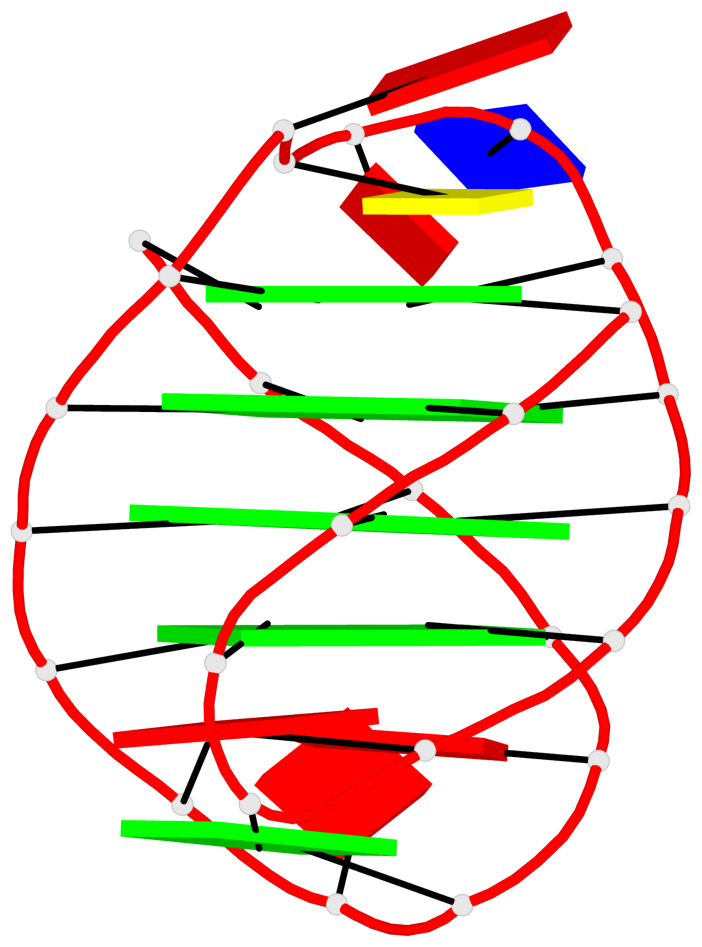
Detailed DSSR results for the G-quadruplex: PDB entry 6ftu
Created and maintained by Xiang-Jun Lu <xiangjun@x3dna.org>
Citation: Please cite the NAR'20 DSSR-PyMOL schematics paper and/or the NAR'15 DSSR method paper.
Summary information
- PDB id
- 6ftu
- Class
- DNA
- Method
- X-ray (2.95 Å)
- Summary
- Structure of a quadruplex forming sequence from d. discoideum
- Reference
- Guedin A, Lin LY, Armane S, Lacroix L, Mergny JL, Thore S, Yatsunyk LA (2018): "Quadruplexes in 'Dicty': crystal structure of a four-quartet G-quadruplex formed by G-rich motif found in the Dictyostelium discoideum genome." Nucleic Acids Res., 46, 5297-5307. doi: 10.1093/nar/gky290.
- Abstract
- Guanine-rich DNA has the potential to fold into non-canonical G-quadruplex (G4) structures. Analysis of the genome of the social amoeba Dictyostelium discoideum indicates a low number of sequences with