Detailed DSSR results for the G-quadruplex: PDB entry 6c65
Created and maintained by Xiang-Jun Lu <xiangjun@x3dna.org>
Citation: Please cite the NAR'20 DSSR-PyMOL schematics paper and/or the NAR'15 DSSR method paper.
Summary information
- PDB id
- 6c65
- Class
- RNA
- Method
- X-ray (2.8 Å)
- Summary
- Crystal structure of the mango-ii-a22u fluorescent aptamer bound to to1-biotin
- Reference
- Trachman 3rd RJ, Abdolahzadeh A, Andreoni A, Cojocaru R, Knutson JR, Ryckelynck M, Unrau PJ, Ferre-D'Amare AR (2018): "Crystal Structures of the Mango-II RNA Aptamer Reveal Heterogeneous Fluorophore Binding and Guide Engineering of Variants with Improved Selectivity and Brightness." Biochemistry, 57, 3544-3548. doi: 10.1021/acs.biochem.8b00399.
- Abstract
- Several RNA aptamers that bind small molecules and enhance their fluorescence have been successfully used to tag and track RNAs in vivo, but these genetically encodable tags have not yet achieved single-fluorophore resolution. Recently, Mango-II, an RNA that binds TO1-Biotin with ∼1 nM affinity and enhances its fluorescence by >1500-fold, was isolated by fluorescence selection from the pool that yielded