Detailed DSSR results for the G-quadruplex: PDB entry 6e8t
Created and maintained by Xiang-Jun Lu <xiangjun@x3dna.org>
Citation: Please cite the NAR'20 DSSR-PyMOL schematics paper and/or the NAR'15 DSSR method paper.
Summary information
- PDB id
- 6e8t
- Class
- RNA
- Method
- X-ray (2.9 Å)
- Summary
- Structure of the mango-iii (a10u) aptamer bound to to1-biotin
- Reference
- Trachman 3rd RJ, Autour A, Jeng SCY, Abdolahzadeh A, Andreoni A, Cojocaru R, Garipov R, Dolgosheina EV, Knutson JR, Ryckelynck M, Unrau PJ, Ferre-D'Amare AR (2019): "Structure and functional reselection of the Mango-III fluorogenic RNA aptamer." Nat. Chem. Biol., 15, 472-479. doi: 10.1038/s41589-019-0267-9.
- Abstract
- Several turn-on RNA aptamers that activate small-molecule fluorophores have been selected in vitro. Among these, the ~30 nucleotide Mango-III is notable because it binds the thiazole orange derivative TO1-Biotin with high affinity and fluoresces brightly (quantum yield 0.55). Uniquely among related aptamers, Mango-III exhibits biphasic thermal melting, characteristic of molecules with tertiary structure. We report crystal structures of TO1-Biotin complexes of Mango-III, a structure-guided mutant Mango-III(A10U), and a functionally reselected mutant iMango-III. The structures reveal a globular architecture arising from an unprecedented pseudoknot-like connectivity between a G-quadruplex and an embedded non-canonical duplex. The fluorophore is restrained into a planar conformation by the