G4-forming potential (249-1055). Therefore, D. discoideum is a perfect model organism to investigate the relationship between the presence of G4s and their biological functions. As a first step in this investigation, we crystallized the dGGGGGAGGGGTACAGGGGTACAGGGG sequence from the putative promoter region of two divergent genes in D. discoideum. According to the crystal structure, this sequence folds into a four-quartet intramolecular antiparallel G4 with two lateral and one diagonal loops. The G-quadruplex core is further stabilized by a G-C Watson-Crick base pair and a A-T-A triad and displays high thermal stability (Tm > 90°C at 100 mM KCl). Biophysical characterization of the native sequence and loop mutants suggests that the DNA adopts the same structure in solution and in crystalline form, and that loop interactions are important for the G4 stability but not for its folding. Four-tetrad G4 structures are sparse. Thus, our work advances understanding of the structural diversity of G-quadruplexes and yields coordinates for in silico drug screening programs and G4 predictive tools.
- G4 notes
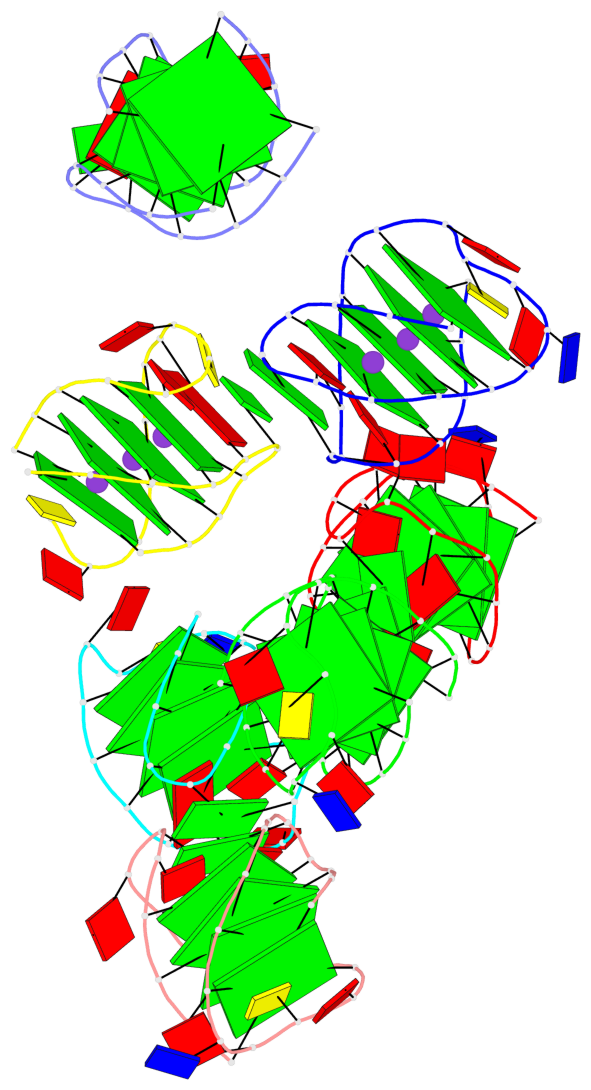
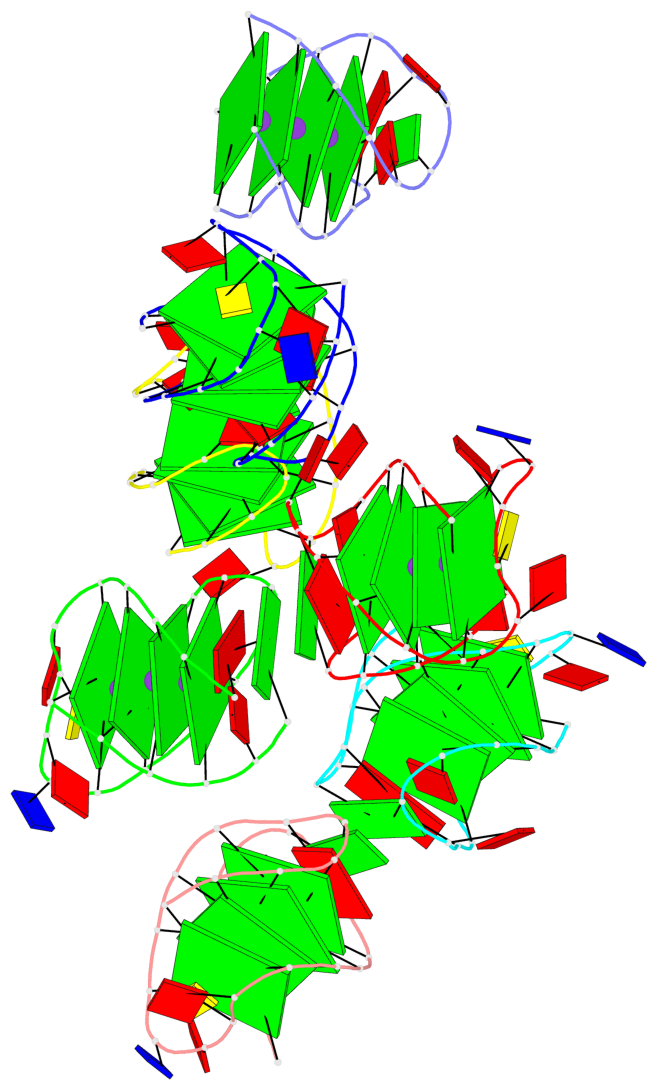
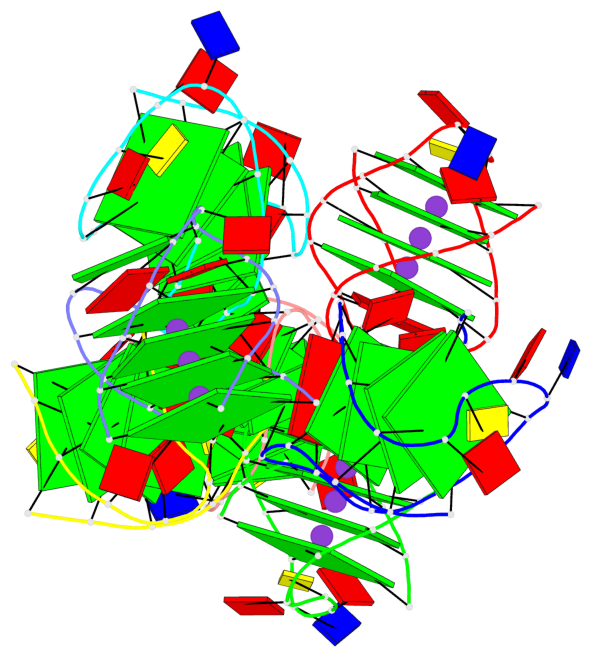
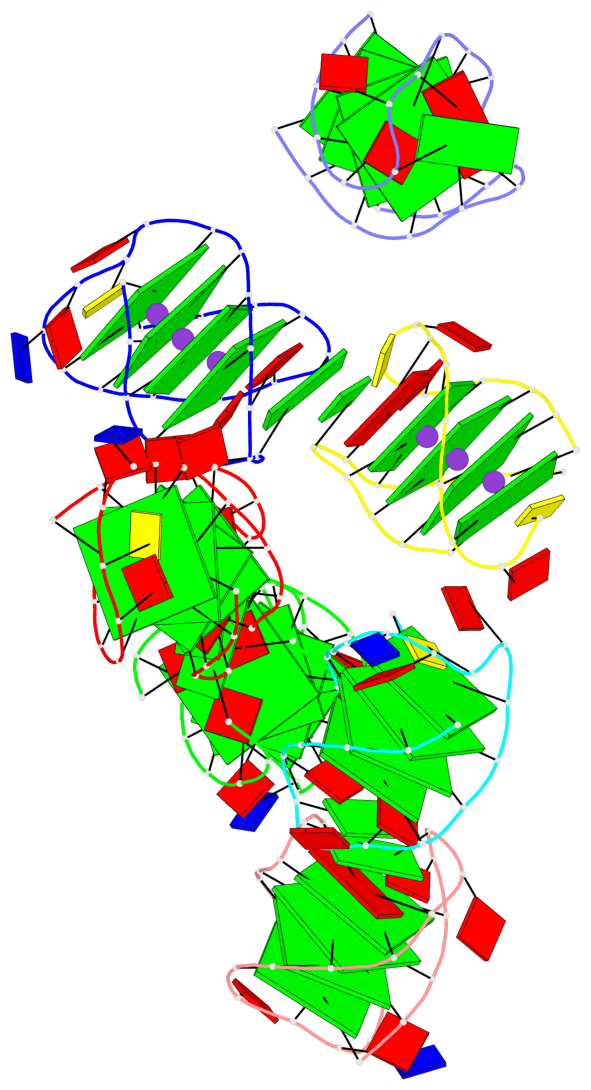
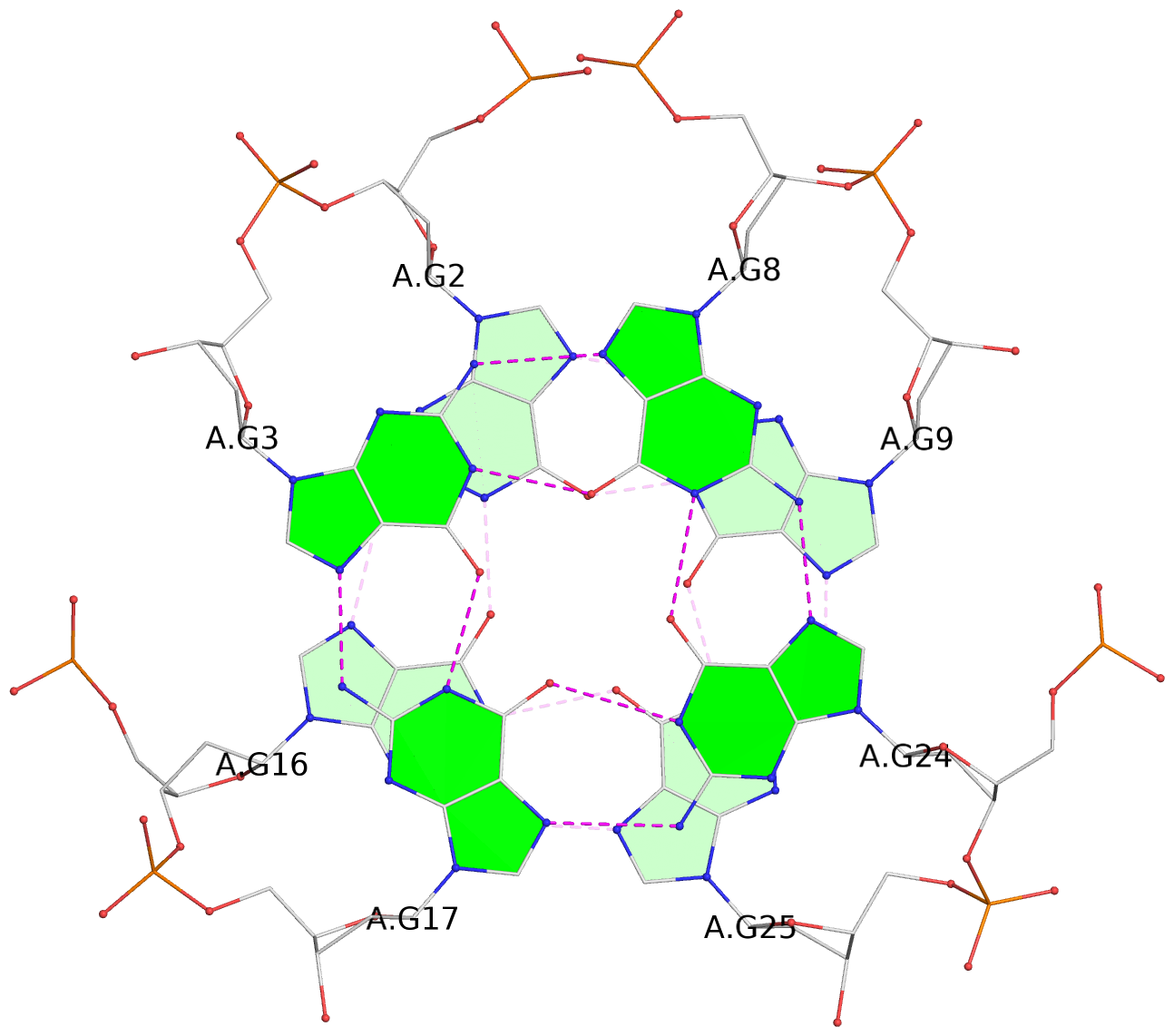
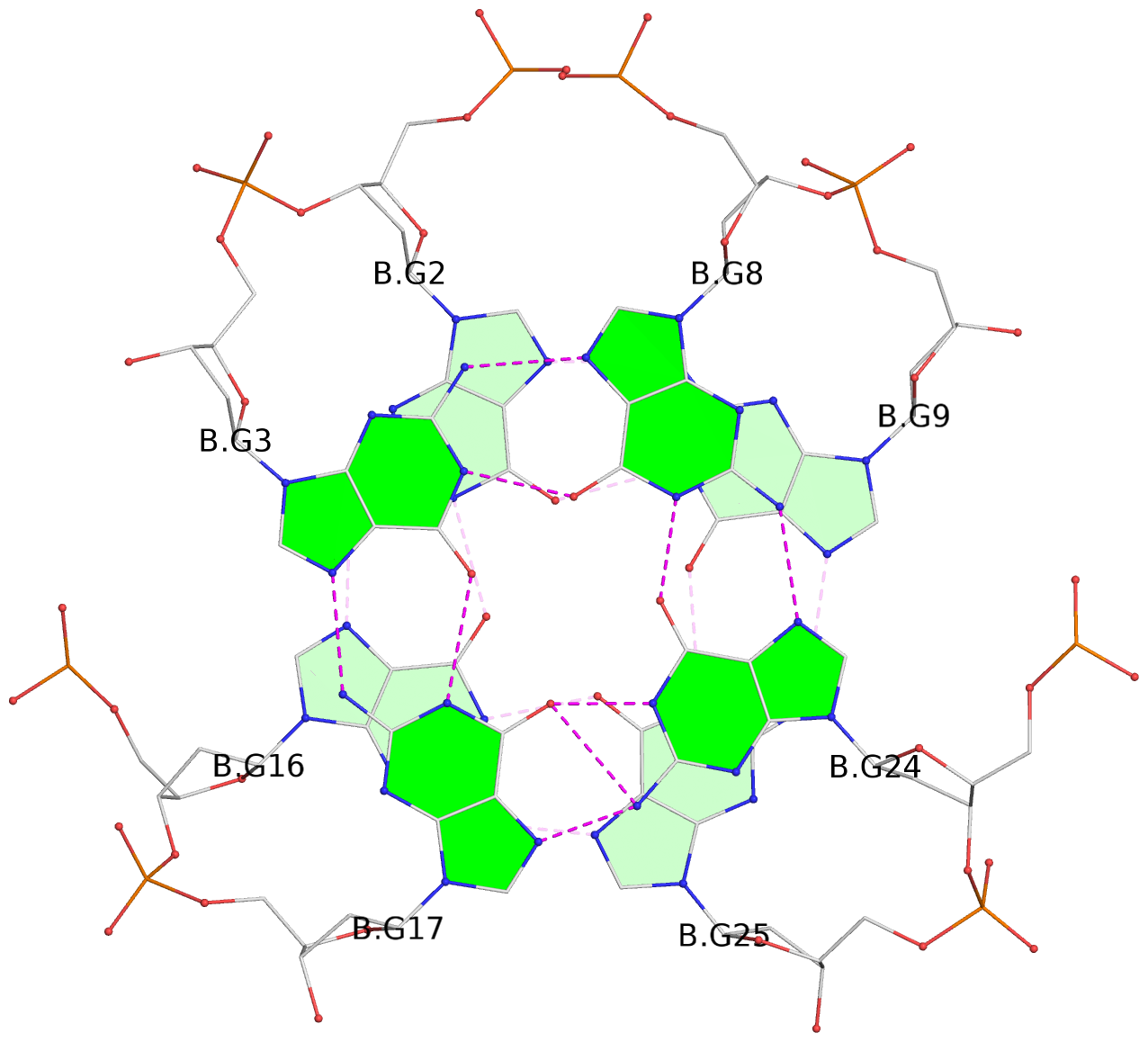
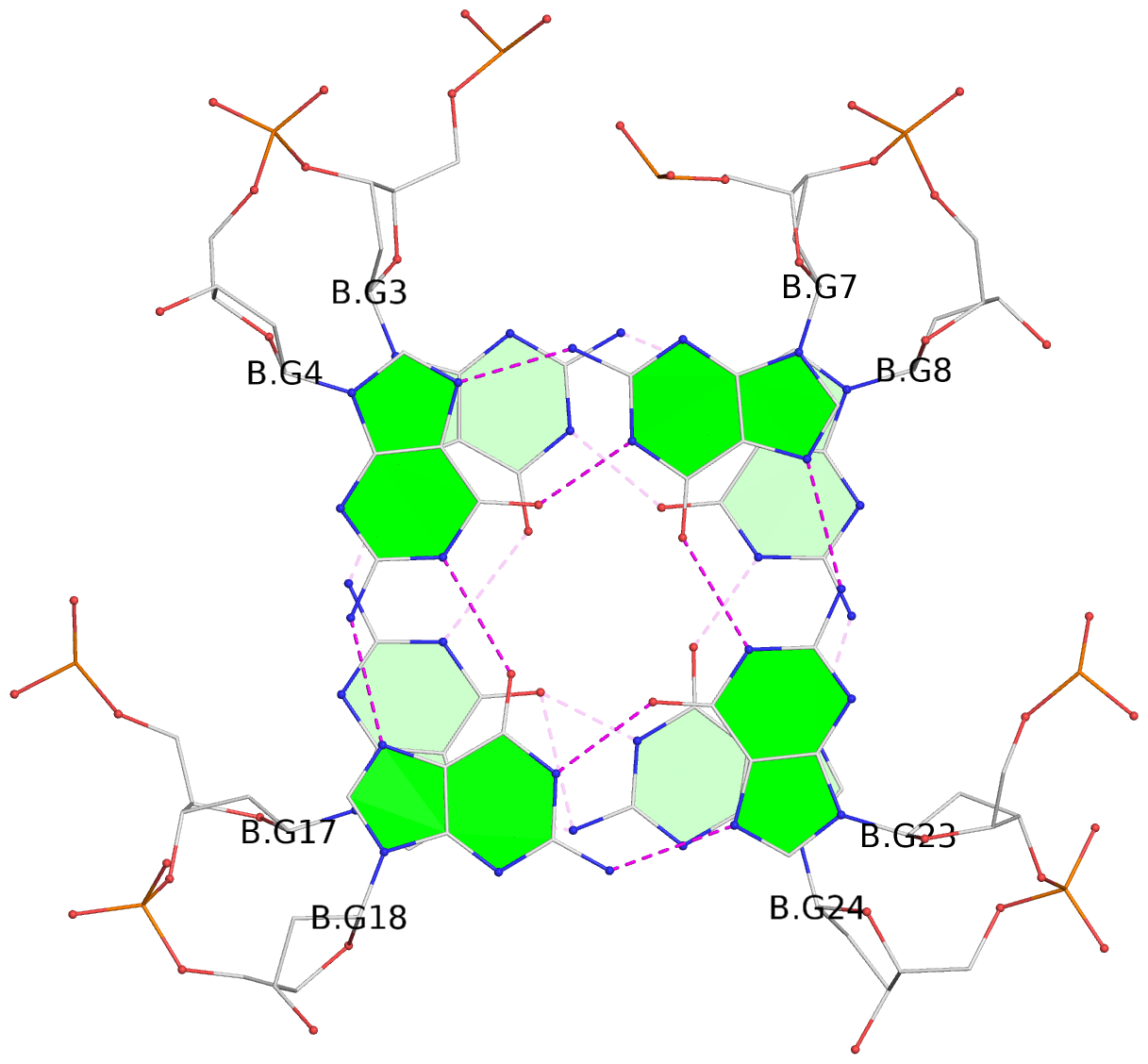
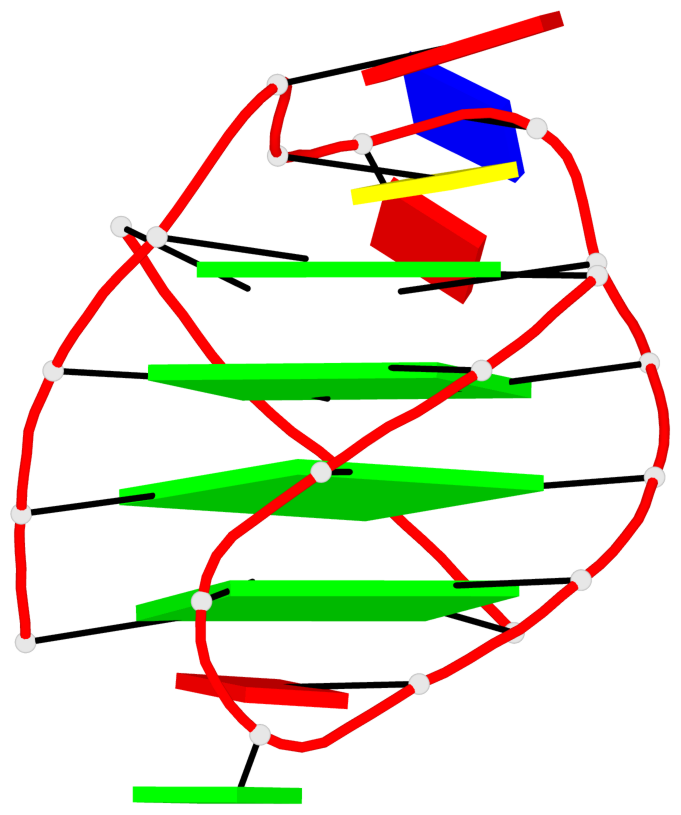
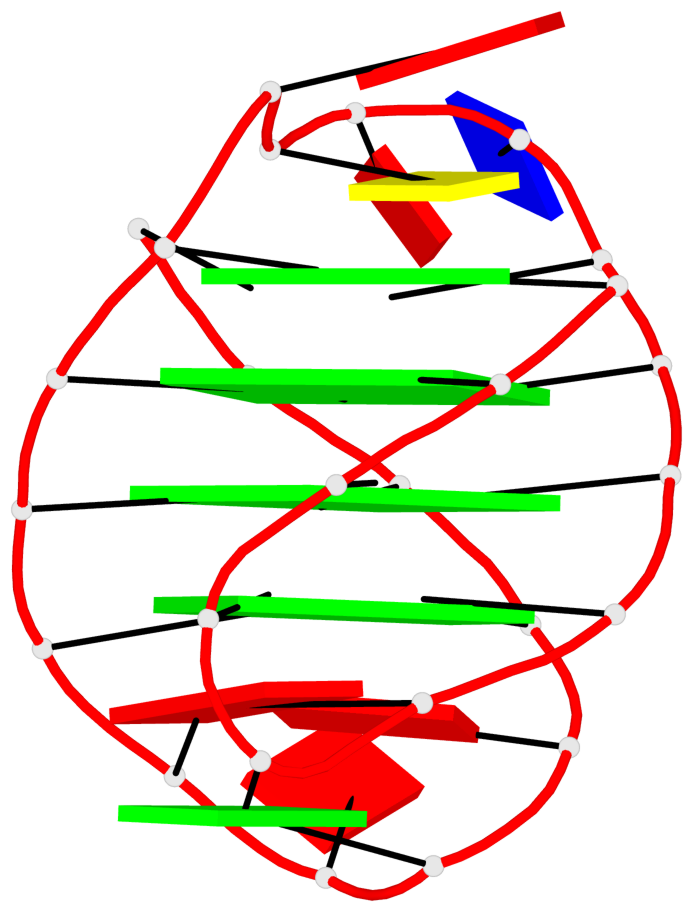
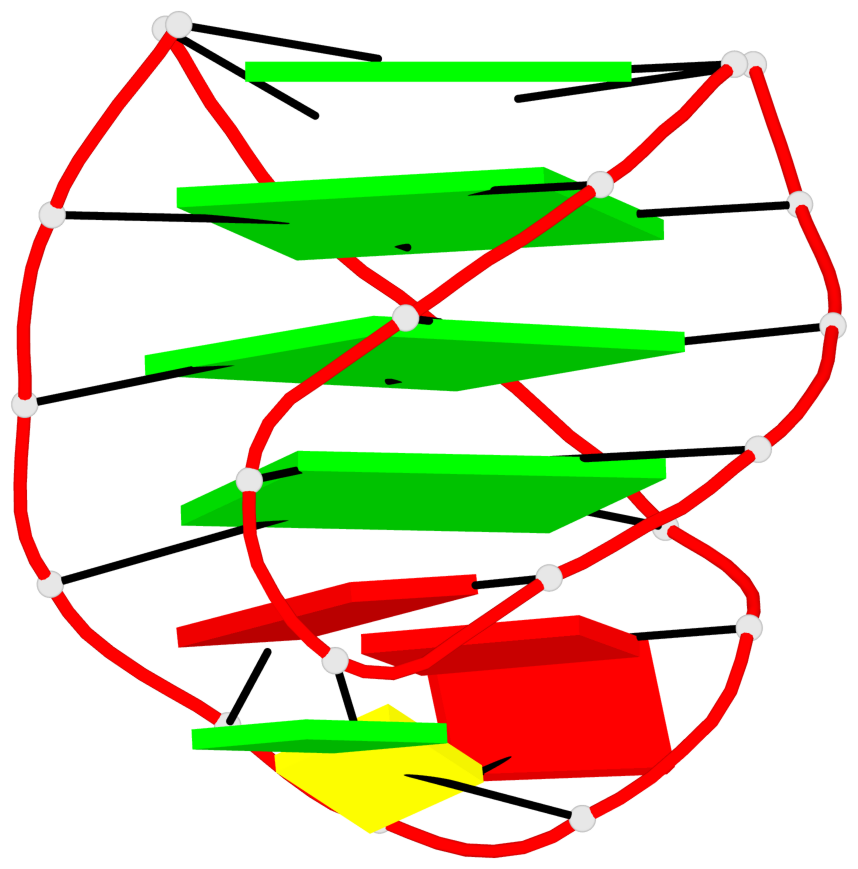
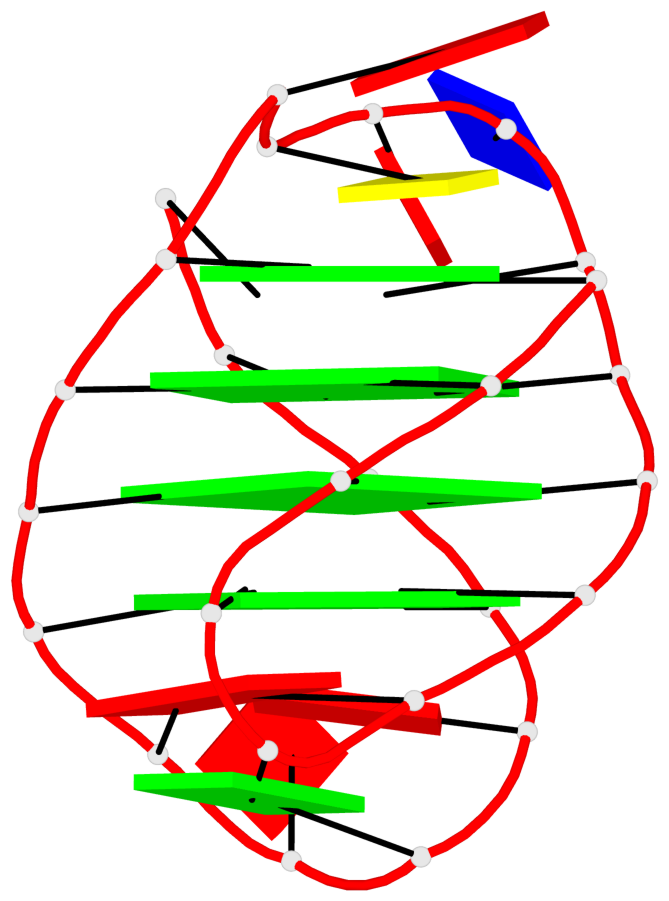
- 28 G-tetrads, 7 G4 helices, 7 G4 stems, 4(+Ln-Lw), (2+2), UUDD; 4(+LnD), basket(2+2), UUDD; 4(+LnD-Lw), basket(2+2), UUDD
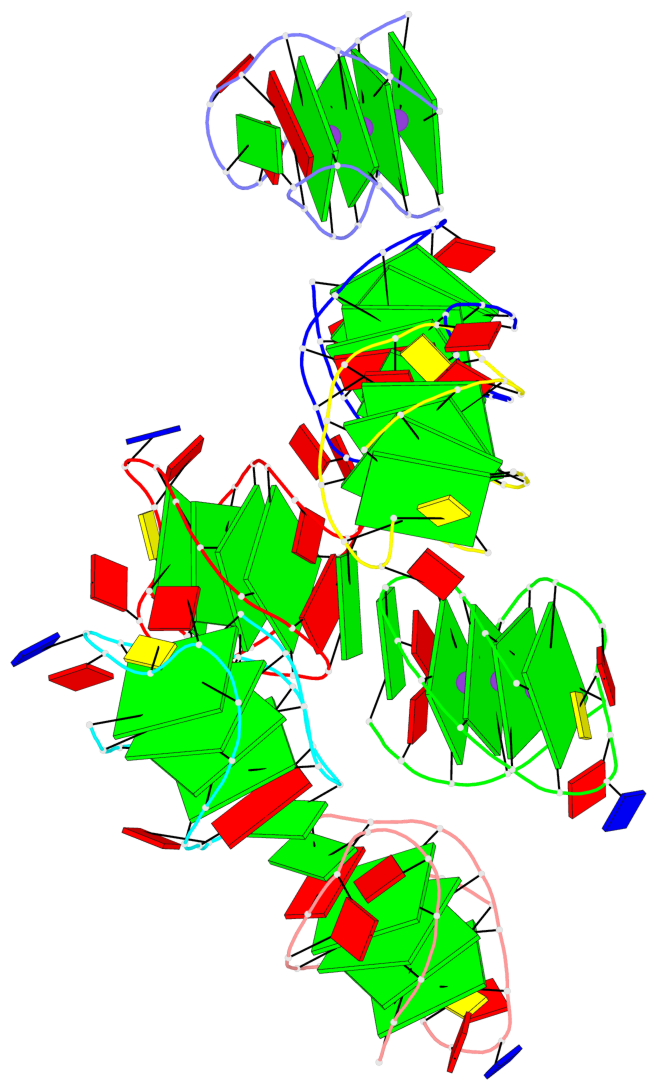
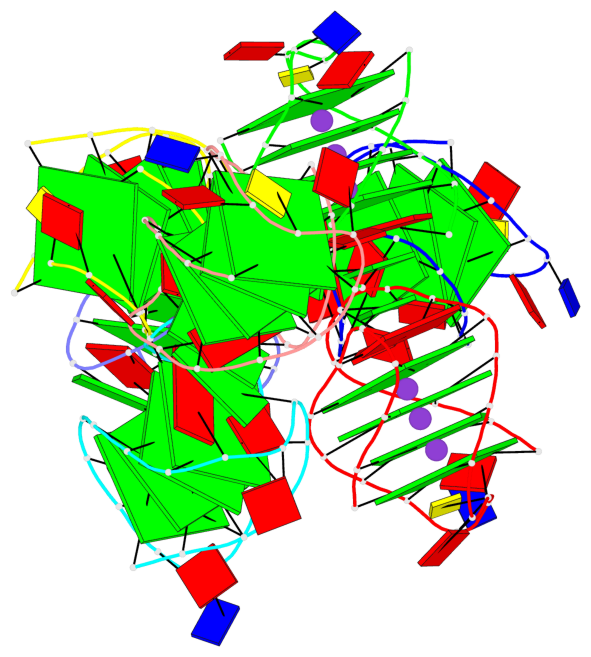
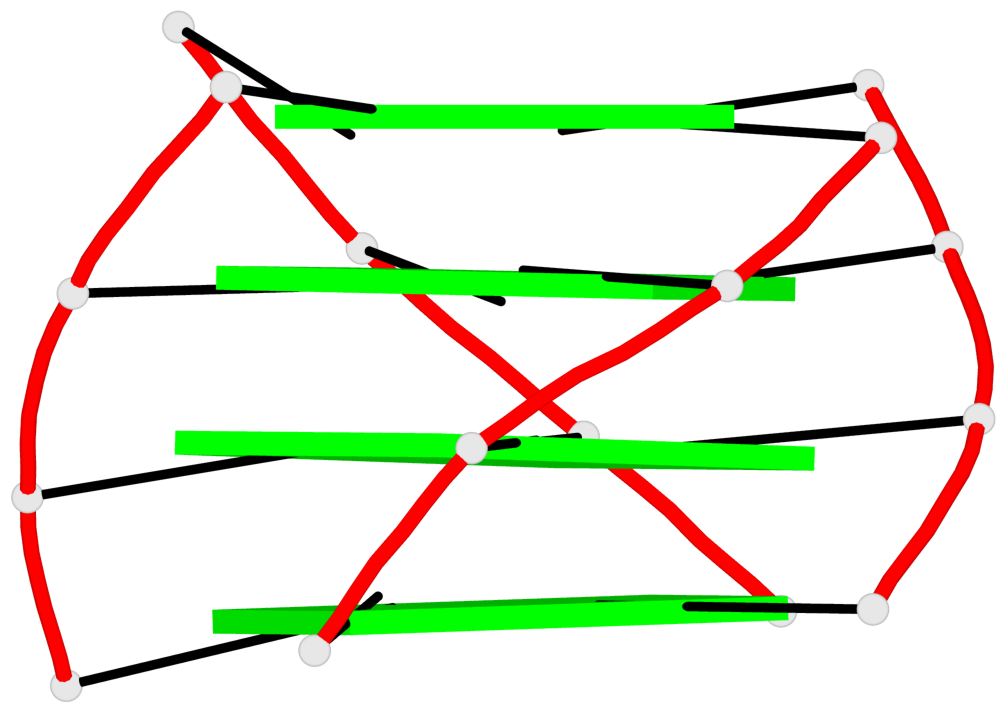
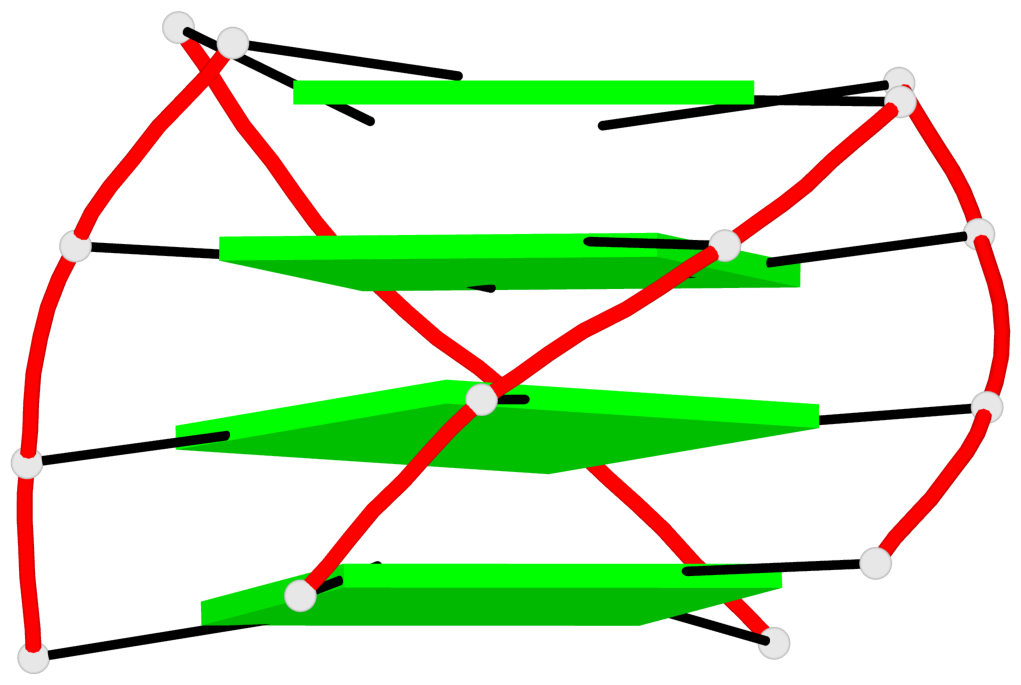
