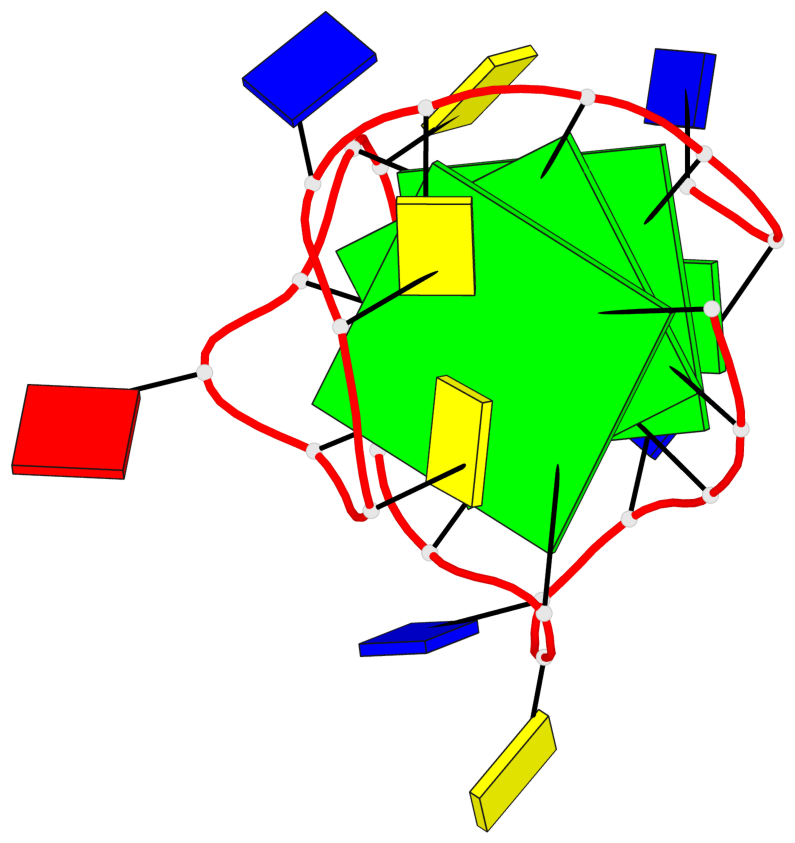
- 3 G-tetrads, 1 G4 helix, 1 G4 stem, 3(-Lw-Ln-P), hybrid-2(3+1), UDUU
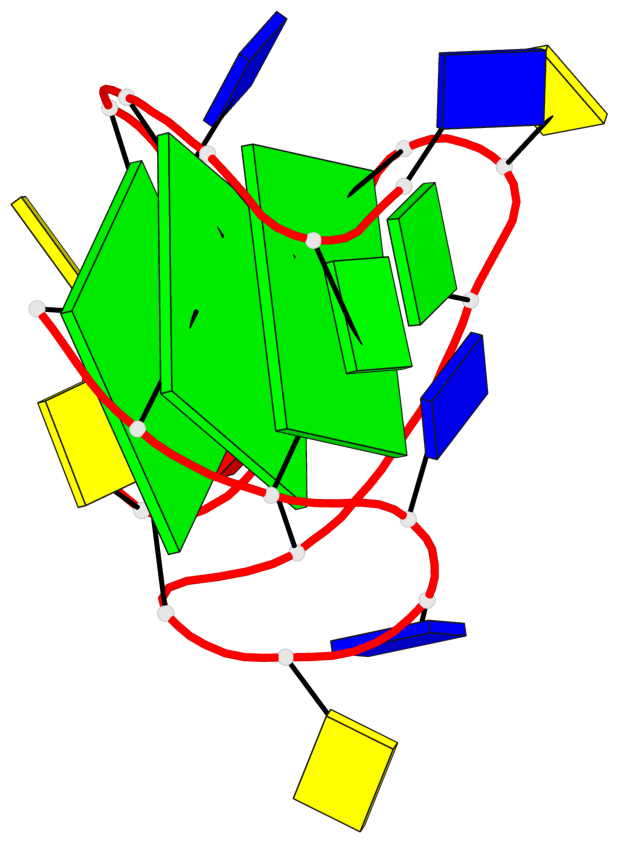
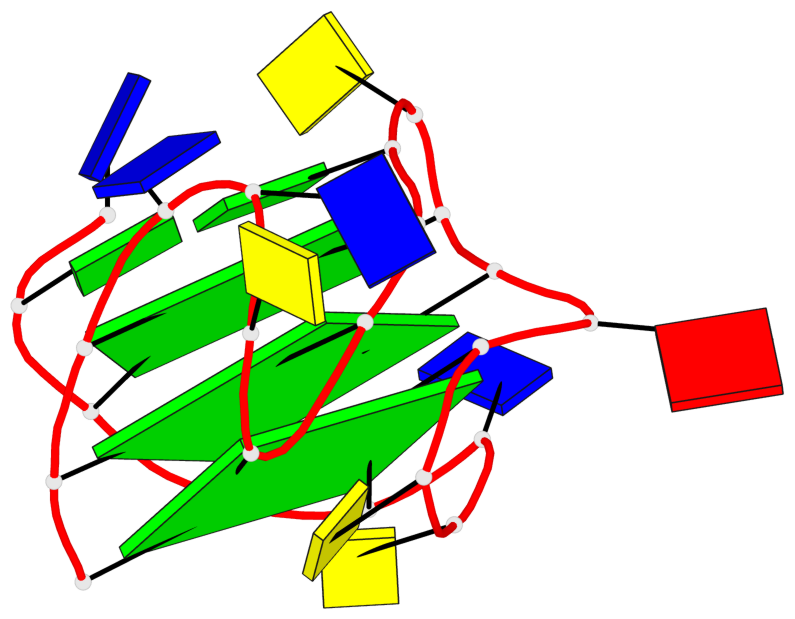
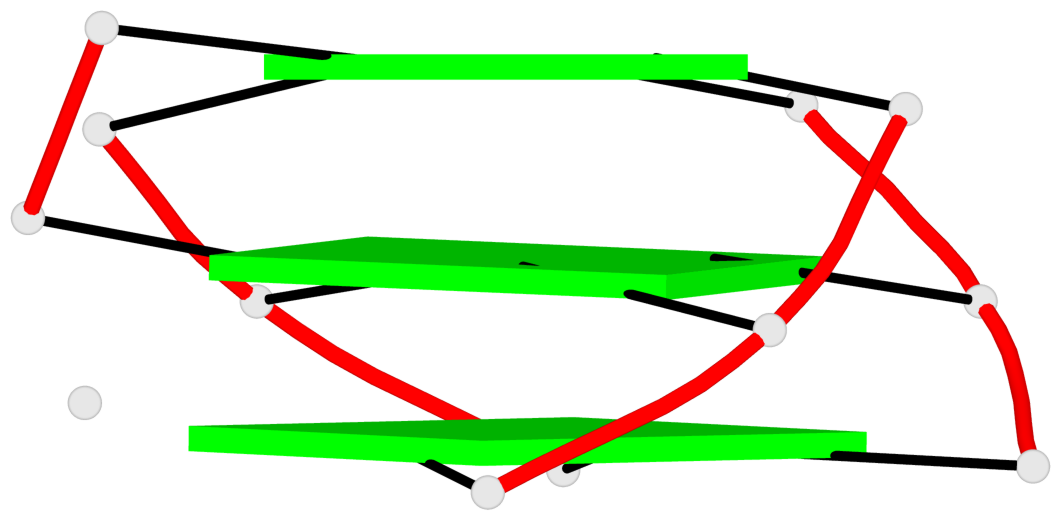
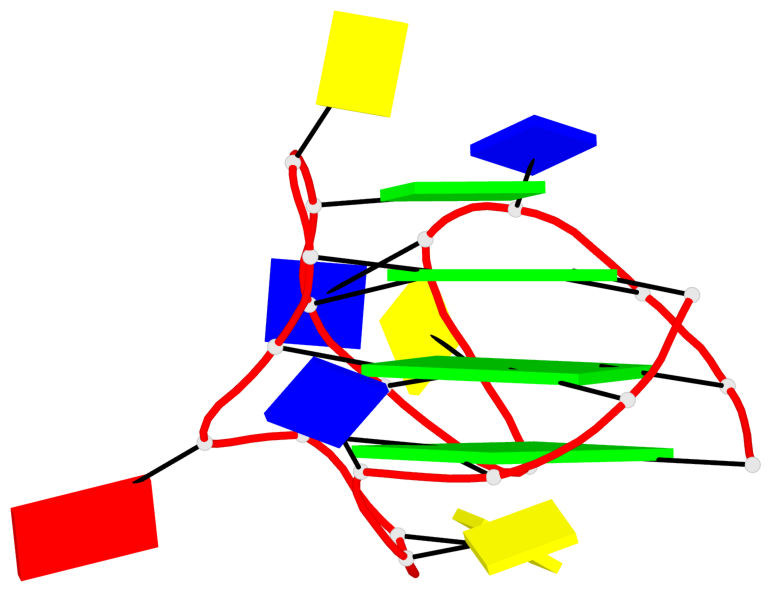
Base-block schematics in six views
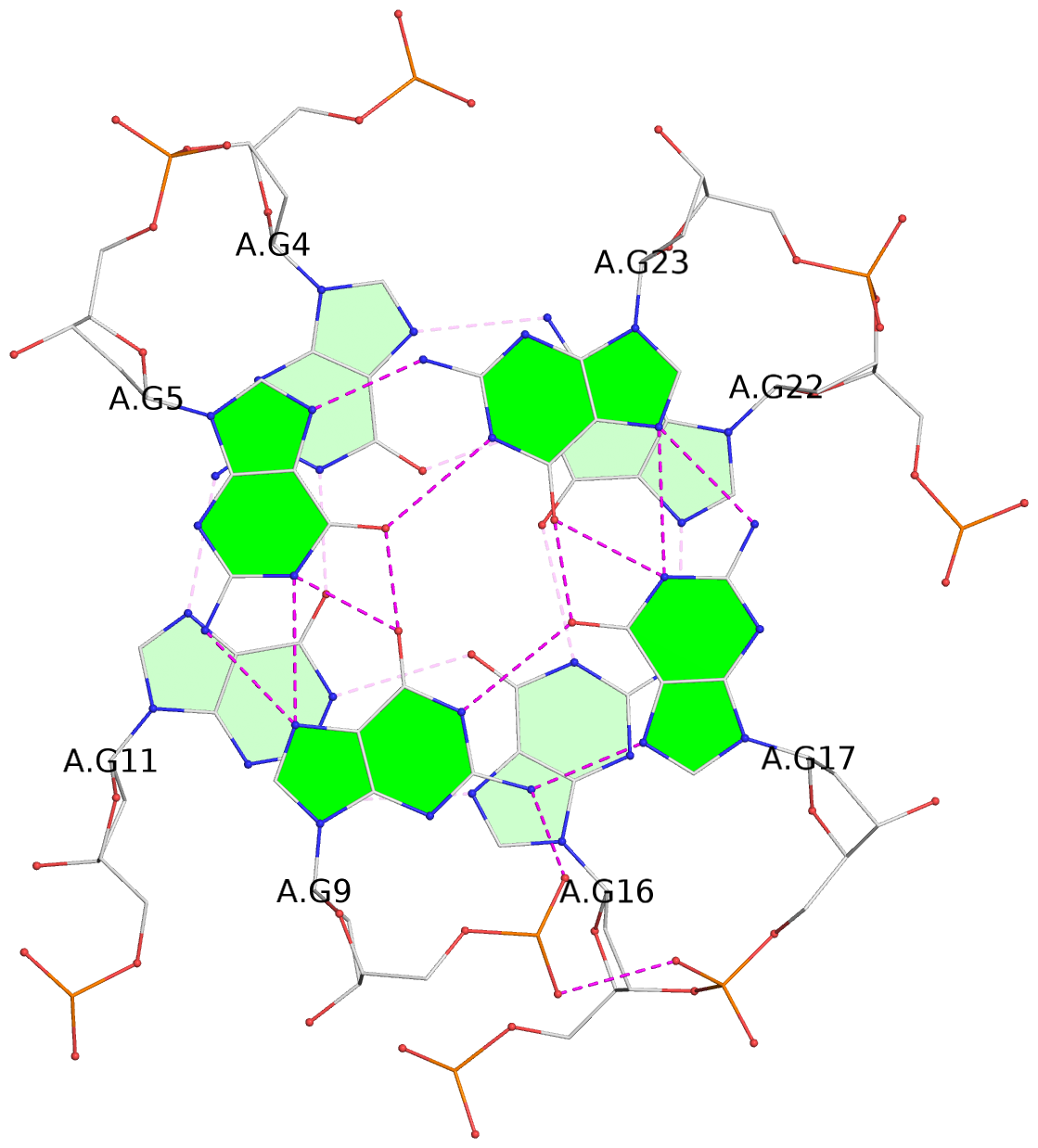
List of 3 G-tetrads
1 glyco-bond=s-ss sugar=---- groove=wn-- planarity=0.137 type=planar nts=4 GGGG A.DG3,A.DG12,A.DG15,A.DG21 2 glyco-bond=-s-- sugar=---- groove=wn-- planarity=0.149 type=planar nts=4 GGGG A.DG4,A.DG11,A.DG16,A.DG22 3 glyco-bond=-s-- sugar=---- groove=wn-- planarity=0.093 type=planar nts=4 GGGG A.DG5,A.DG9,A.DG17,A.DG23
List of 1 G4-helix
In DSSR, a G4-helix is defined by stacking interactions of G-tetrads, regardless of backbone connectivity, and may contain more than one G4-stem.
Helix#1, 3 G-tetrad layers, INTRA-molecular, with 1 stem
List of 1 G4-stem
In DSSR, a G4-stem is defined as a G4-helix with backbone connectivity. Bulges are also allowed along each of the four strands.