G-quadruplex, a lone, long-range trans Watson-Crick pair (whose A10U mutation increases quantum yield to 0.66), and a pyrimidine perpendicular to the nucleobase planes of those motifs. The improved iMango-III and Mango-III(A10U) fluoresce ~50% brighter than enhanced green fluorescent protein, making them suitable tags for live cell RNA visualization.
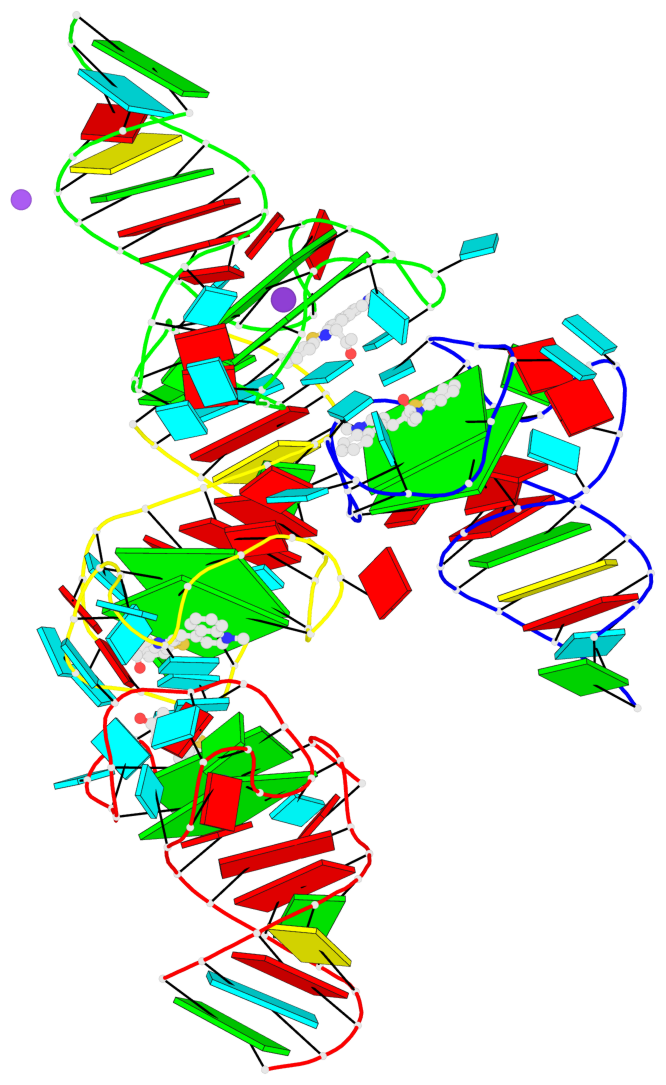
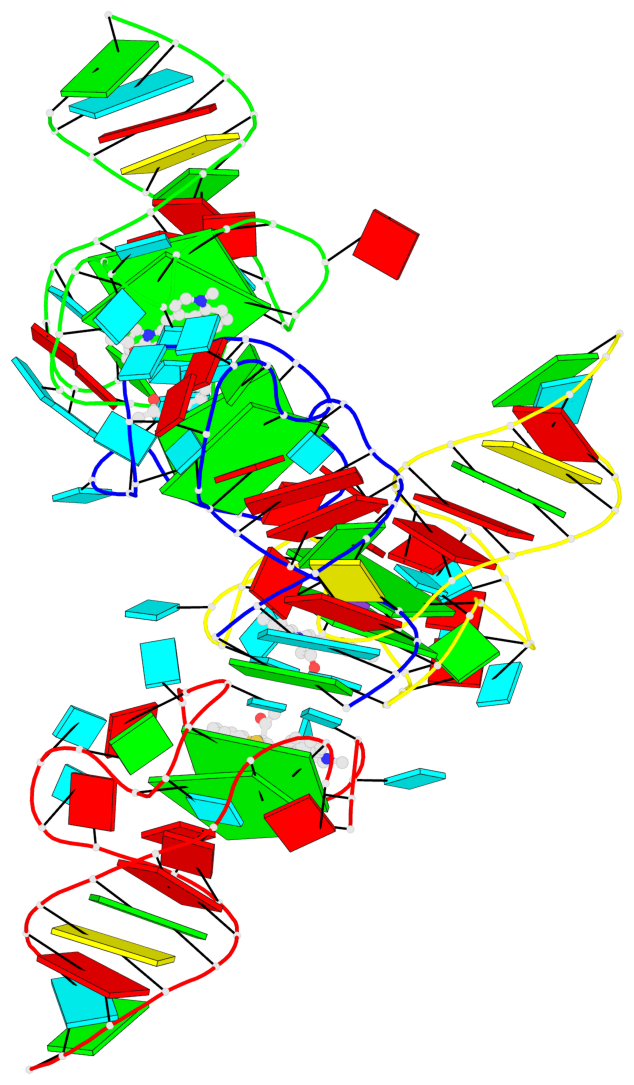
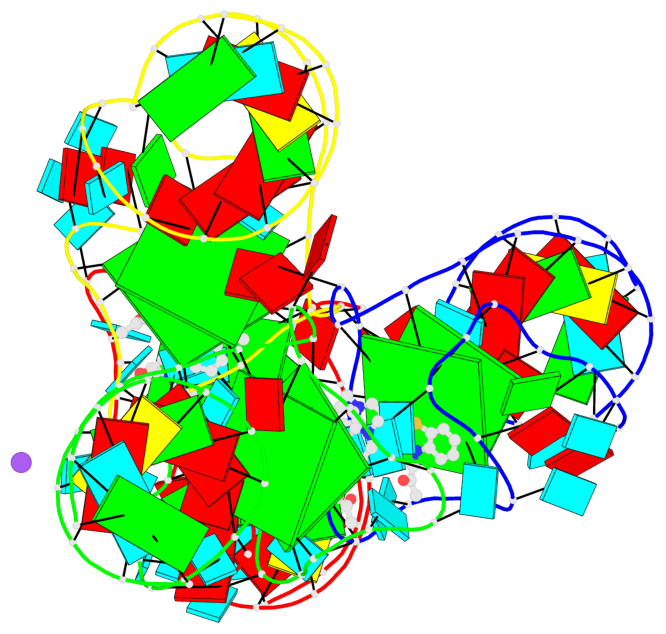
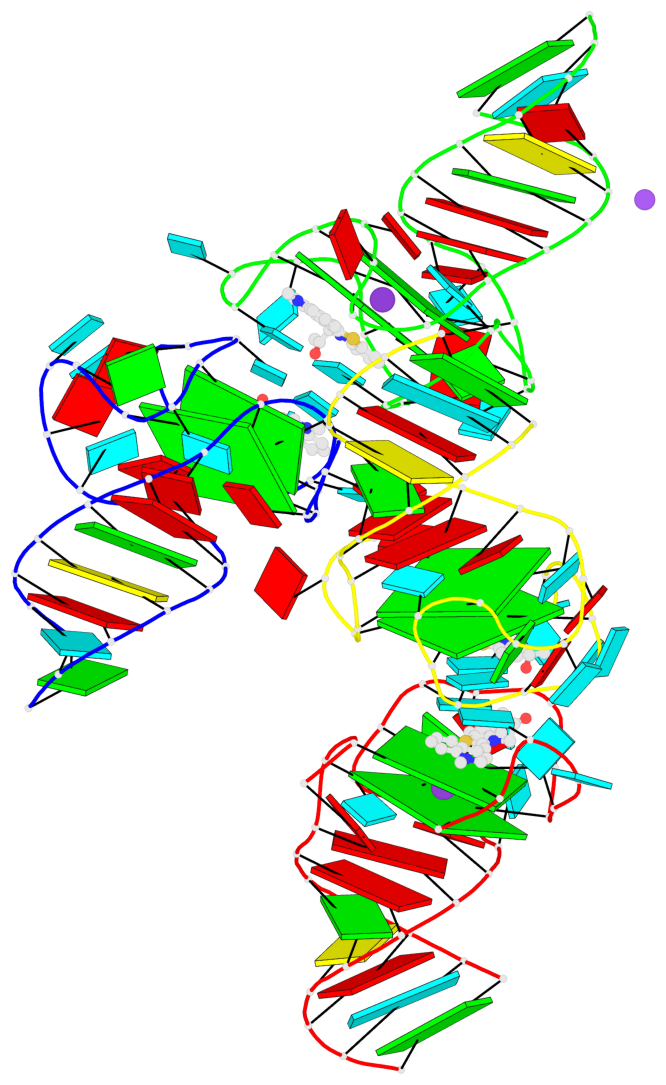