the original RNA Mango. We determined the crystal structures of Mango-II in complex with two fluorophores, TO1-Biotin and TO3-Biotin, and found that despite their high affinity, the ligands adopt multiple distinct conformations, indicative of a binding pocket with modest stereoselectivity. Mutational analysis of the binding site led to Mango-II(A22U), which retains high affinity for TO1-Biotin but now discriminates >5-fold against TO3-biotin. Moreover, fluorescence enhancement of TO1-Biotin increases by 18%, while that of TO3-Biotin decreases by 25%. Crystallographic, spectroscopic, and analogue studies show that the A22U mutation improves conformational homogeneity and shape complementarity of the fluorophore-RNA interface. Our work demonstrates that even after extensive functional selection, aptamer RNAs can be further improved through structure-guided engineering.
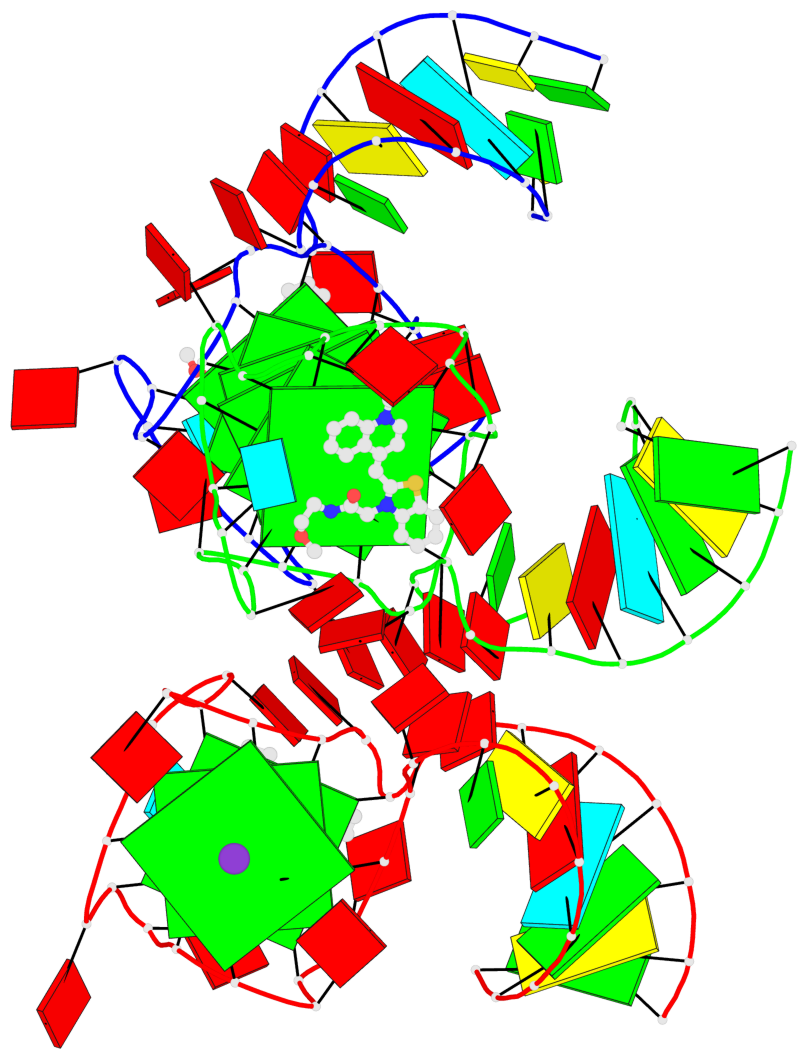
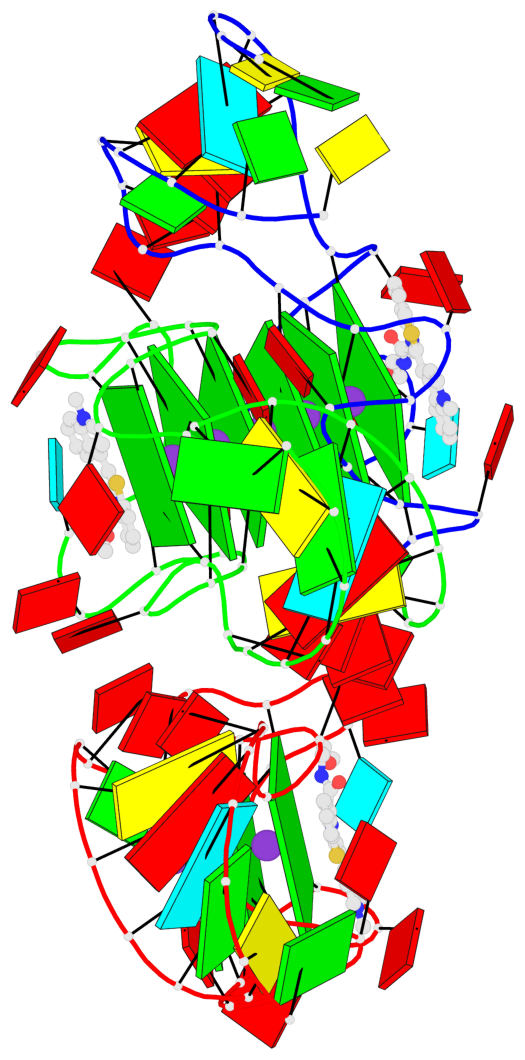
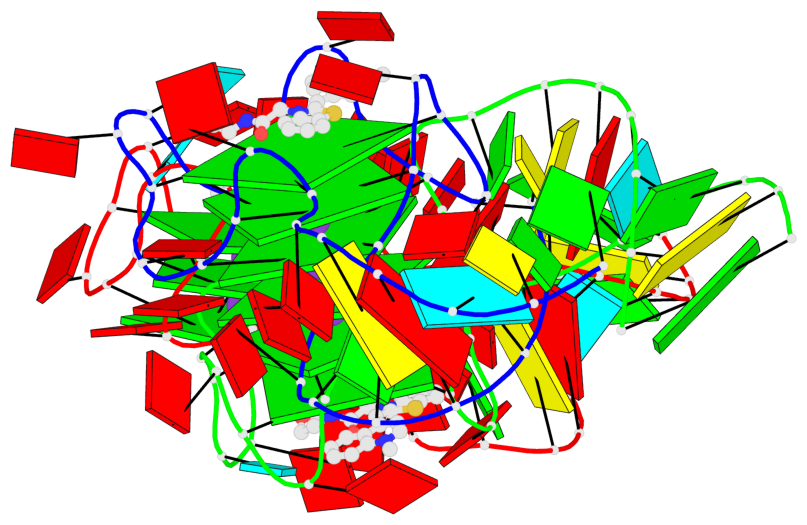
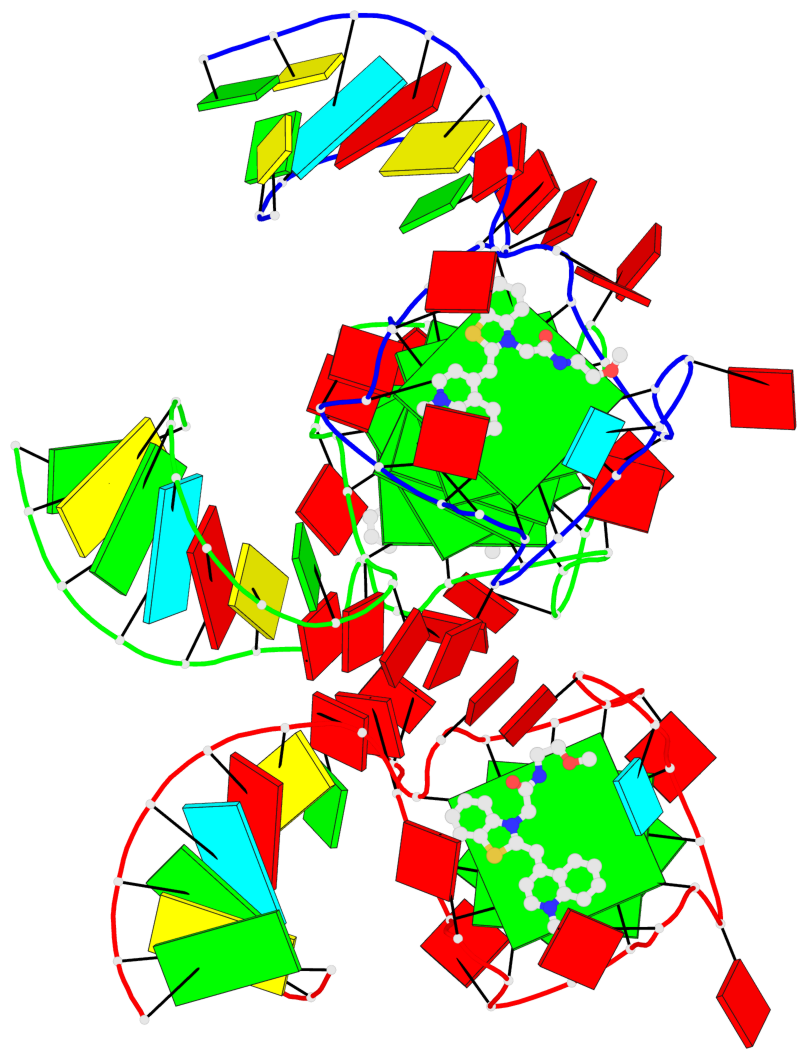
- G4 notes
- 9 G-tetrads, 2 G4 helices, 3 G4 stems, 1 G4 coaxial stack, 2(-P-P-P), parallel(4+0), UUUU, coaxial interfaces: 5'/5'