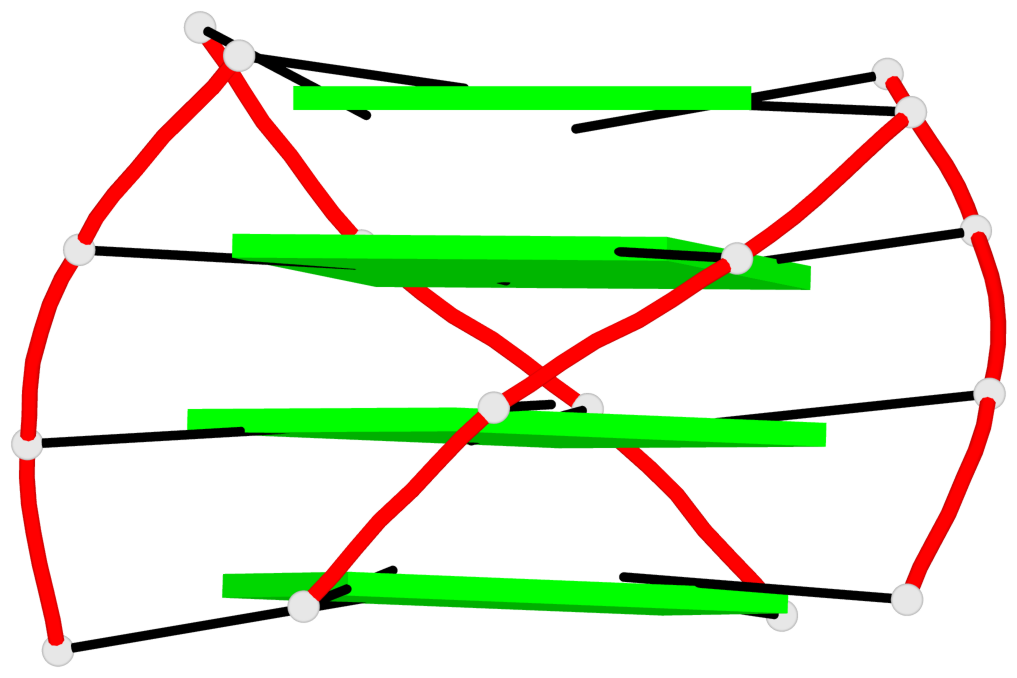
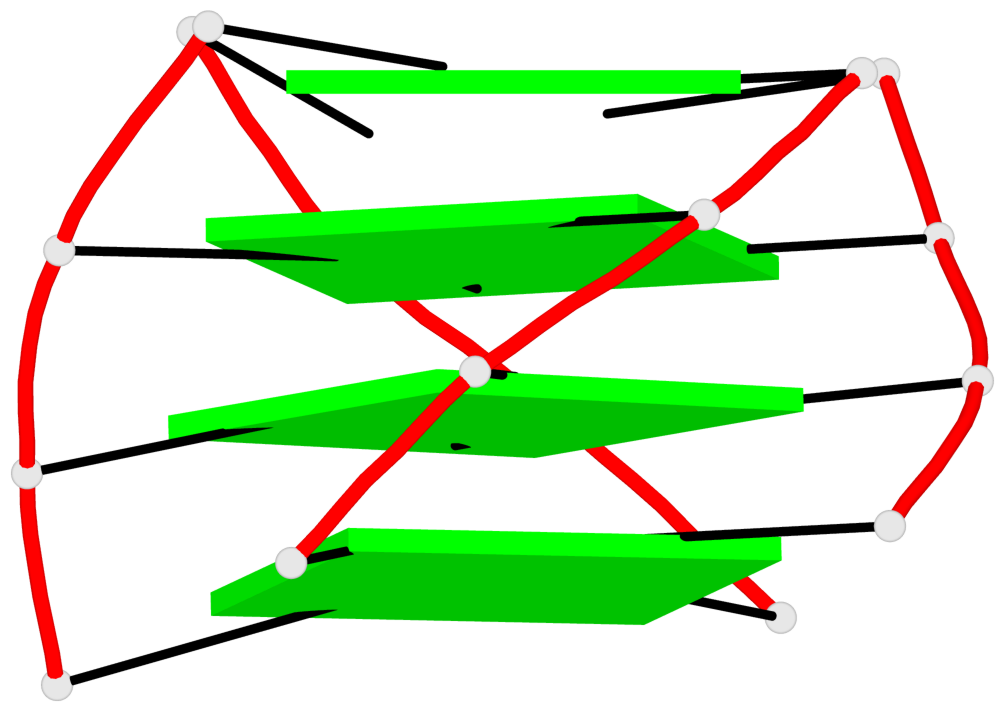
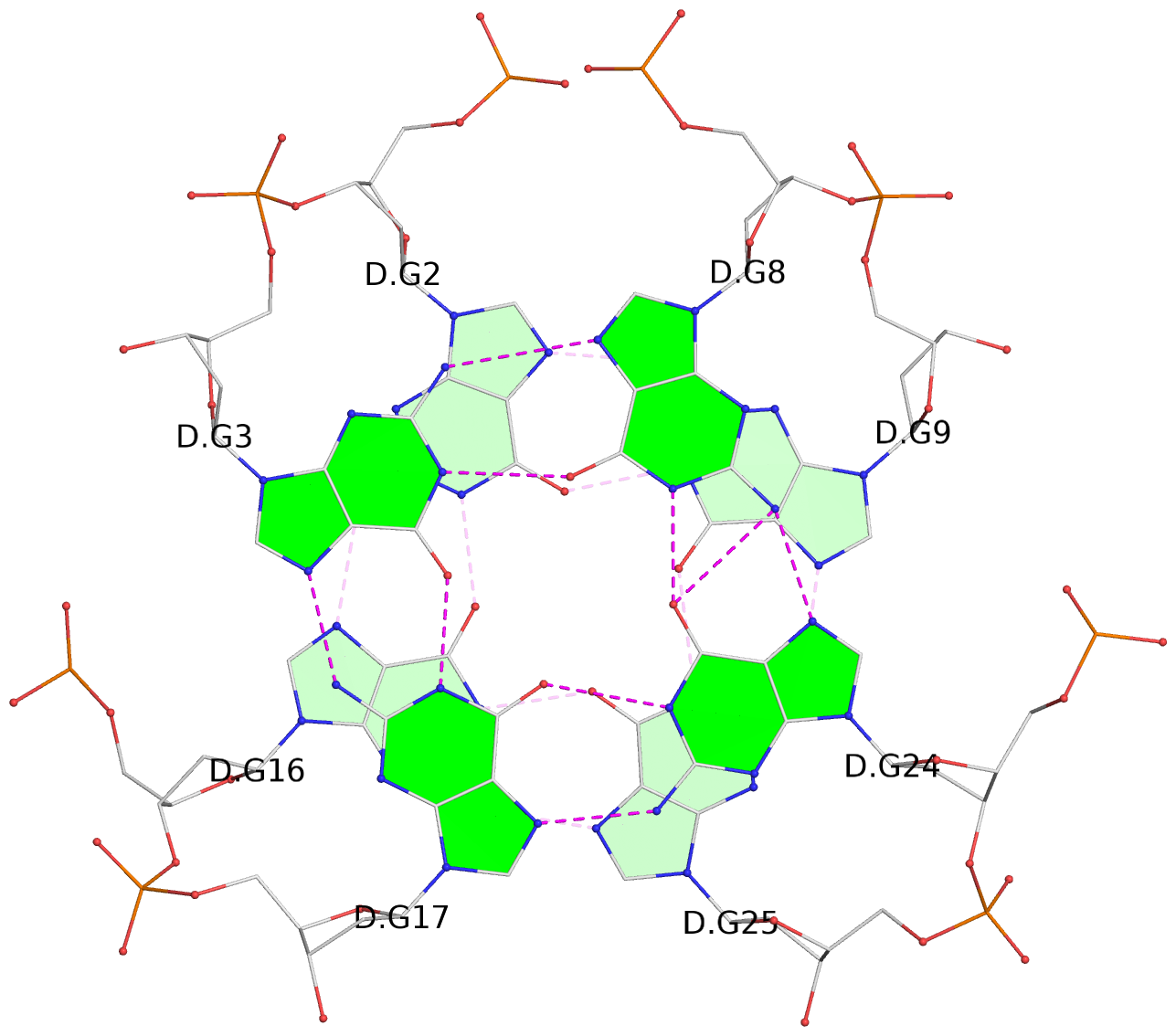
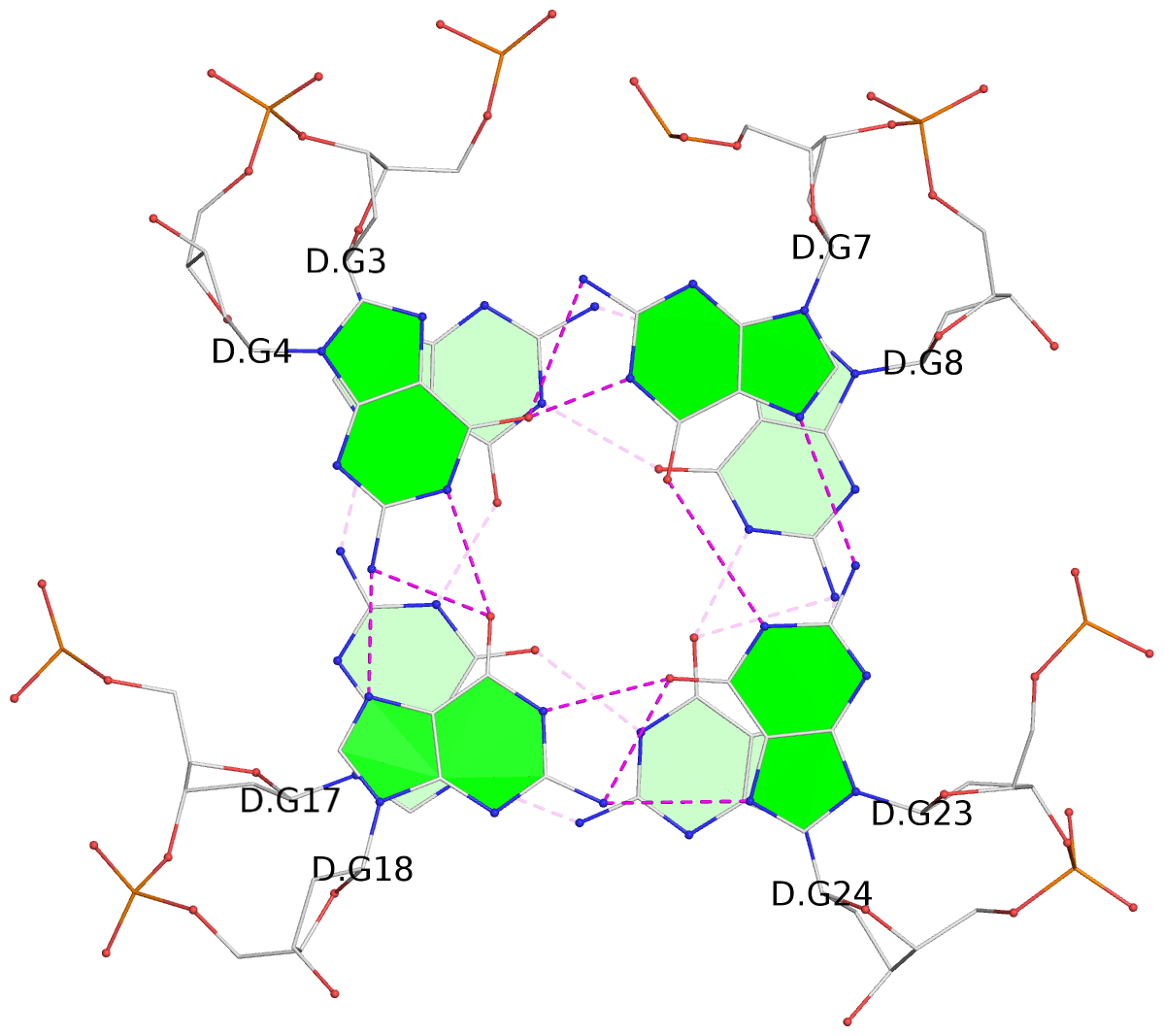
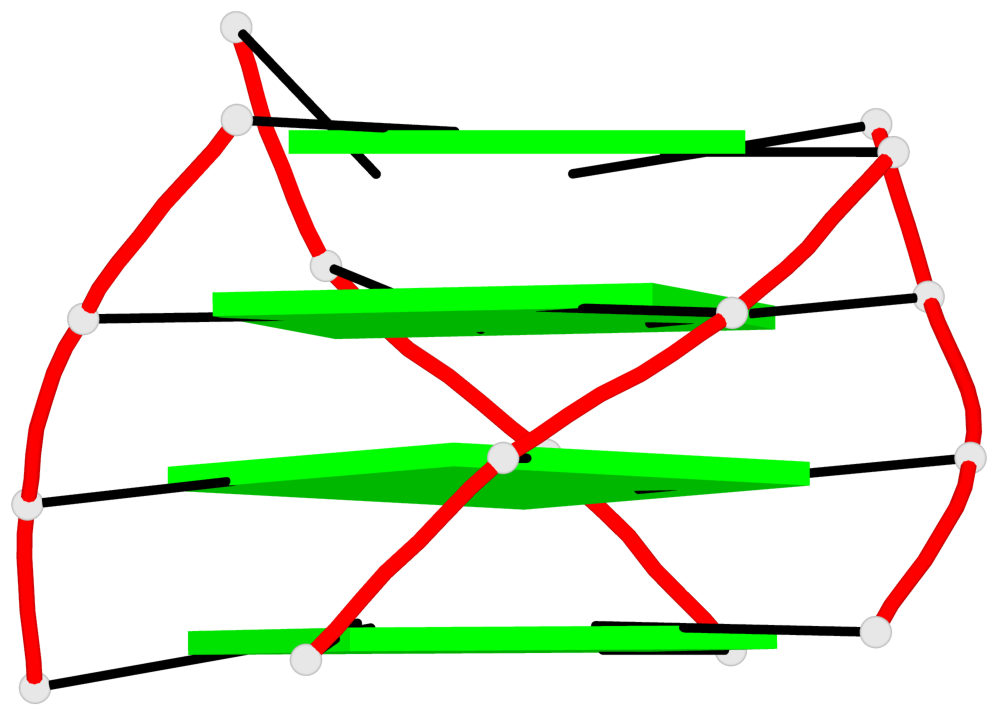
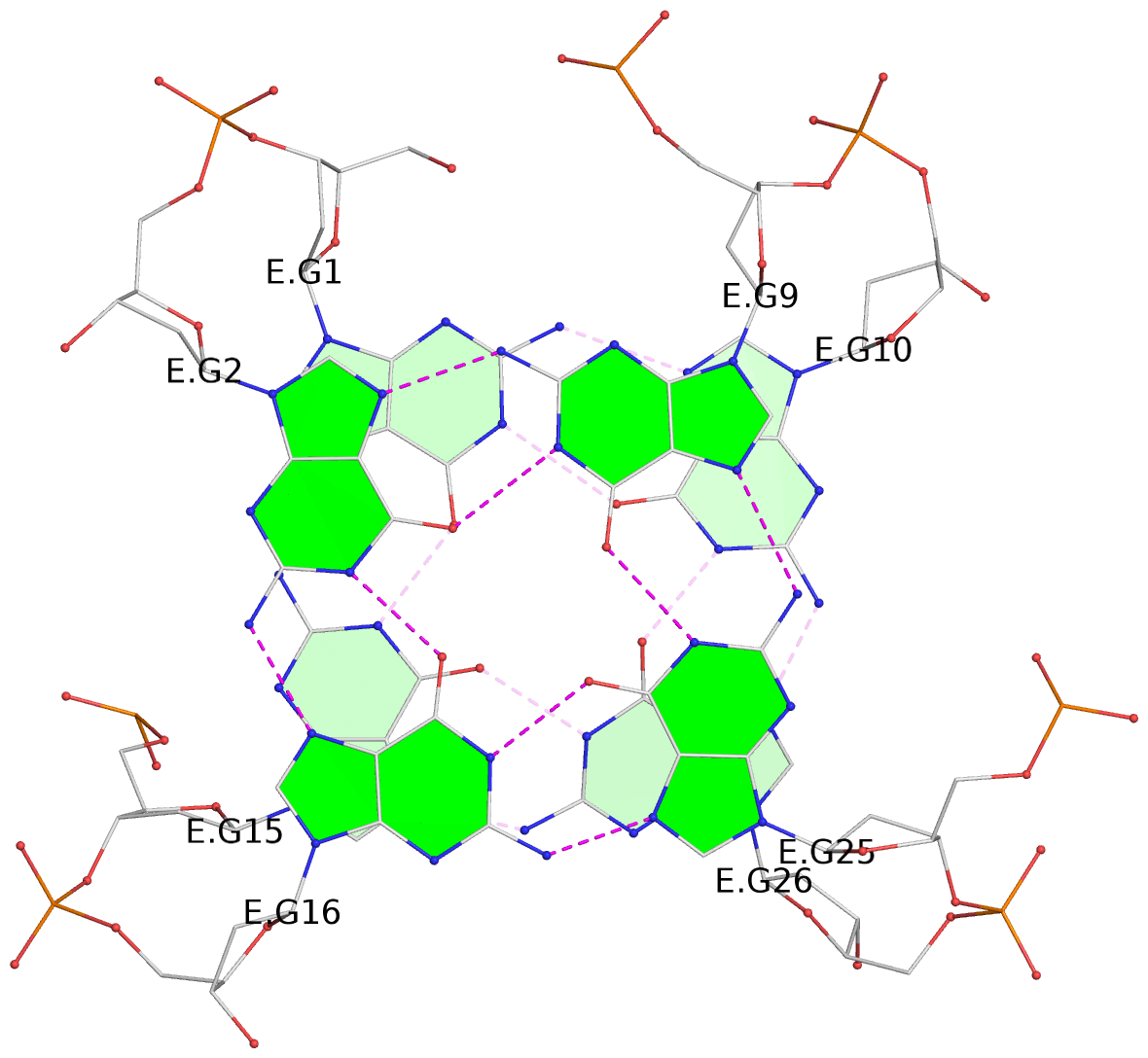
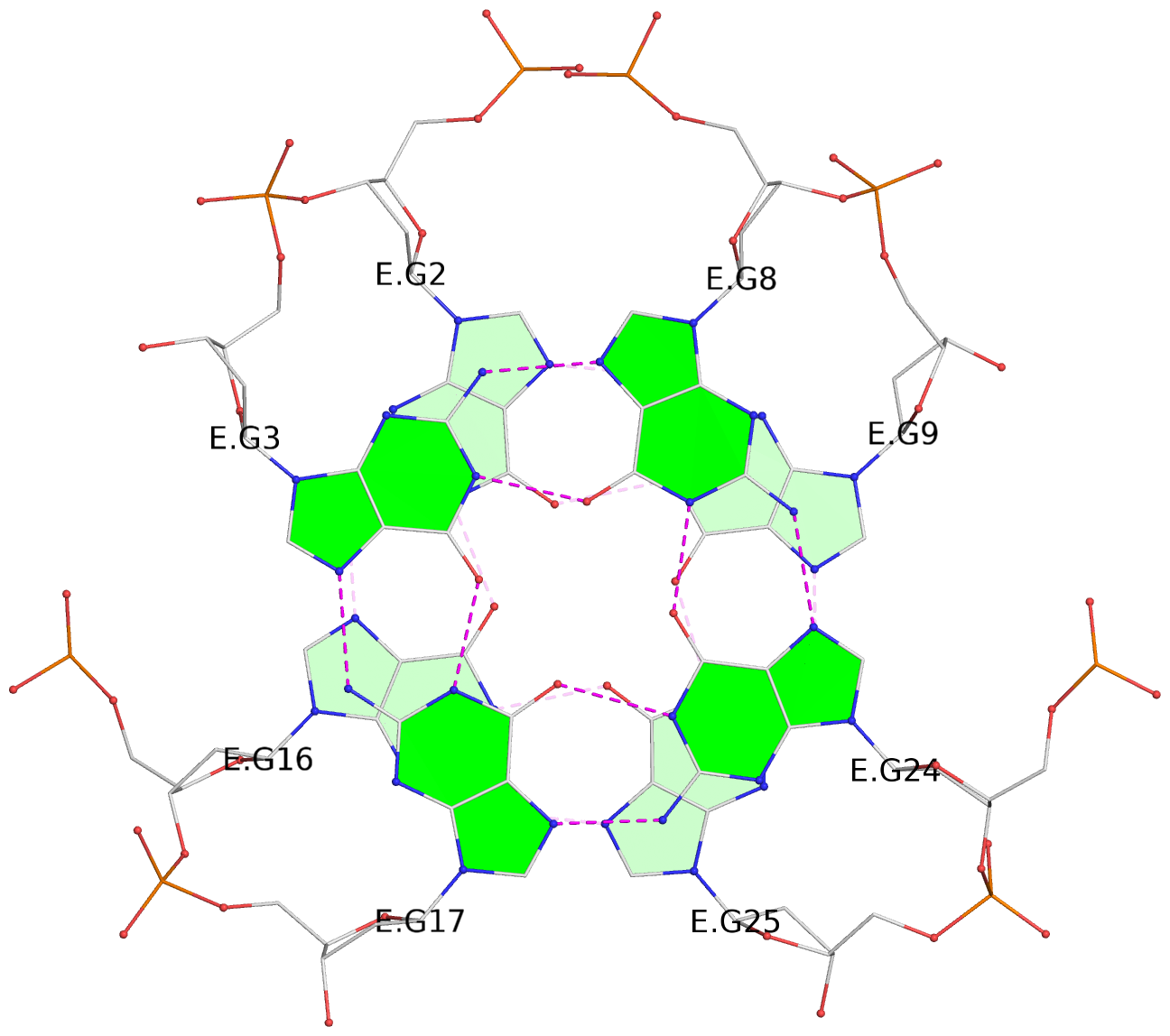
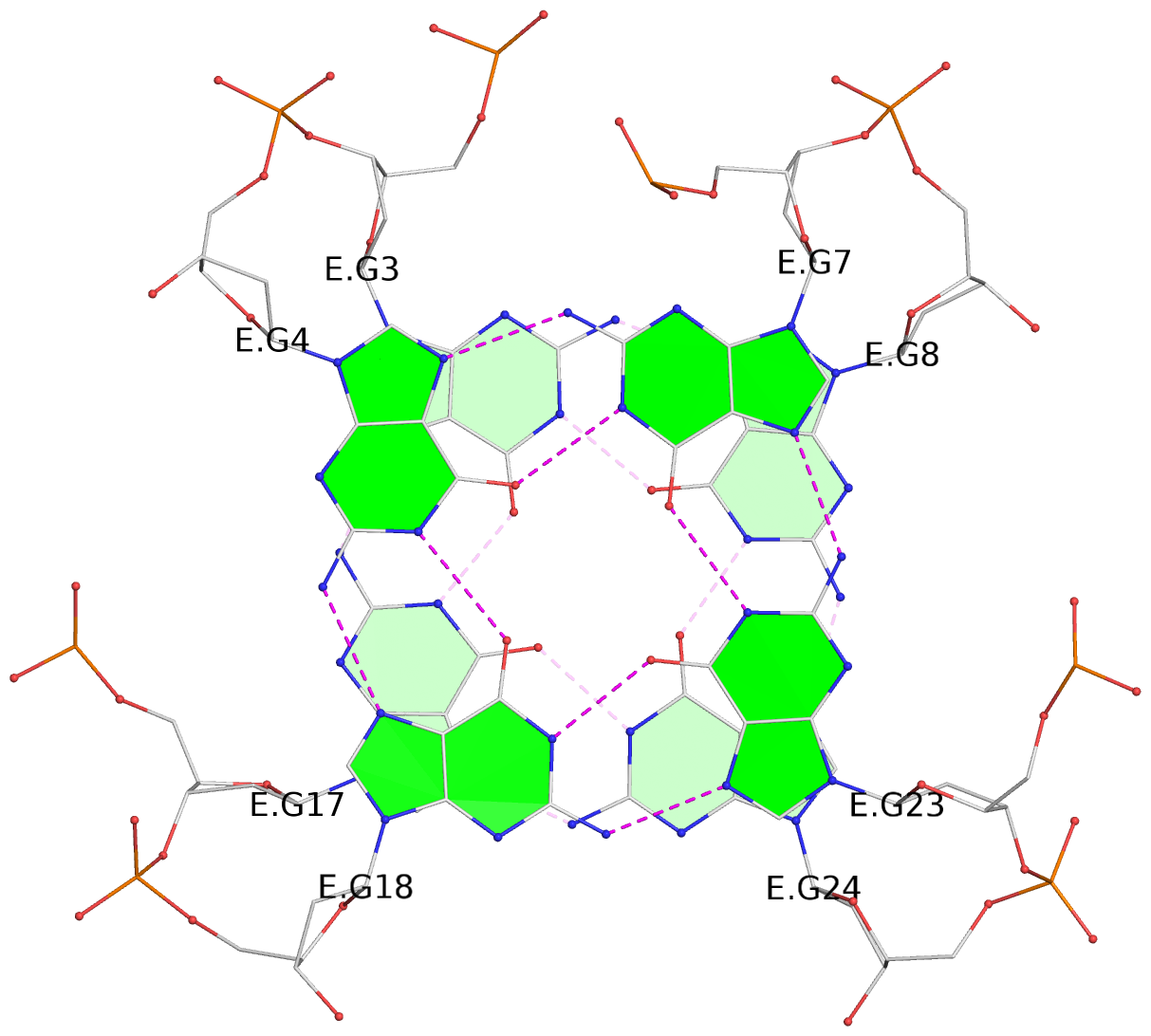
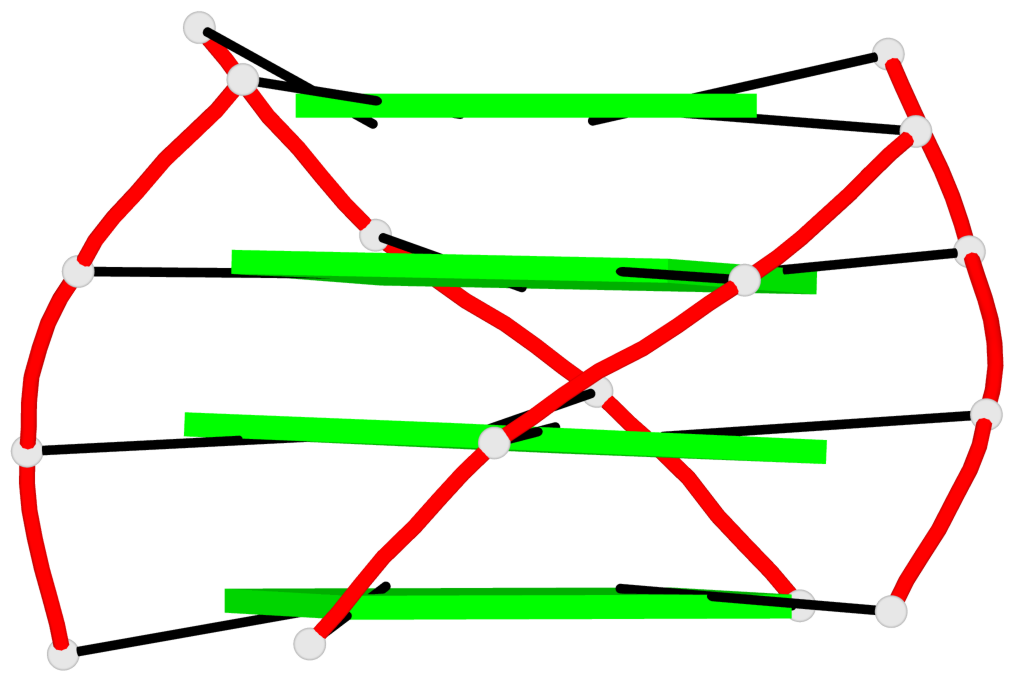
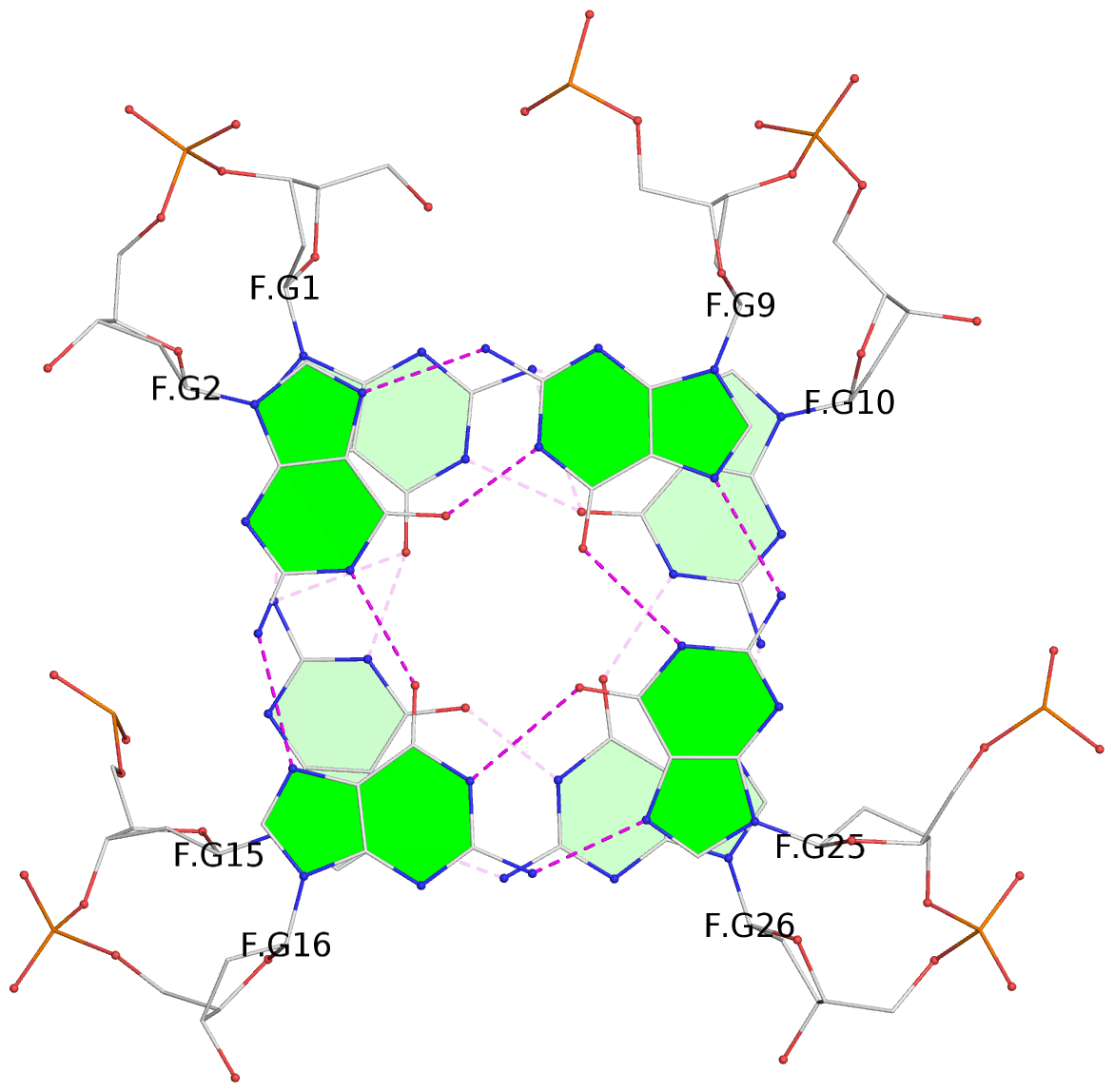
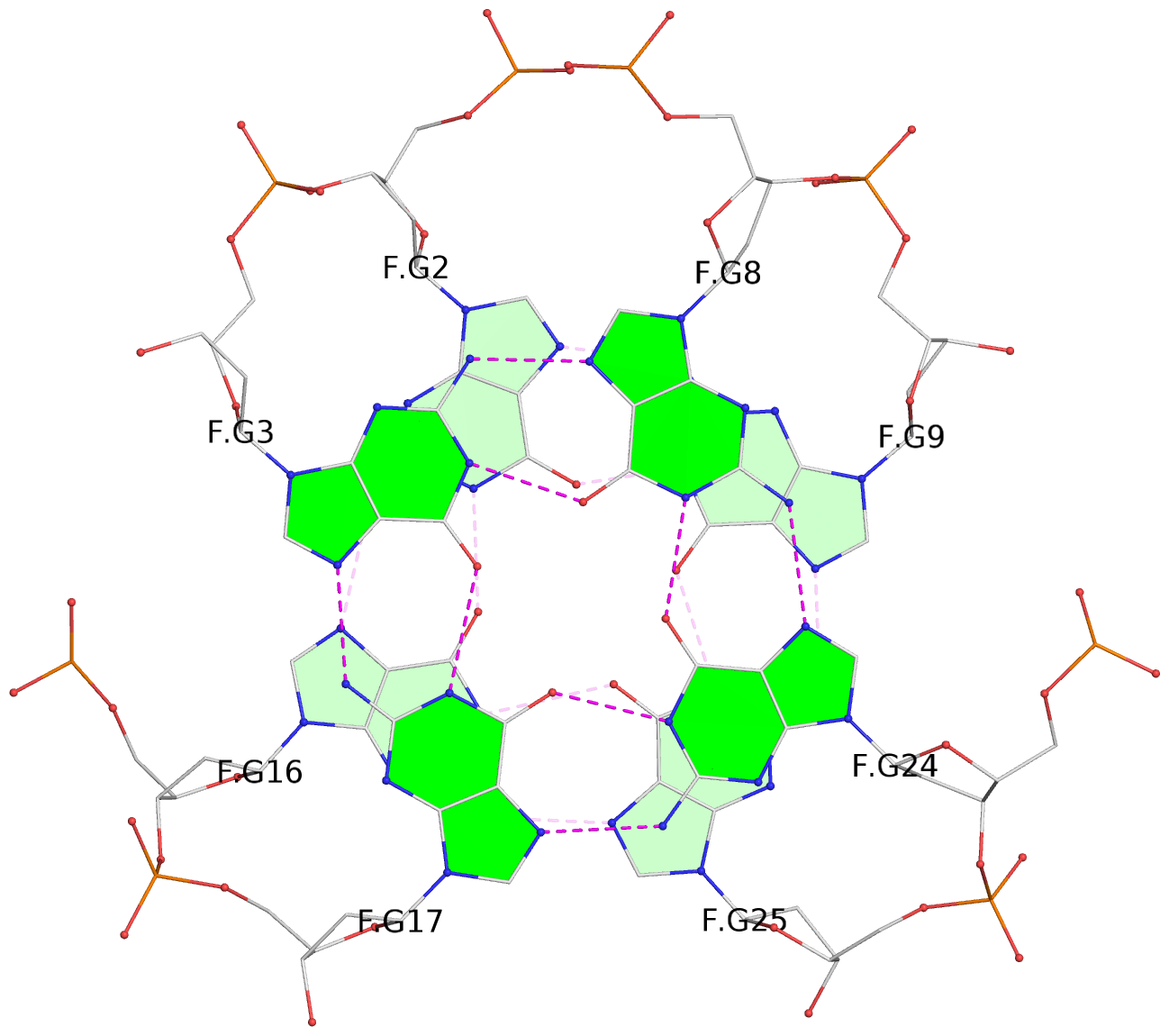
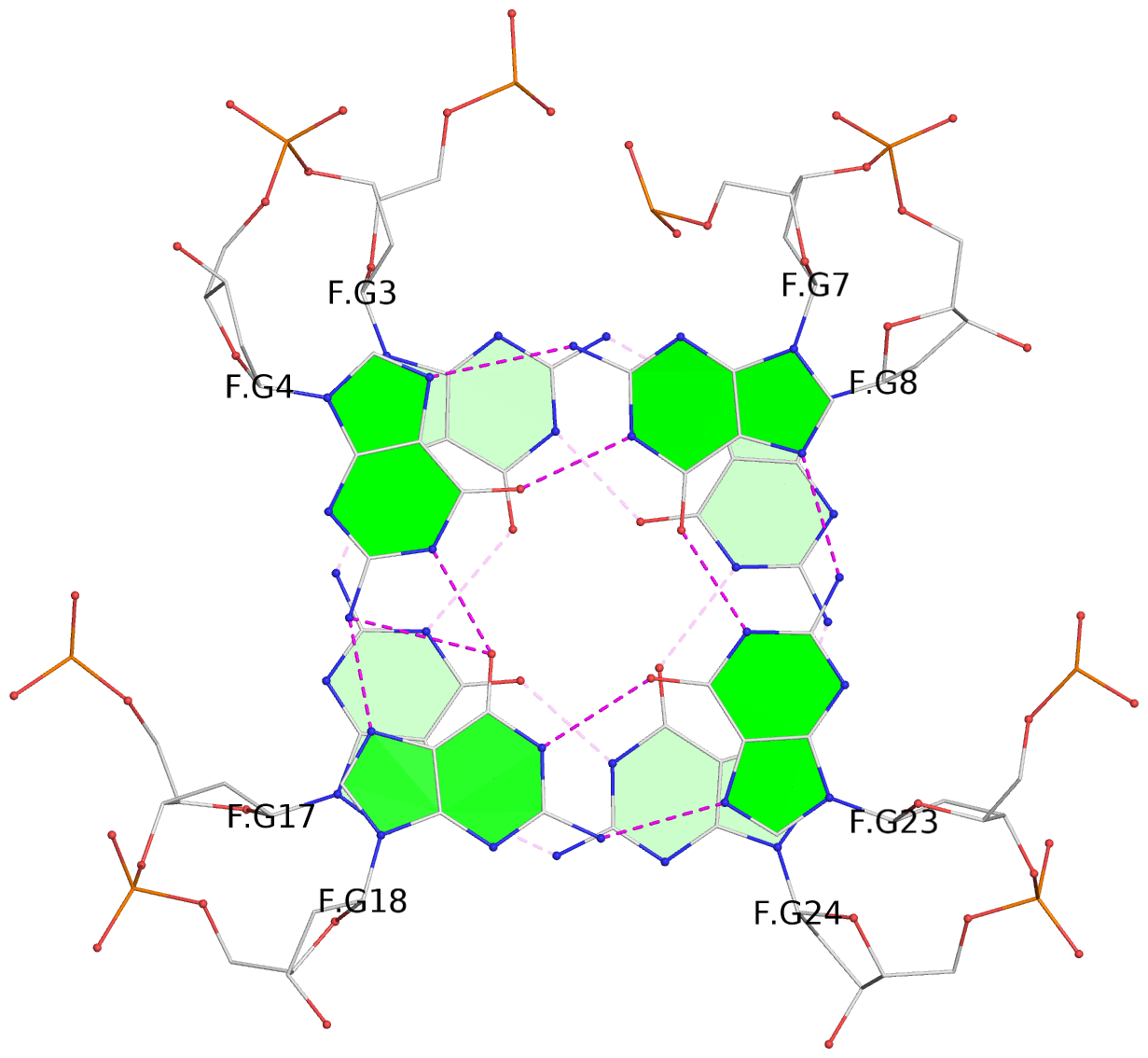
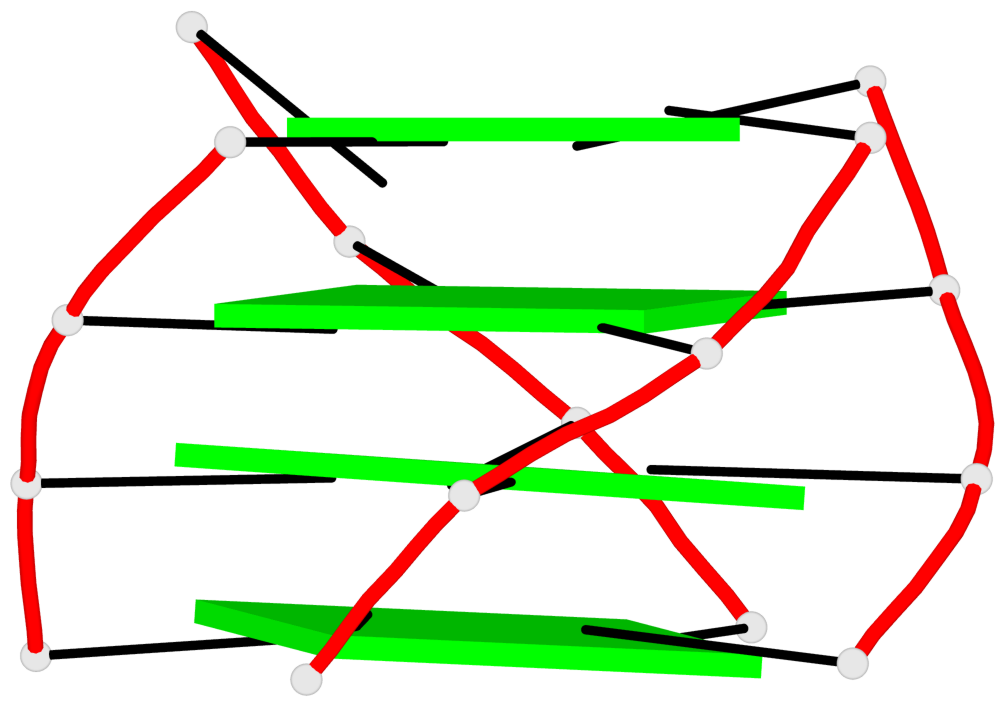
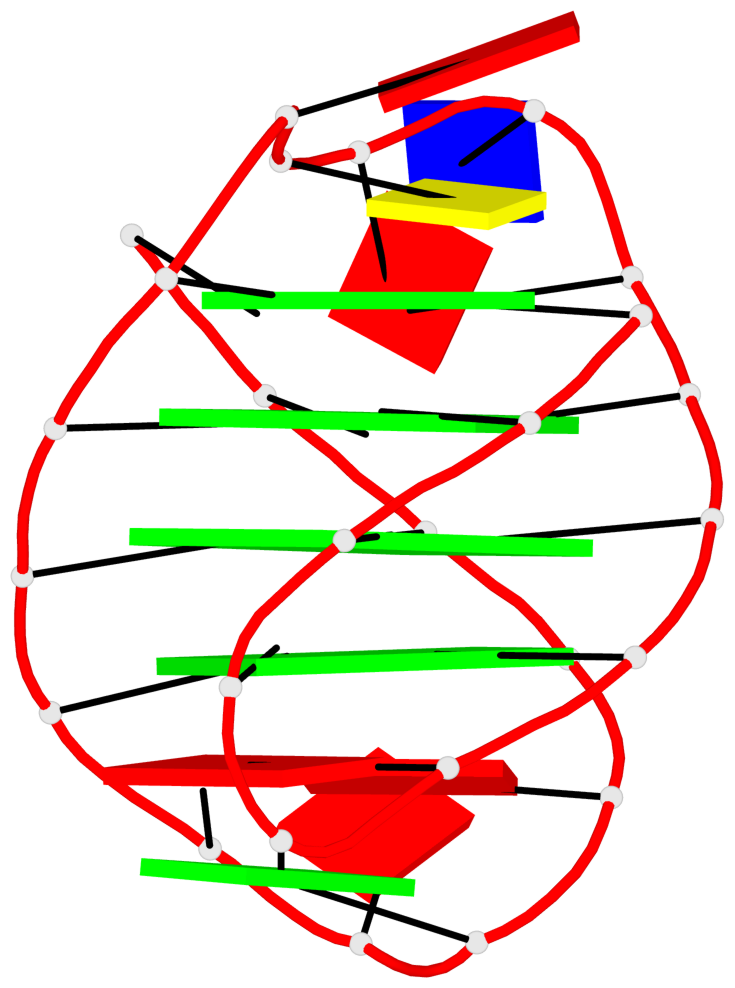
Base-block schematics in six views
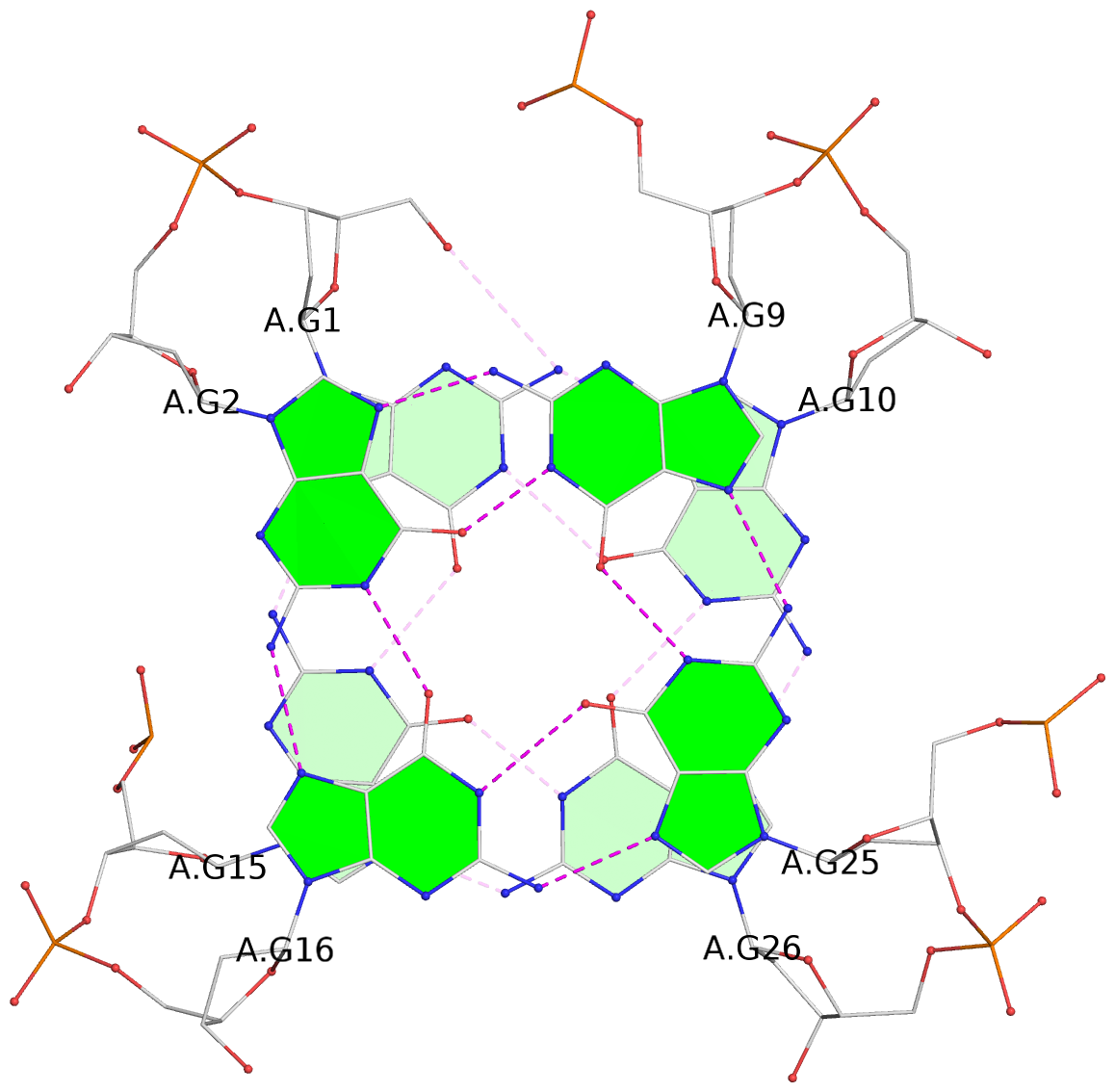
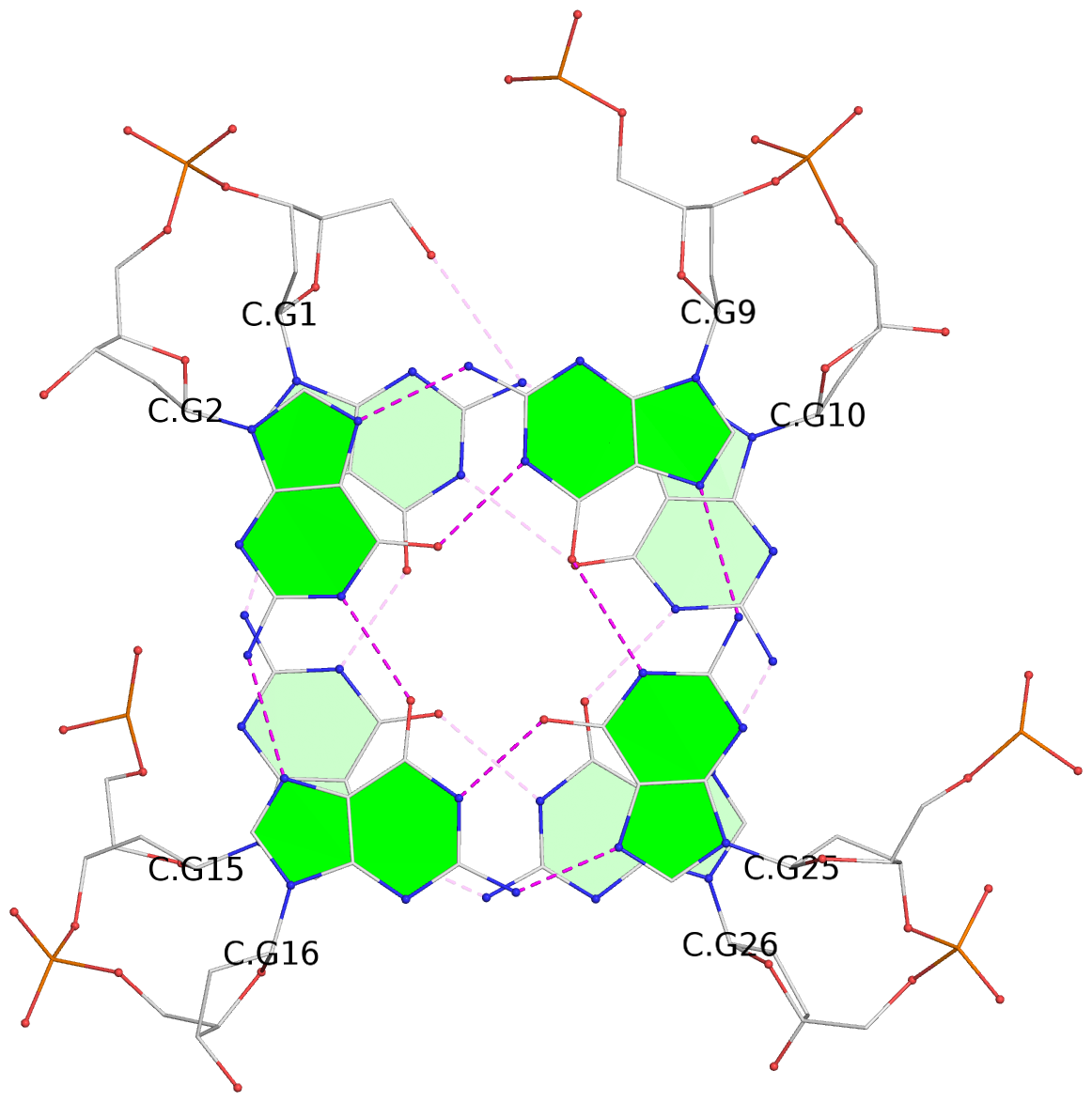
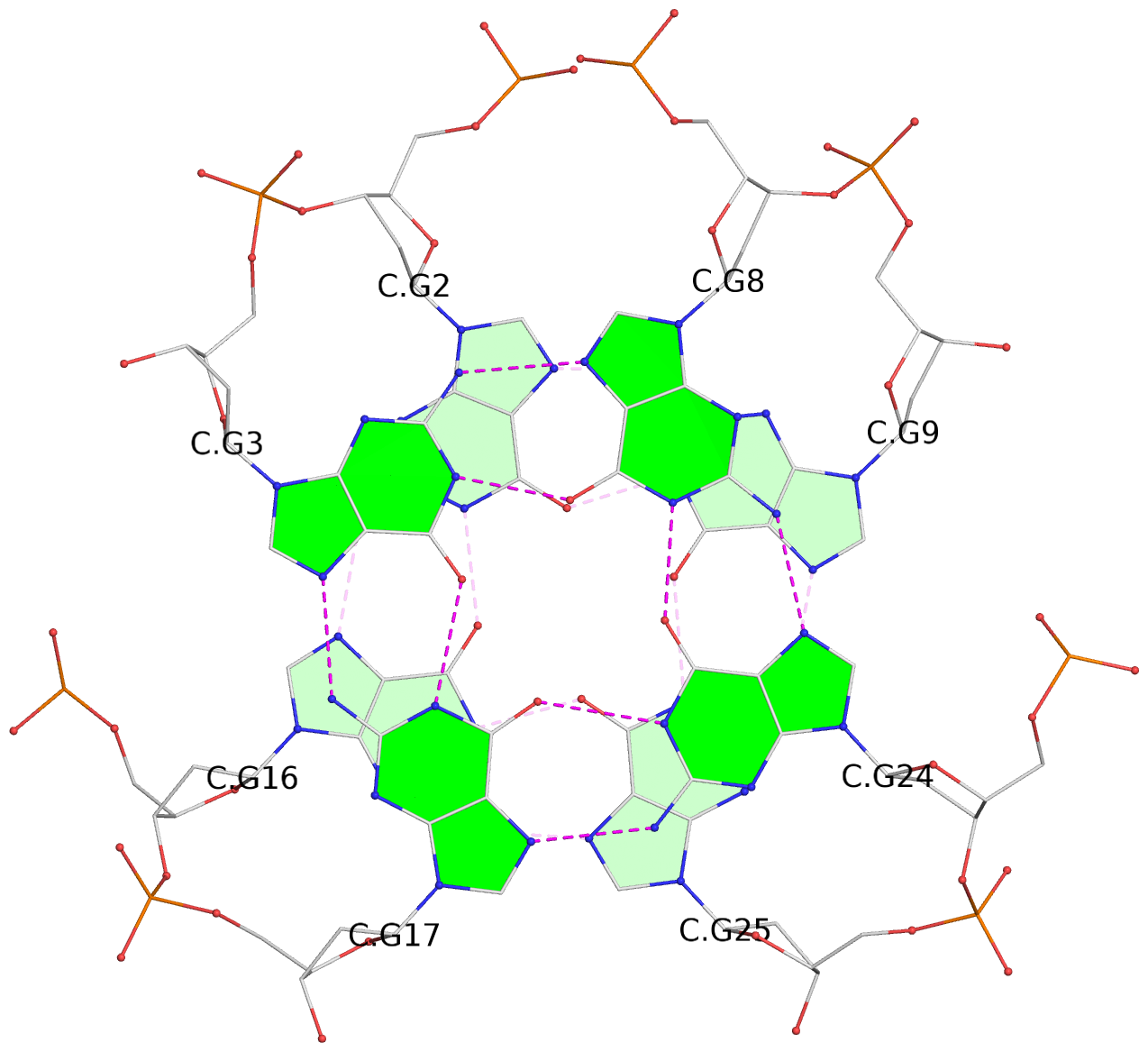
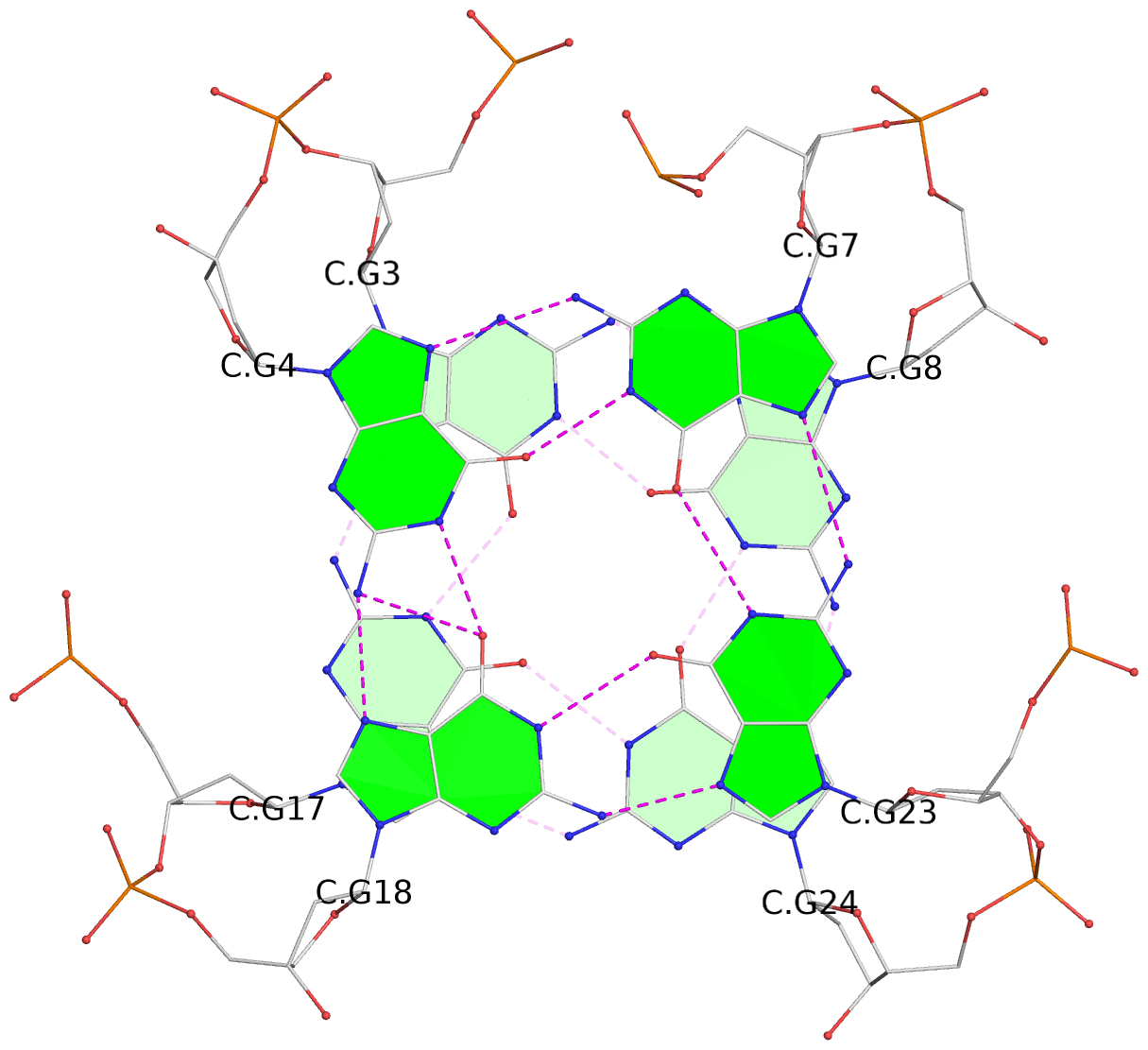
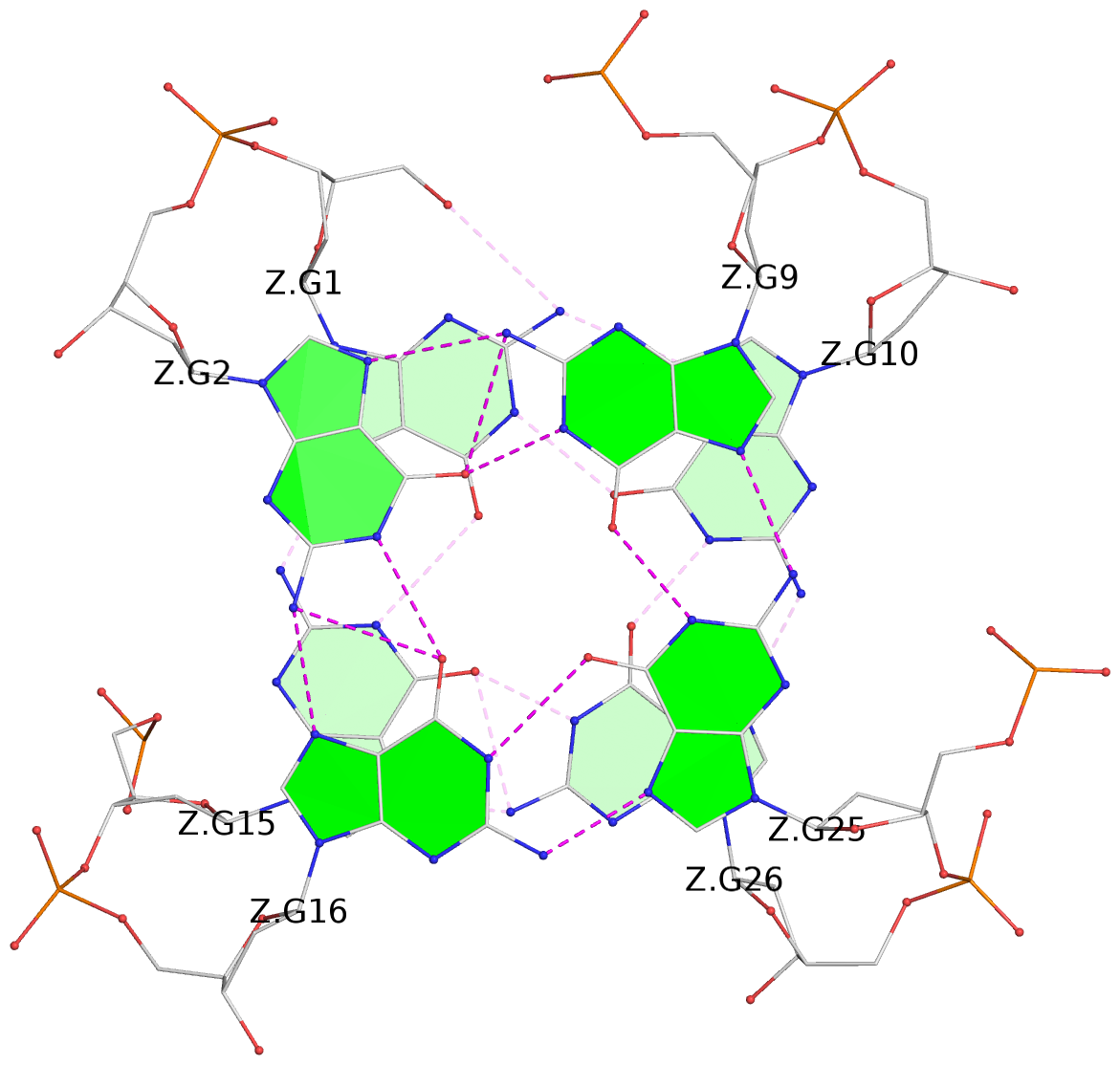
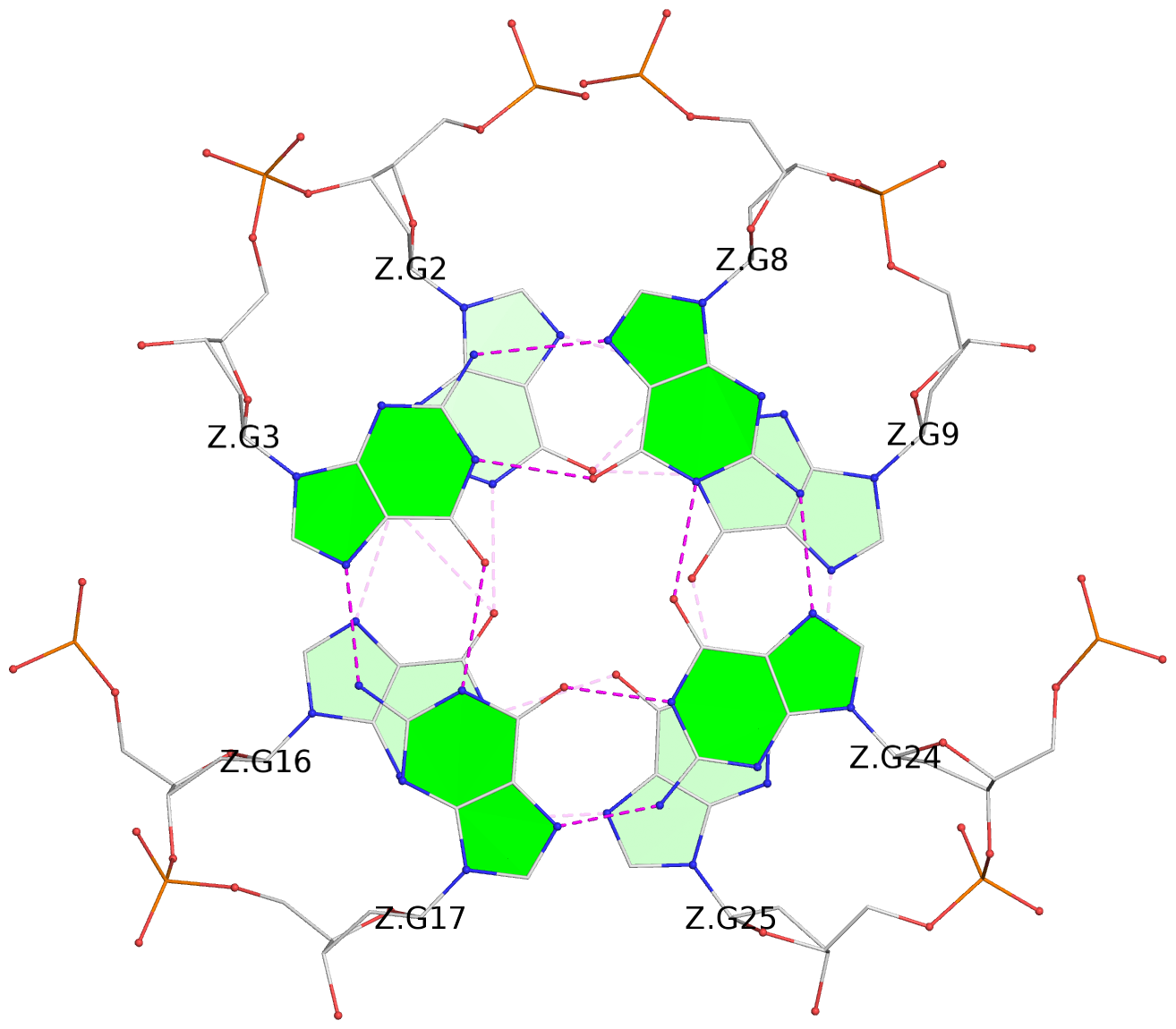
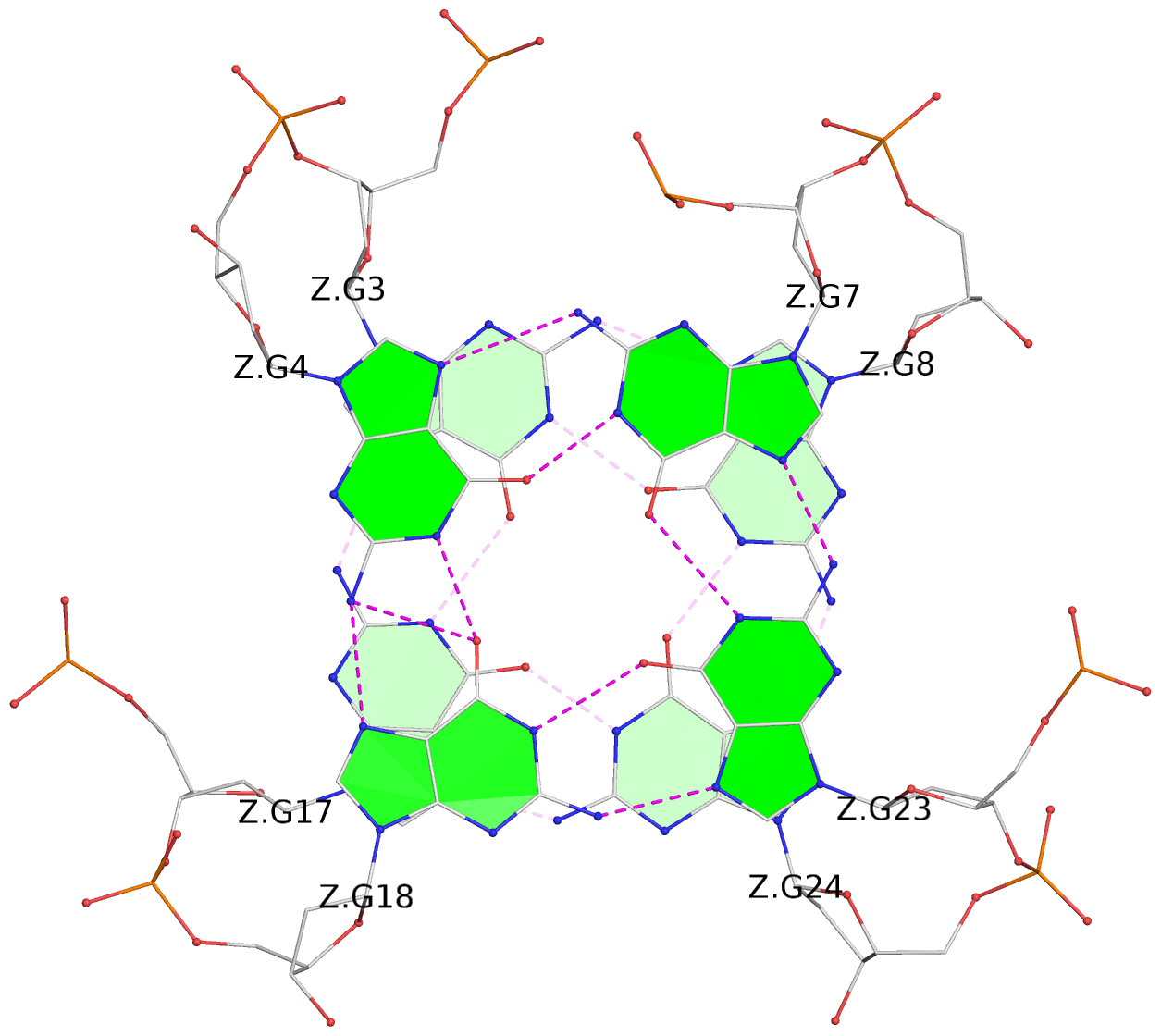
List of 28 G-tetrads