- G4 notes
- 8 G-tetrads, 4 G4 helices, 4 G4 stems, 2(-P-Lw), hybrid-1(3+1), UUUD; 2(-P-P-Lw), hybrid-2R(3+1), UUUD
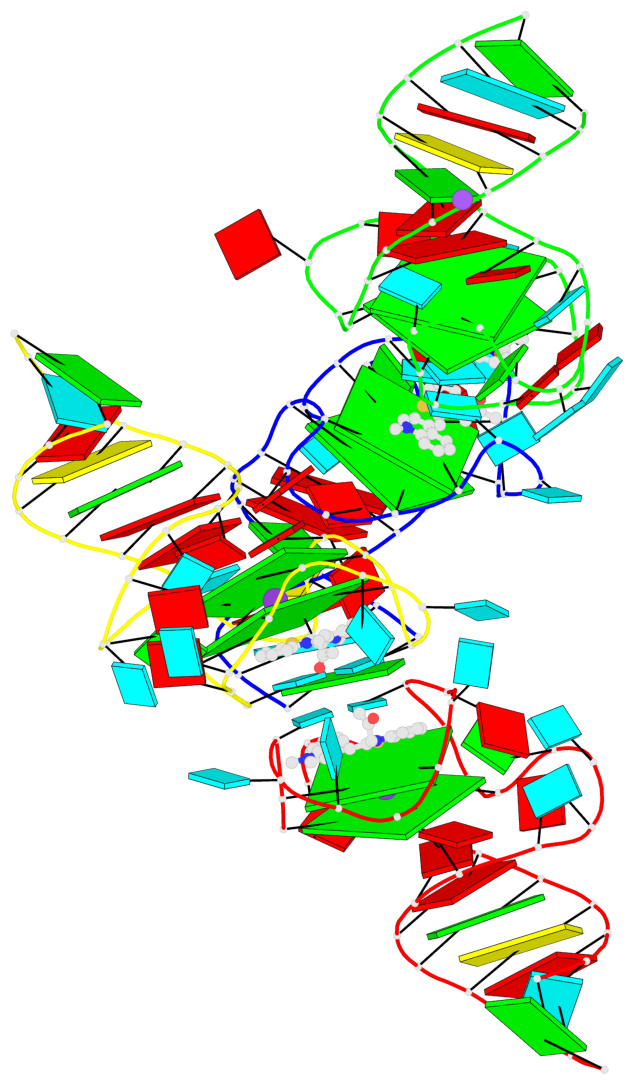
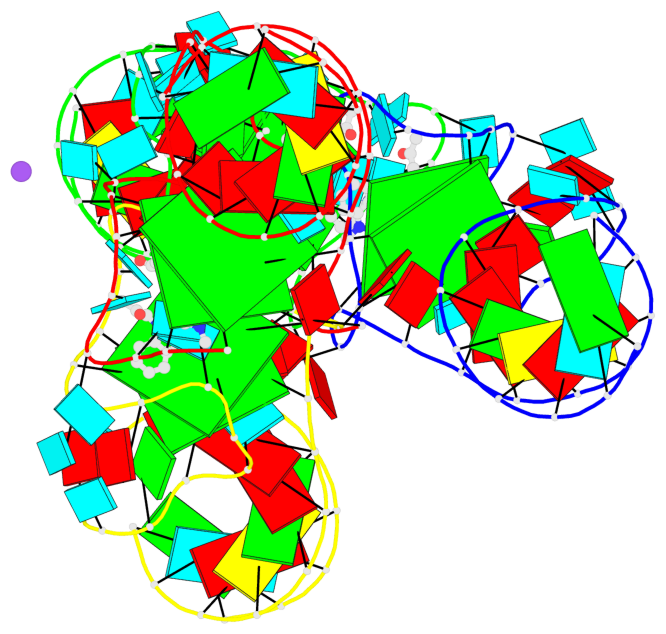
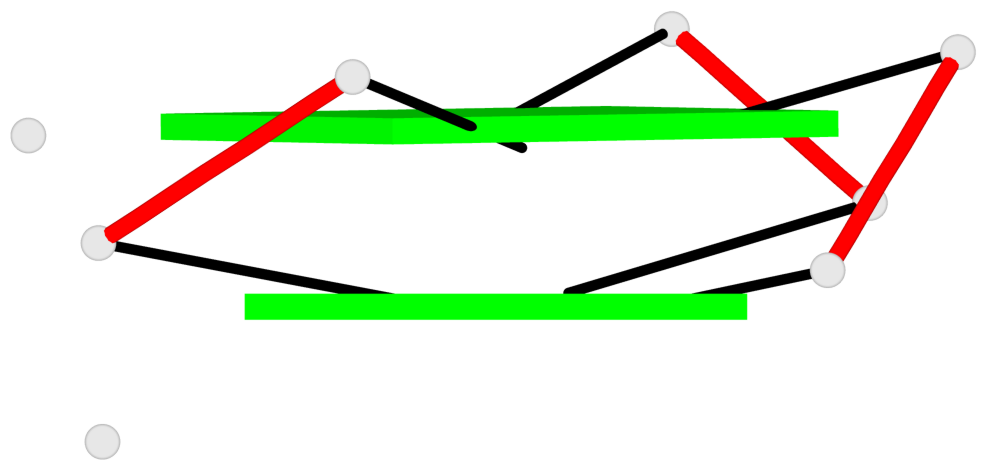
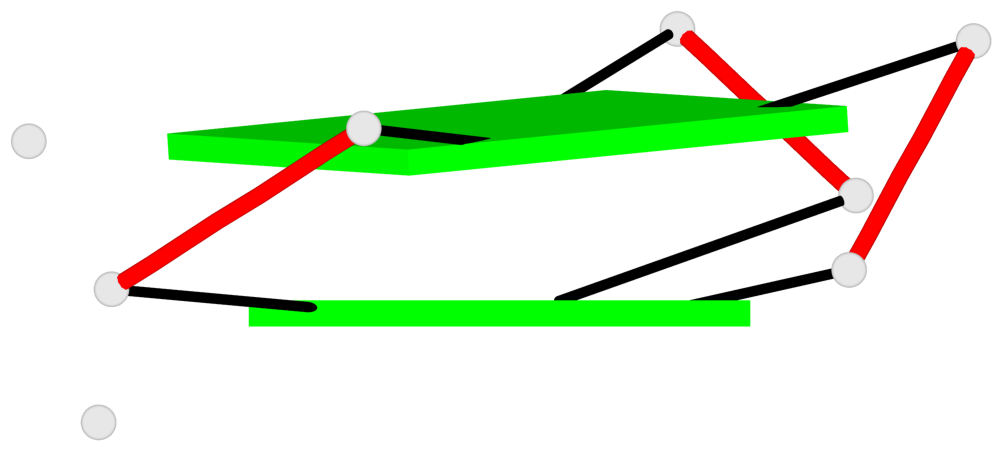
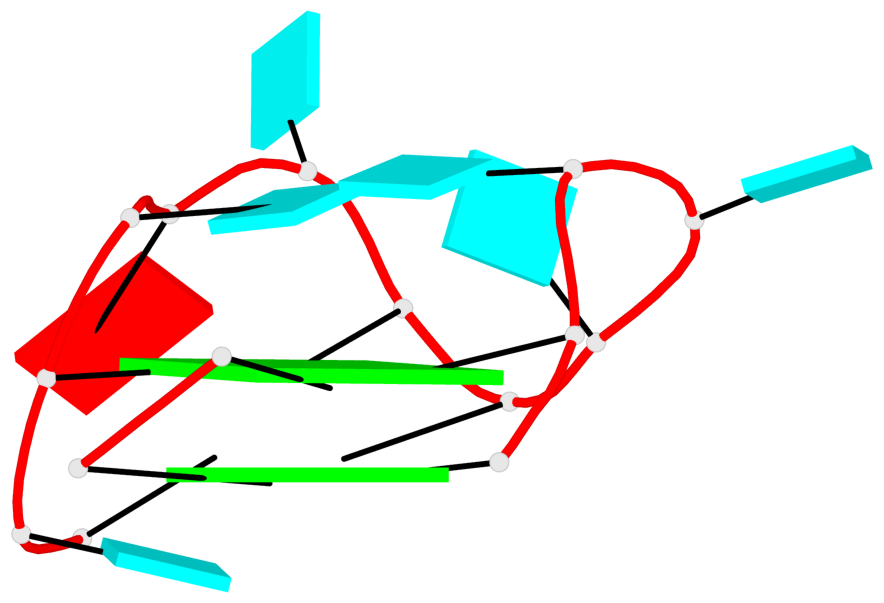
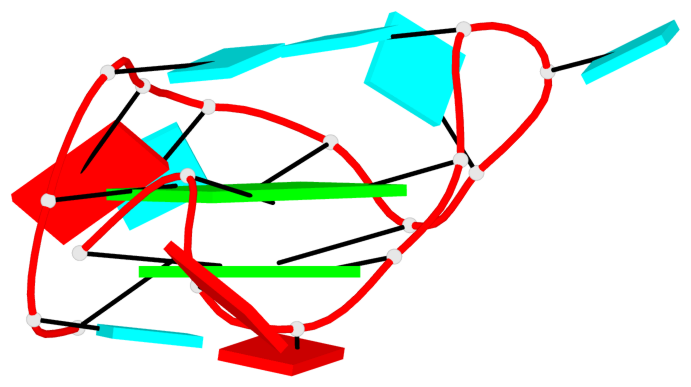
Base-block schematics in six views
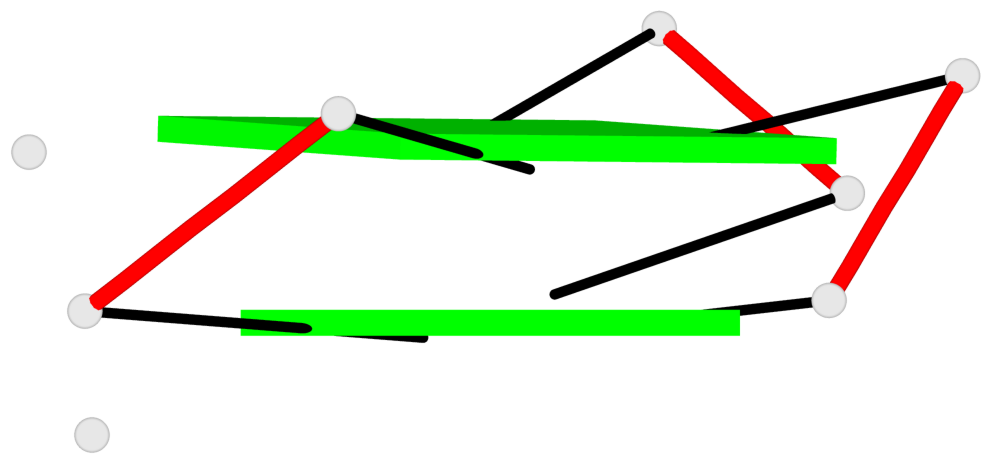
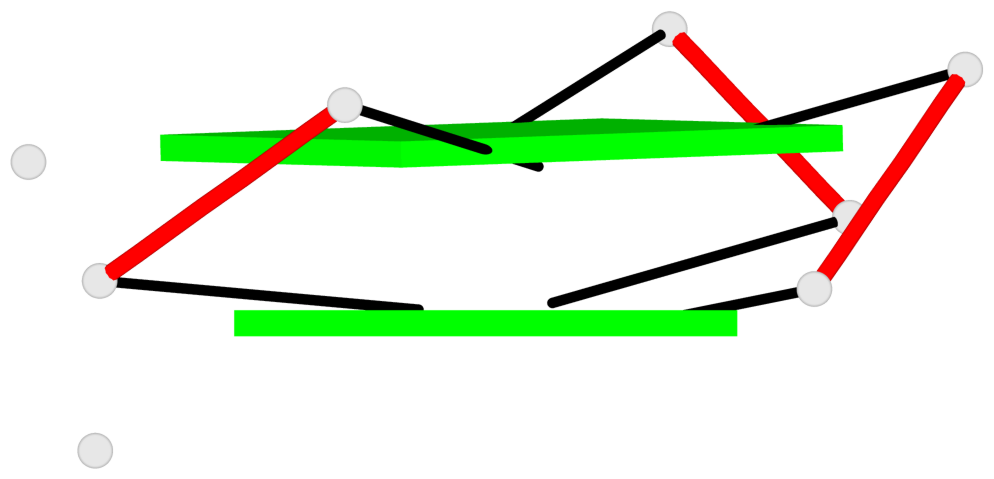