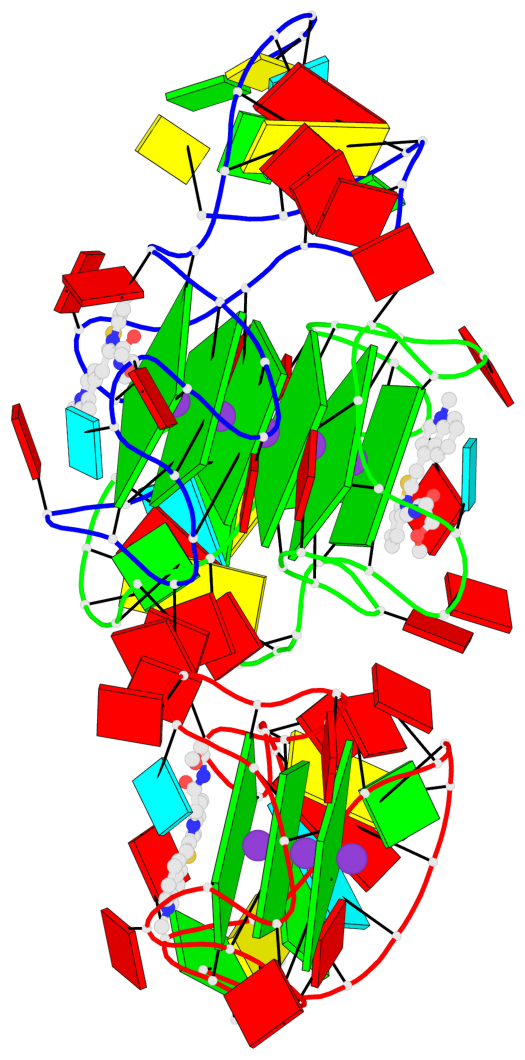
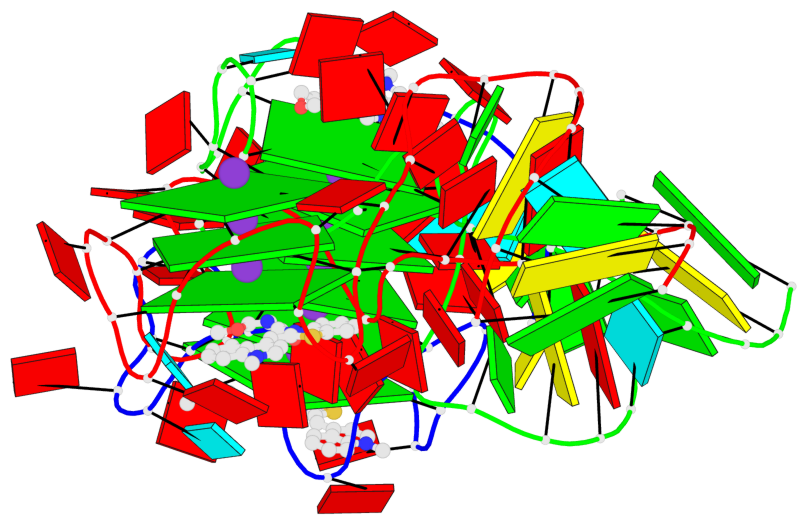
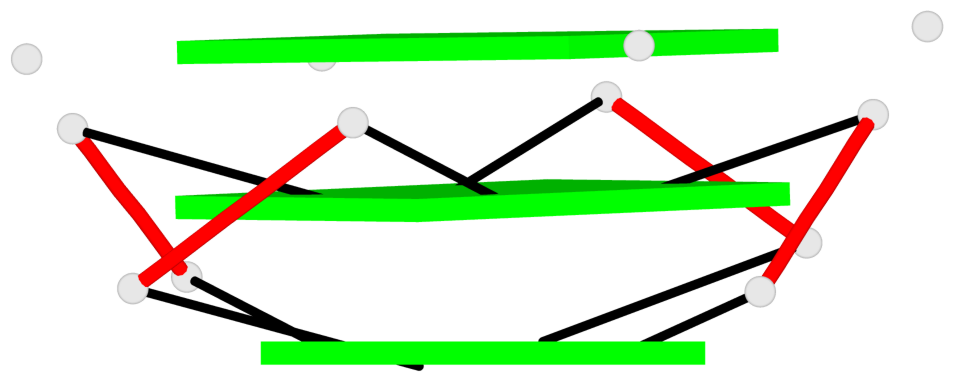
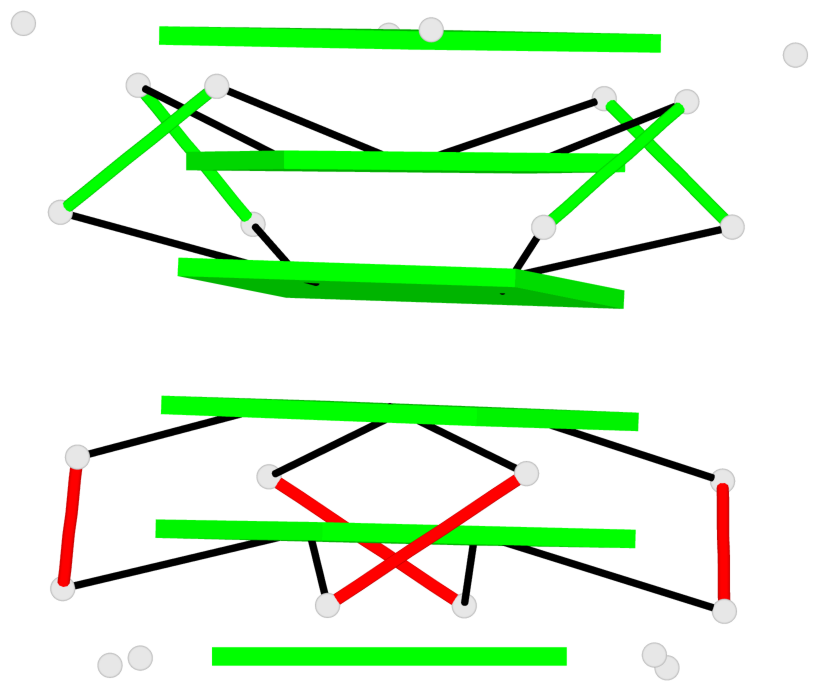
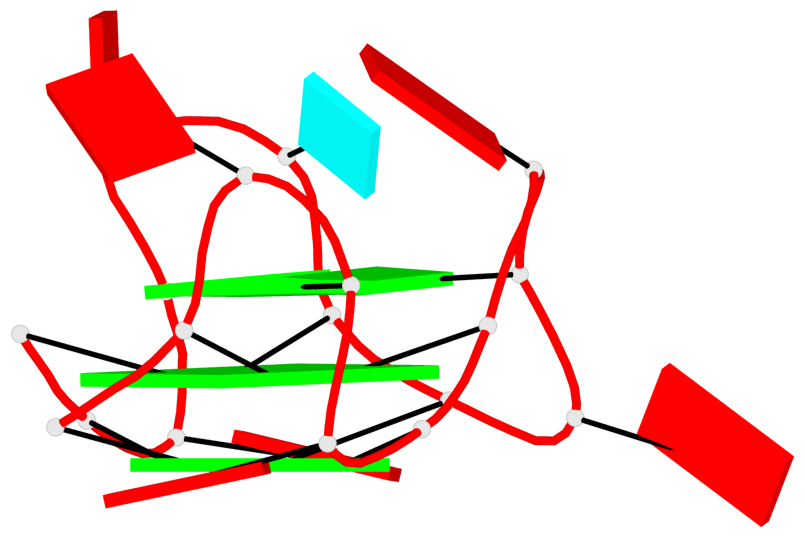
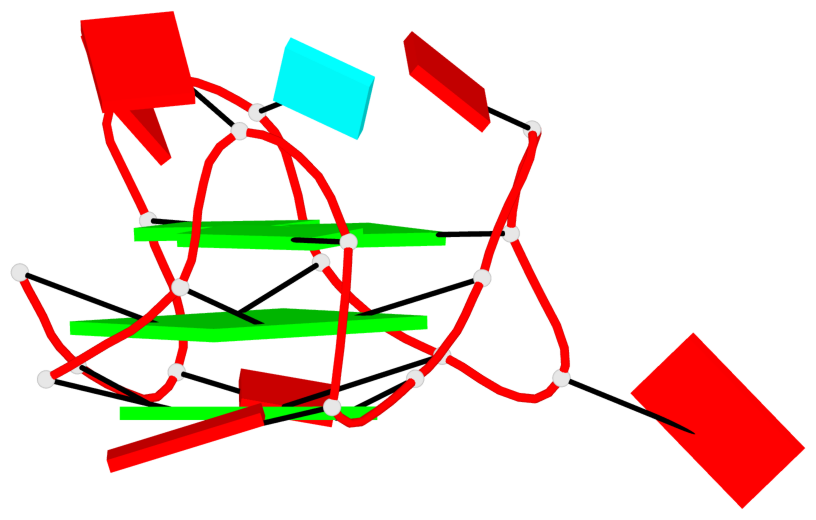
Base-block schematics in six views
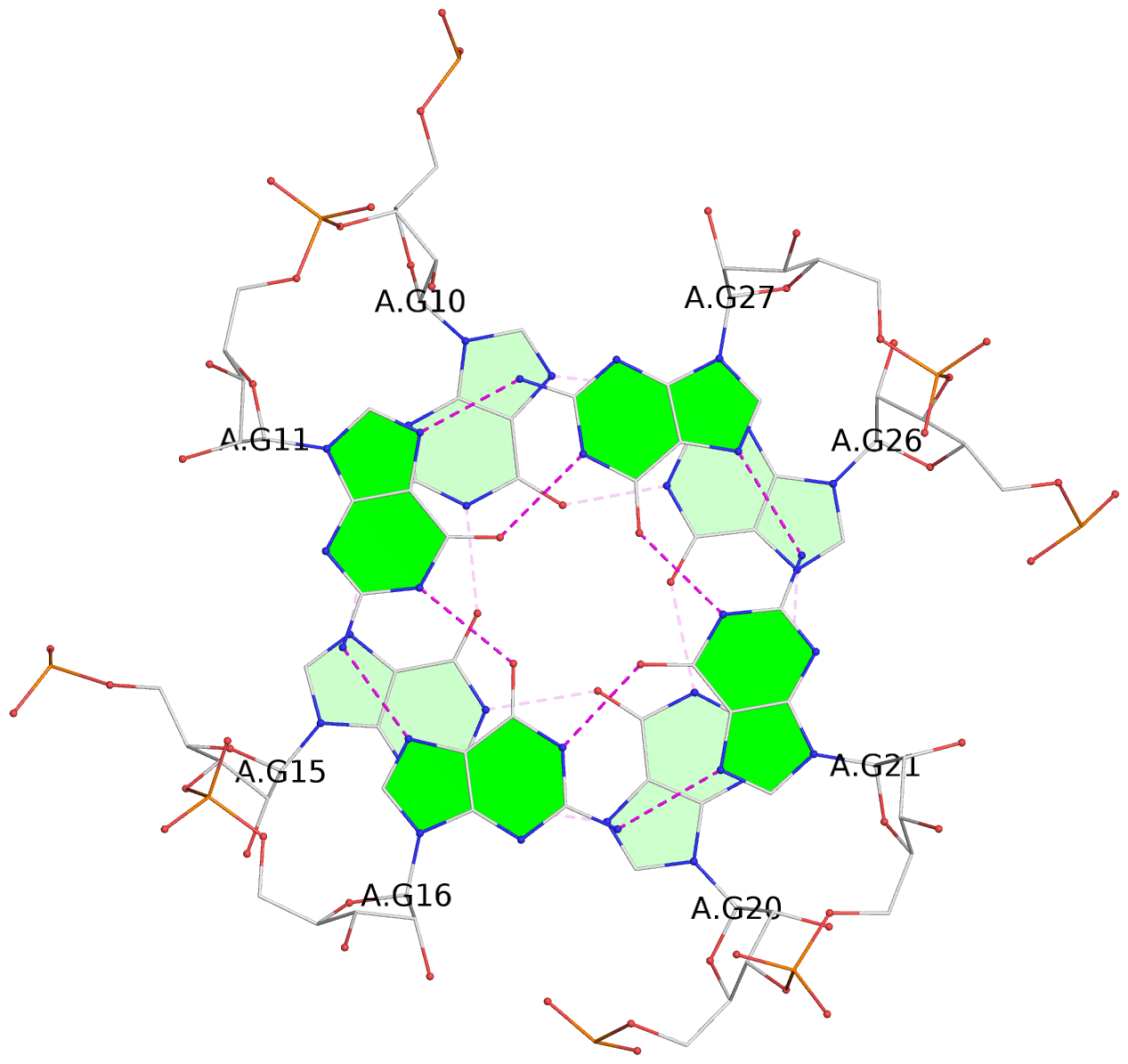
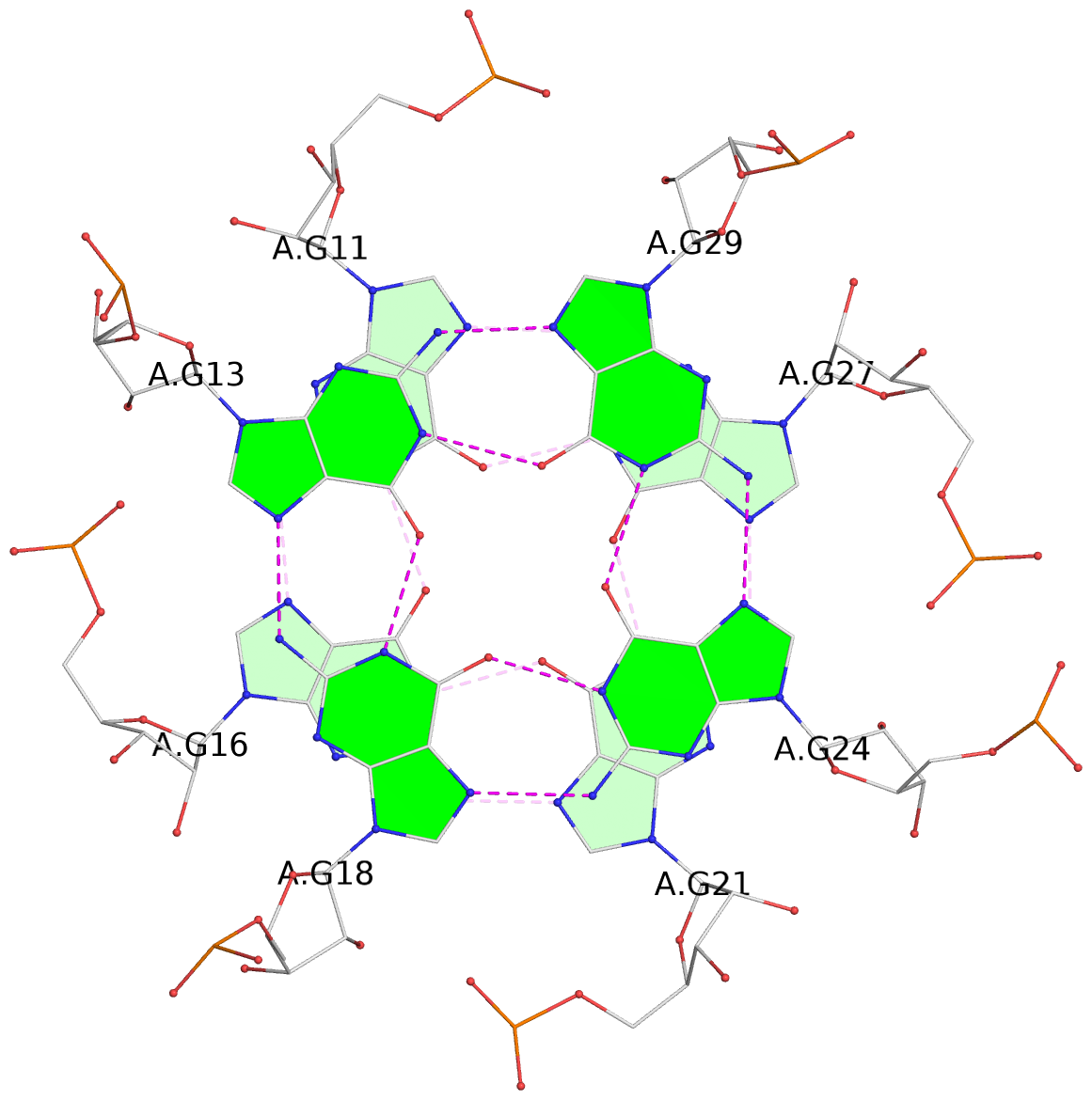
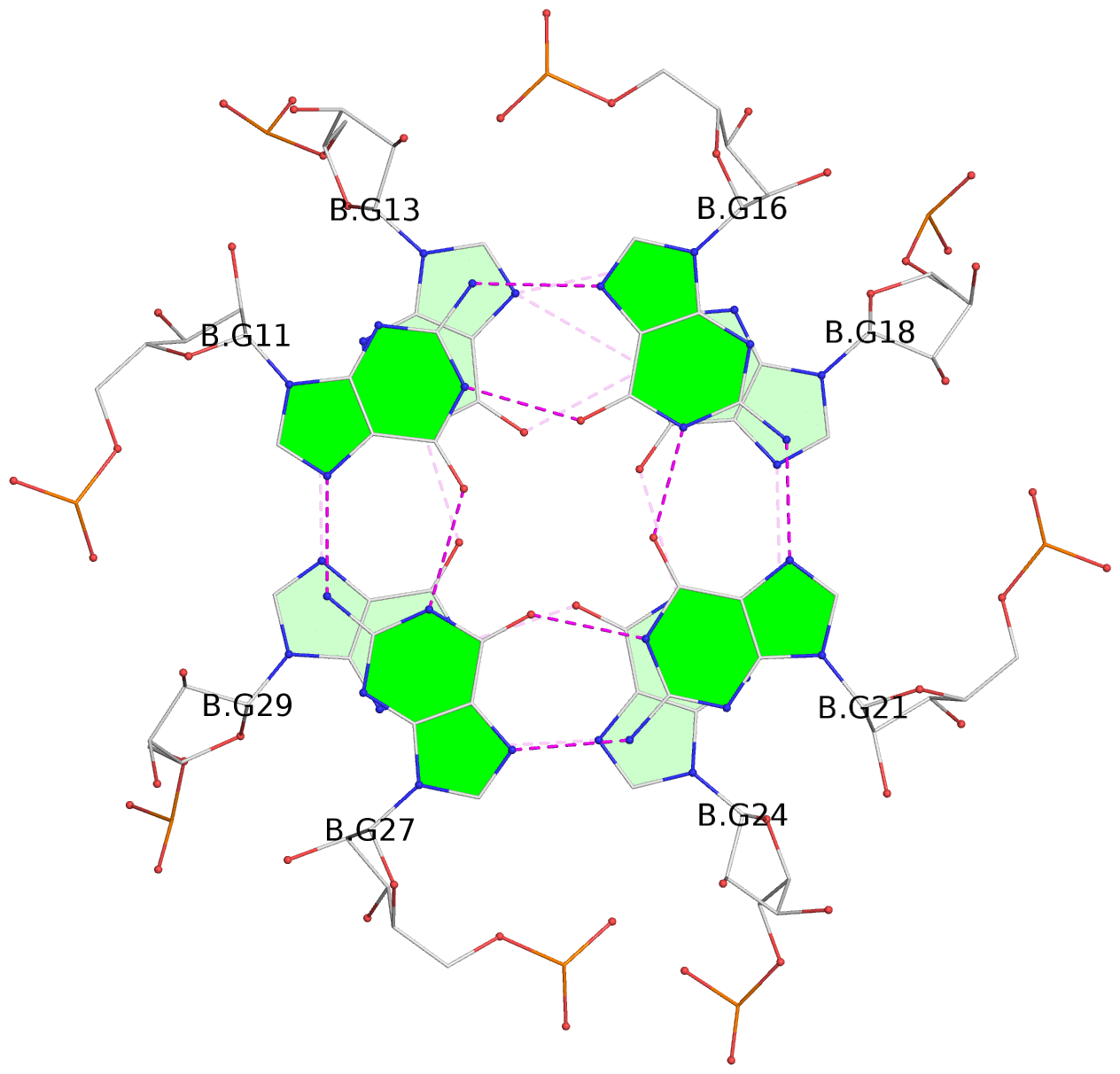
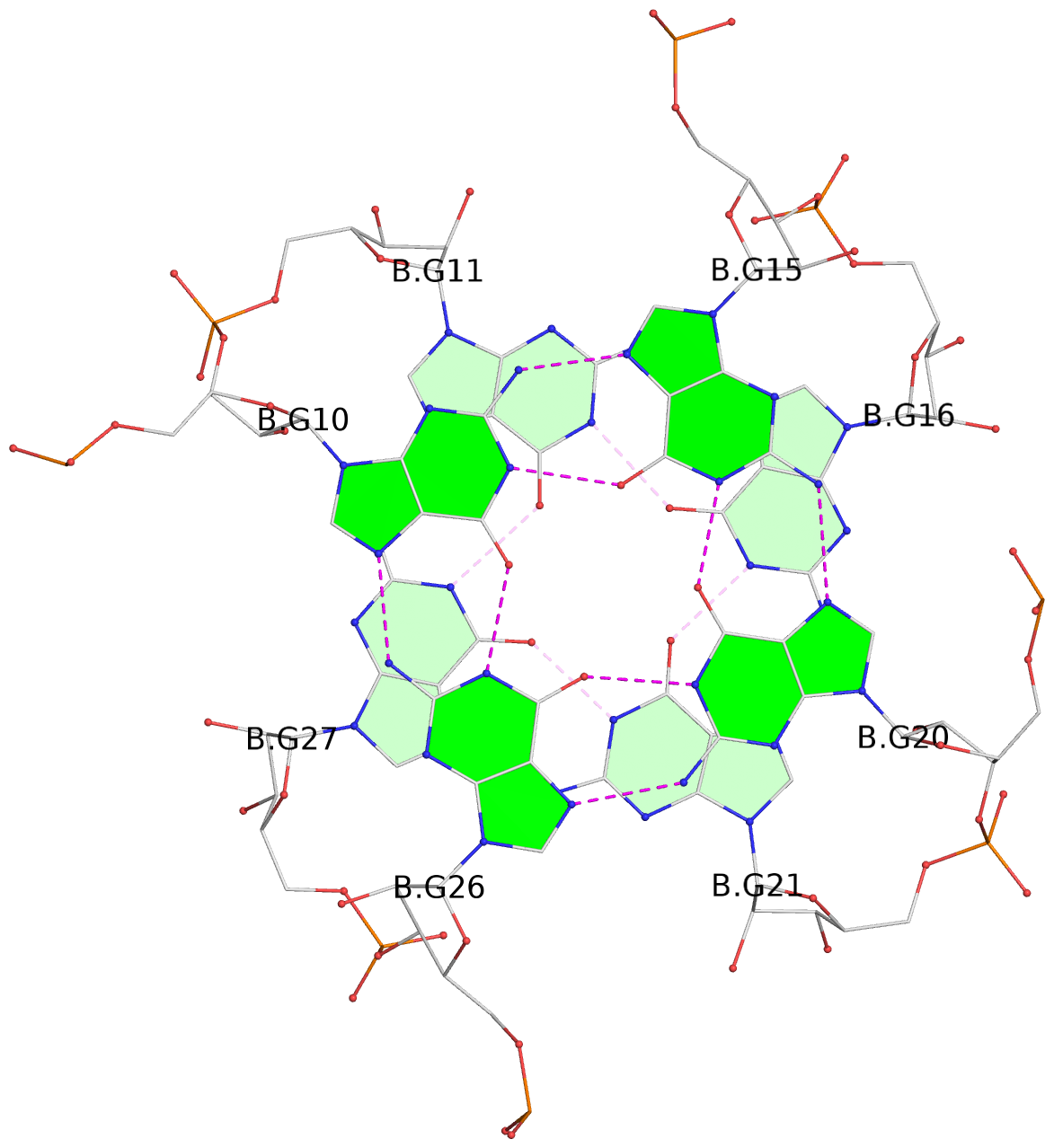
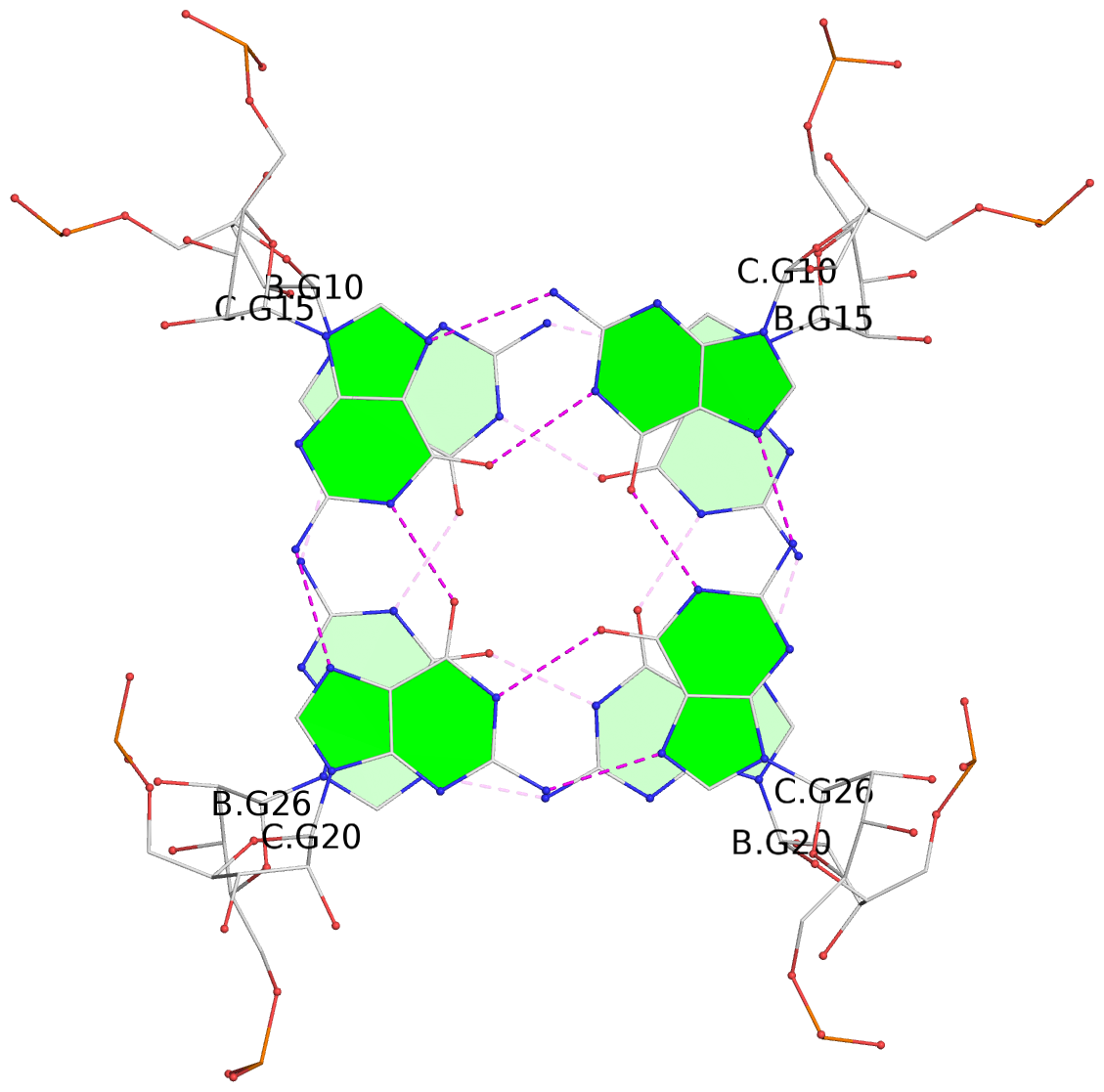
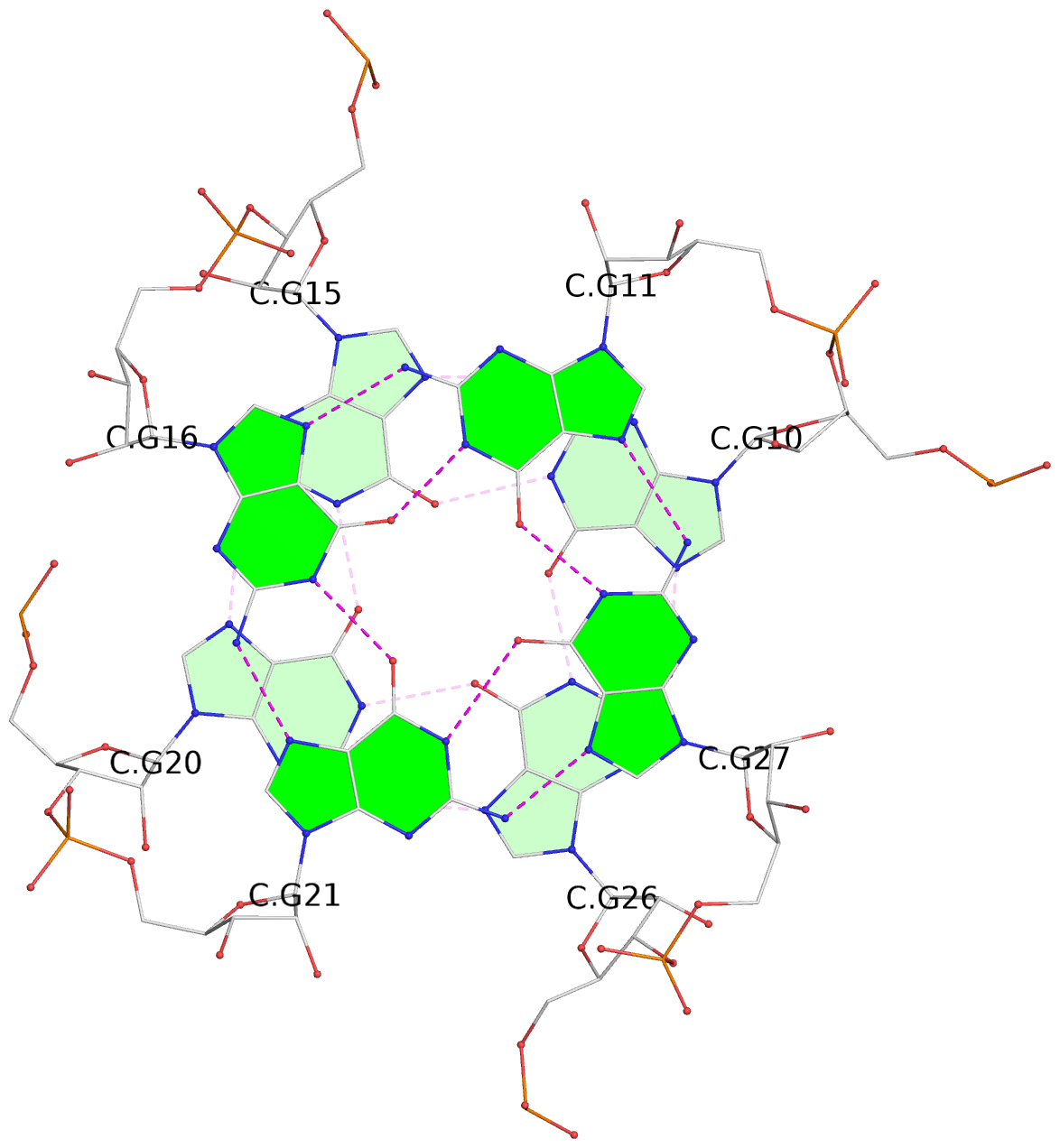
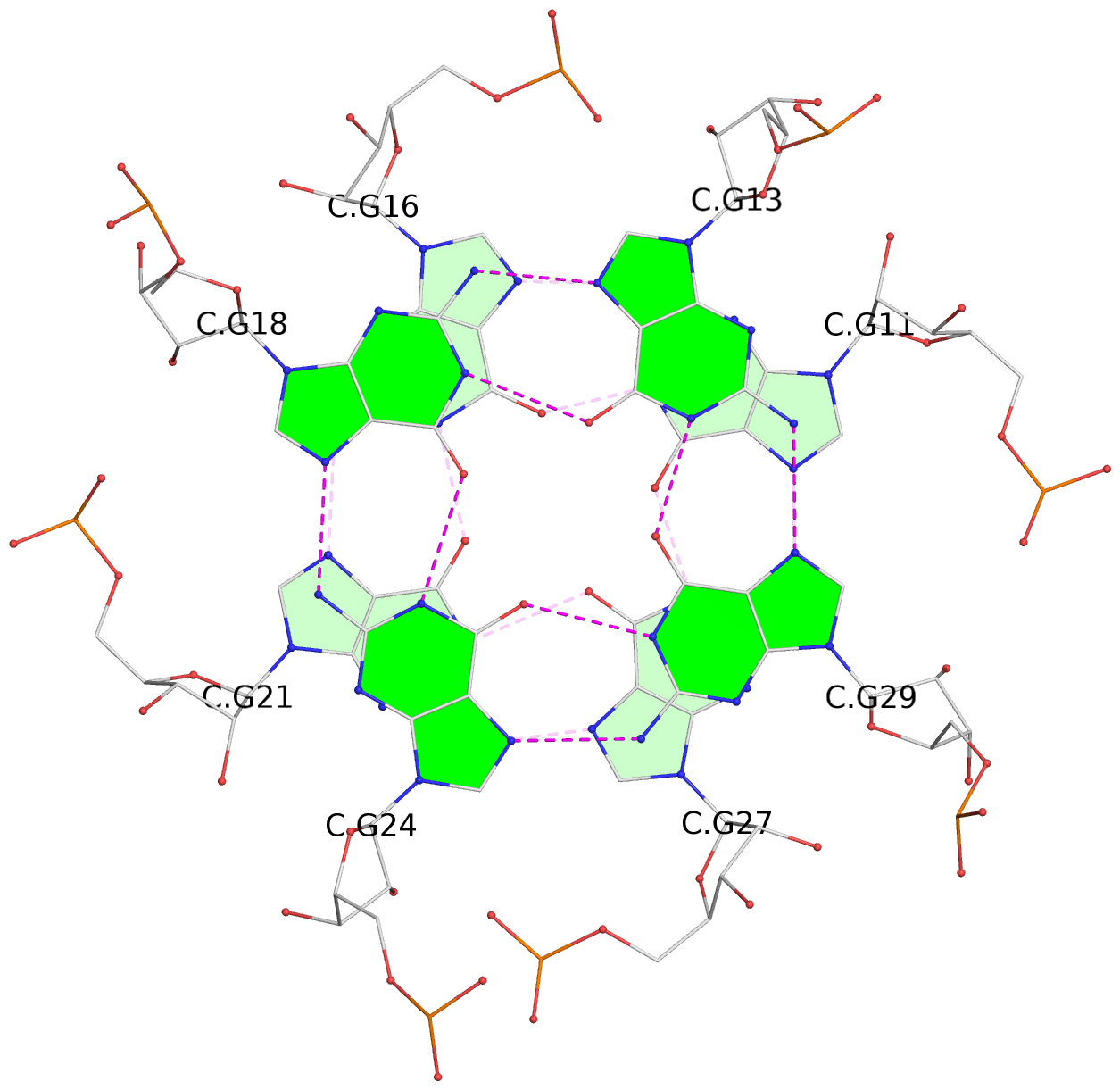