1 glyco-bond=ss-- sugar=---- groove=-w-n planarity=0.227 type=other nts=4 GGGG A.DG1,A.DG15,A.DG26,A.DG10 2 glyco-bond=--ss sugar=---- groove=-w-n planarity=0.140 type=planar nts=4 GGGG A.DG2,A.DG16,A.DG25,A.DG9 3 glyco-bond=ss-- sugar=---- groove=-w-n planarity=0.143 type=planar nts=4 GGGG A.DG3,A.DG17,A.DG24,A.DG8 4 glyco-bond=--ss sugar=3--- groove=-w-n planarity=0.207 type=other nts=4 GGGG A.DG4,A.DG18,A.DG23,A.DG7 5 glyco-bond=ss-- sugar=---. groove=-w-n planarity=0.328 type=bowl nts=4 GGGG B.DG1,B.DG15,B.DG26,B.DG10 6 glyco-bond=--ss sugar=---- groove=-w-n planarity=0.137 type=planar nts=4 GGGG B.DG2,B.DG16,B.DG25,B.DG9 7 glyco-bond=ss-- sugar=---- groove=-w-n planarity=0.099 type=planar nts=4 GGGG B.DG3,B.DG17,B.DG24,B.DG8 8 glyco-bond=--ss sugar=---- groove=-w-n planarity=0.192 type=other nts=4 GGGG B.DG4,B.DG18,B.DG23,B.DG7 9 glyco-bond=ss-- sugar=---. groove=-w-n planarity=0.257 type=other nts=4 