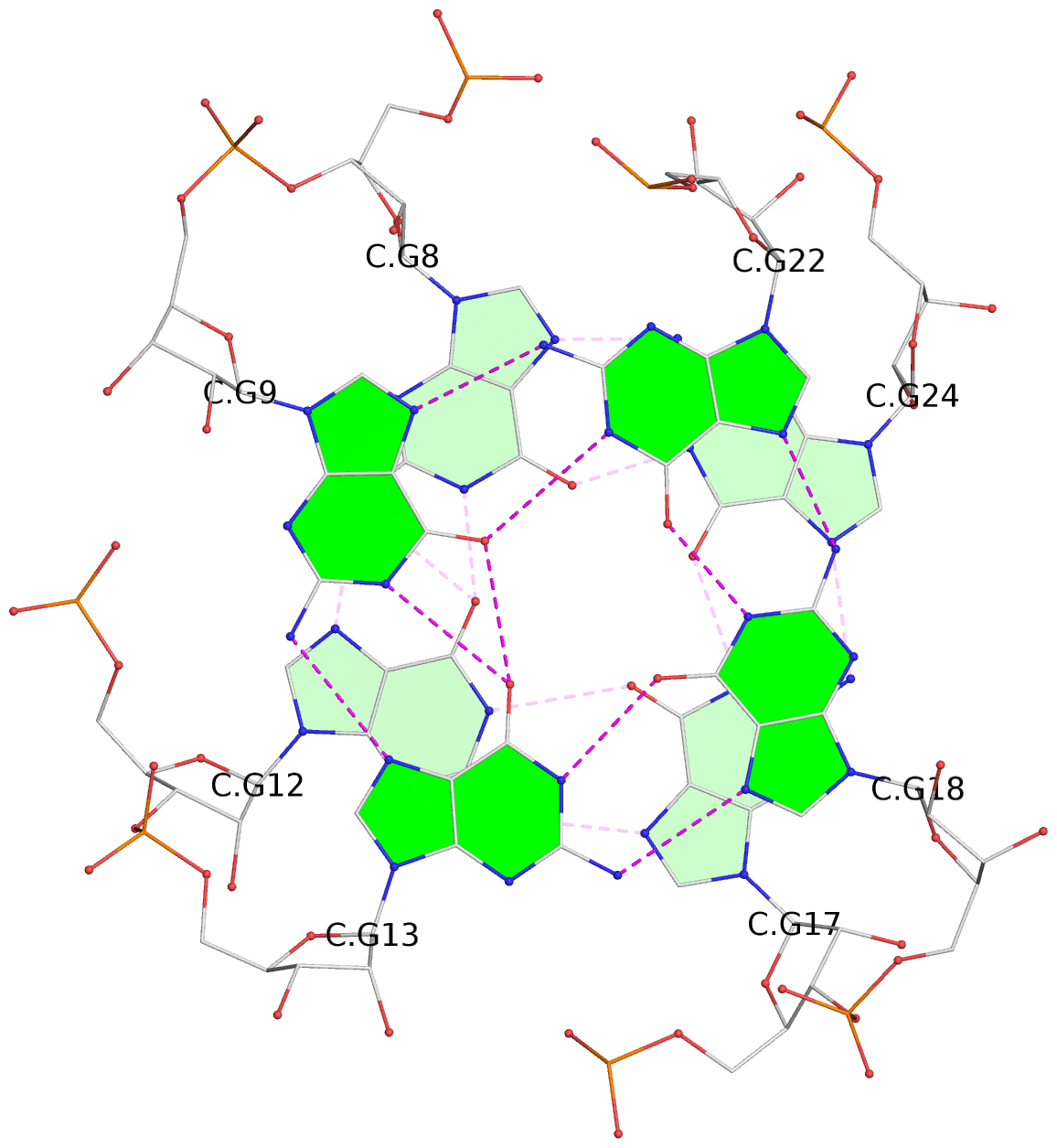
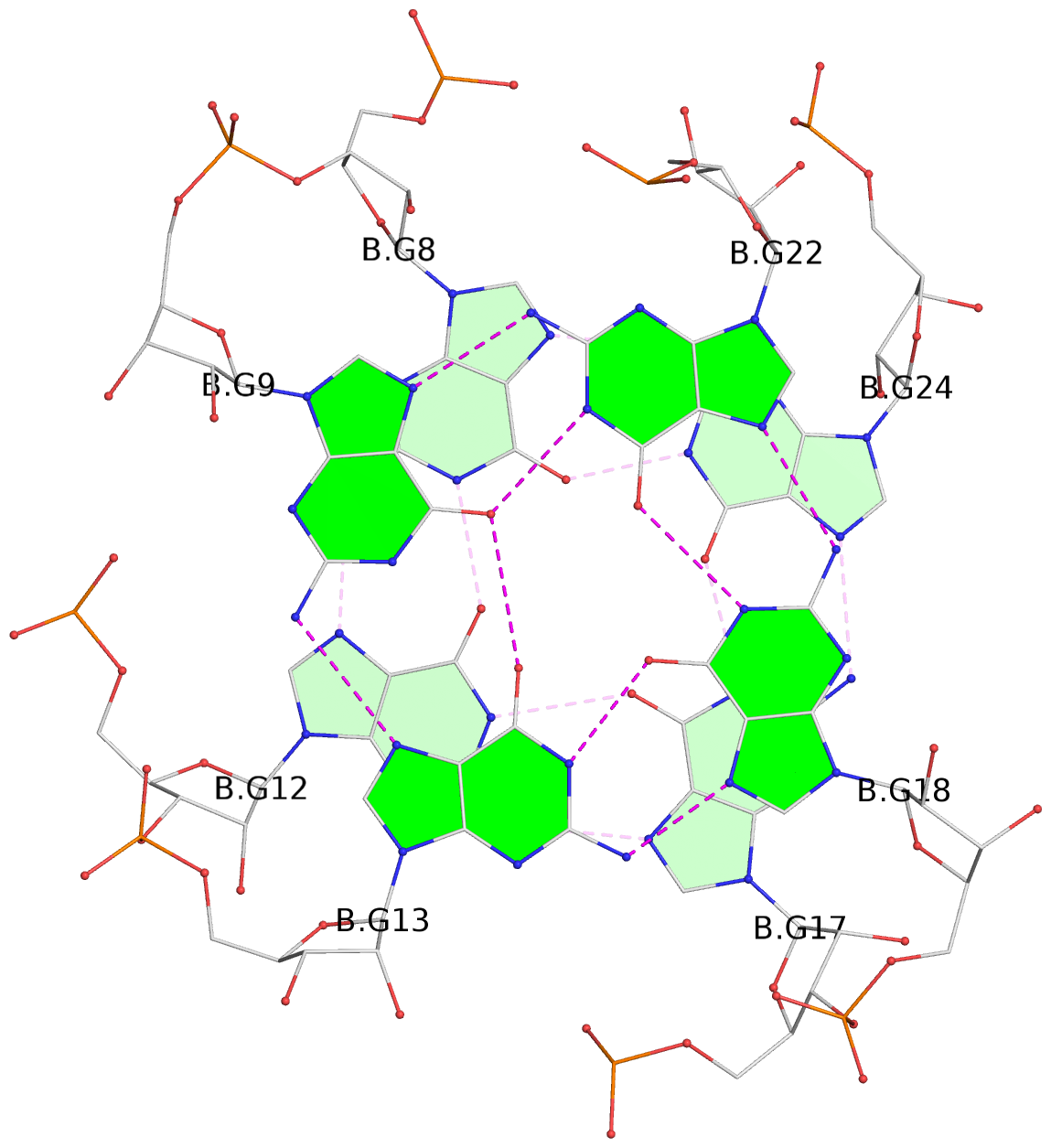
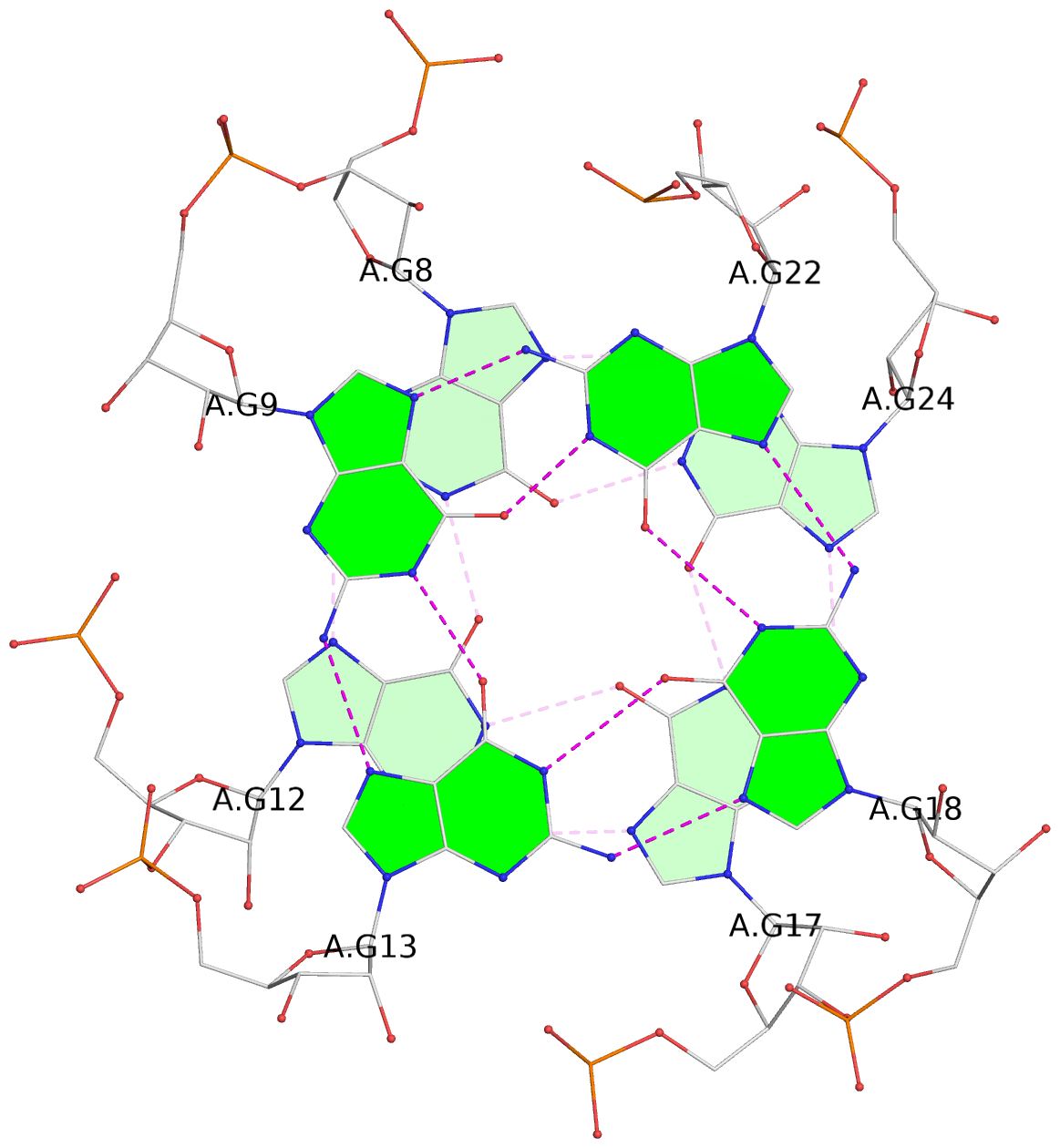
List of 8 G-tetrads
1 glyco-bond=---s sugar=-33- groove=--wn planarity=0.436 type=other nts=4 GGGG D.G8,D.G12,D.G17,D.G24 2 glyco-bond=---s sugar=-3-3 groove=--wn planarity=0.194 type=other nts=4 GGGG D.G9,D.G13,D.G18,D.G22 3 glyco-bond=---s sugar=-33- groove=--wn planarity=0.426 type=other nts=4 GGGG C.G8,C.G12,C.G17,C.G24 4 glyco-bond=---s sugar=-3-3 groove=--wn planarity=0.242 