List of 9 G-tetrads
1 glyco-bond=---- sugar=-333 groove=---- planarity=0.242 type=other nts=4 GGGG A.G10,A.G15,A.G20,A.G26 2 glyco-bond=---- sugar=3333 groove=---- planarity=0.135 type=planar nts=4 GGGG A.G11,A.G16,A.G21,A.G27 3 glyco-bond=---- sugar=---- groove=---- planarity=0.106 type=planar nts=4 GGGG A.G13,A.G29,A.G24,A.G18 4 glyco-bond=---- sugar=-3-3 groove=---- planarity=0.321 type=other nts=4 GGGG B.G10,B.G15,B.G20,B.G26 5 glyco-bond=---- sugar=3333 groove=---- planarity=0.083 type=planar nts=4 GGGG B.G11,B.G16,B.G21,B.G27 6 glyco-bond=---- sugar=---- groove=---- planarity=0.070 type=planar nts=4 GGGG B.G13,B.G29,B.G24,B.G18 7 glyco-bond=---- sugar=-333 groove=---- planarity=0.283 type=other nts=4 GGGG C.G10,C.G15,C.G20,C.G26 8 glyco-bond=---- sugar=3333 groove=---- planarity=0.106 type=planar nts=4 GGGG C.G11,C.G16,C.G21,C.G27 9 glyco-bond=---- sugar=---- groove=---- planarity=0.109 type=planar nts=4 GGGG C.G13,C.G29,C.G24,C.G18
List of 2 G4-helices
In DSSR, a G4-helix is defined by stacking interactions of G-tetrads, regardless of backbone connectivity, and may contain more than one G4-stem.
Helix#1, 3 G-tetrad layers, INTRA-molecular, with 1 stem
Helix#2, 6 G-tetrad layers, inter-molecular, with 2 stems
List of 3 G4-stems
In DSSR, a G4-stem is defined as a G4-helix with backbone connectivity. Bulges are also allowed along each of the four strands.
Stem#1, 2 G-tetrad layers, 3 loops, INTRA-molecular, UUUU, parallel, 2(-P-P-P), parallel(4+0)
Stem#2, 2 G-tetrad layers, 3 loops, INTRA-molecular, UUUU, parallel, 2(-P-P-P), parallel(4+0)
Stem#3, 2 G-tetrad layers, 3 loops, INTRA-molecular, UUUU, parallel, 2(-P-P-P), parallel(4+0)
List of 1 G4 coaxial stack
1 G4 helix#2 contains 2 G4 stems: [#2,#3] [5'/5']