GGGG C.DG1,C.DG15,C.DG26,C.DG10 10 glyco-bond=--ss sugar=---- groove=-w-n planarity=0.134 type=planar nts=4 GGGG C.DG2,C.DG16,C.DG25,C.DG9 11 glyco-bond=ss-- sugar=---- groove=-w-n planarity=0.132 type=planar nts=4 GGGG C.DG3,C.DG17,C.DG24,C.DG8 12 glyco-bond=--ss sugar=3-.- groove=-w-n planarity=0.200 type=other nts=4 GGGG C.DG4,C.DG18,C.DG23,C.DG7 13 glyco-bond=ss-- sugar=--.. groove=-w-n planarity=0.425 type=bowl nts=4 GGGG D.DG1,D.DG15,D.DG26,D.DG10 14 glyco-bond=--ss sugar=---- groove=-w-n planarity=0.235 type=other nts=4 GGGG D.DG2,D.DG16,D.DG25,D.DG9 15 glyco-bond=ss-- sugar=---- groove=-w-n planarity=0.147 type=planar nts=4 GGGG D.DG3,D.DG17,D.DG24,D.DG8 16 glyco-bond=--ss sugar=3--- groove=-w-n planarity=0.247 type=other nts=4 GGGG D.DG4,D.DG18,D.DG23,D.DG7 17 glyco-bond=ss-- sugar=--.