type=other nts=4 GGGG C.G9,C.G13,C.G18,C.G22 5 glyco-bond=---s sugar=-33- groove=--wn planarity=0.431 type=other nts=4 GGGG B.G8,B.G12,B.G17,B.G24 6 glyco-bond=---s sugar=-3-3 groove=--wn planarity=0.201 type=saddle nts=4 GGGG B.G9,B.G13,B.G18,B.G22 7 glyco-bond=---s sugar=-33- groove=--wn planarity=0.382 type=other nts=4 GGGG A.G8,A.G12,A.G17,A.G24 8 glyco-bond=---s sugar=-3-3 groove=--wn planarity=0.233 type=other nts=4 GGGG A.G9,A.G13,A.G18,A.G22
List of 4 G4-helices
In DSSR, a G4-helix is defined by stacking interactions of G-tetrads, regardless of backbone connectivity, and may contain more than one G4-stem.
Helix#1, 2 G-tetrad layers, INTRA-molecular, with 1 stem
Helix#2, 2 G-tetrad layers, INTRA-molecular, with 1 stem
Helix#3, 2 G-tetrad layers, INTRA-molecular, with 1 stem
Helix#4, 2 G-tetrad layers, INTRA-molecular, with 1 stem
List of 4 G4-stems
In DSSR, a G4-stem is defined as a G4-helix with backbone connectivity. Bulges are also allowed along each of the four strands.