- groove=-w-n planarity=0.407 type=bowl nts=4 GGGG E.DG1,E.DG15,E.DG26,E.DG10 18 glyco-bond=--ss sugar=---- groove=-w-n planarity=0.219 type=other nts=4 GGGG E.DG2,E.DG16,E.DG25,E.DG9 19 glyco-bond=ss-- sugar=---- groove=-w-n planarity=0.146 type=planar nts=4 GGGG E.DG3,E.DG17,E.DG24,E.DG8 20 glyco-bond=--ss sugar=---- groove=-w-n planarity=0.201 type=other nts=4 GGGG E.DG4,E.DG18,E.DG23,E.DG7 21 glyco-bond=ss-- sugar=---- groove=-w-n planarity=0.248 type=bowl nts=4 GGGG F.DG1,F.DG15,F.DG26,F.DG10 22 glyco-bond=--ss sugar=---- groove=-w-n planarity=0.130 type=planar nts=4 GGGG F.DG2,F.DG16,F.DG25,F.DG9 23 glyco-bond=ss-- sugar=---. groove=-w-n planarity=0.111 type=planar nts=4 GGGG F.DG3,F.DG17,F.DG24,F.DG8 24 glyco-bond=--ss sugar=3--- groove=-w-n planarity=0.196 type=other nts=4 GGGG F.DG4,F.DG18,F.DG23,F.DG7 25 glyco-bond=ss-- sugar=---. groove=-w-n planarity=0.497 type=bowl nts=4 GGGG Z.DG1,Z.DG15,Z.DG26,Z.DG10 26 glyco-bond=--ss sugar=---- groove=-w-n planarity=0.303 type=other nts=4 GGGG Z.DG2,Z.DG16,Z.DG25,Z.DG9 27 glyco-bond=ss-- sugar=---- groove=-w-n planarity=0.169 type=other nts=4 GGGG Z.DG3,Z.DG17,Z.DG24,Z.DG8 28 glyco-bond=--ss sugar=3--- groove=-w-n planarity=0.250 type=other nts=4 GGGG Z.DG4,Z.DG18,Z.DG23,Z.DG7
List of 7 G4-helices
In DSSR, a G4-helix is defined by stacking interactions of G-tetrads, regardless of backbone connectivity, and may contain more than one G4-stem.
Helix#1, 4 G-tetrad layers, INTRA-molecular, with 1 stem
Helix#2, 4 G-tetrad layers, INTRA-molecular, with 1 stem
Helix#3, 4 G-tetrad layers, INTRA-molecular, with 1 stem
Helix#4, 4 G-tetrad layers, INTRA-molecular, with 1 stem
Helix#5, 4 G-tetrad layers, INTRA-molecular, with 1 stem
Helix#6, 4 G-tetrad layers, INTRA-molecular, with 1 stem
Helix#7, 4 G-tetrad layers, INTRA-molecular, with 1 stem
List of 7 G4-stems
In DSSR, a G4-stem is defined as a G4-helix with backbone connectivity. Bulges are also allowed along each of the four strands.