Detailed DSSR results for the G-quadruplex: PDB entry 6isw
Created and maintained by Xiang-Jun Lu <xiangjun@x3dna.org>
Citation: Please cite the NAR'20 DSSR-PyMOL schematics paper and/or the NAR'15 DSSR method paper.
Summary information
- PDB id
- 6isw
- Class
- DNA
- Method
- X-ray (2.3 Å)
- Summary
- Structure of human telomeric DNA with 5-selenophene-modified deoxyuridine at residue 12
- Reference
- Nuthanakanti A, Ahmed I, Khatik SY, Saikrishnan K, Srivatsan SG (2019): "Probing G-quadruplex topologies and recognition concurrently in real time and 3D using a dual-app nucleoside probe." Nucleic Acids Res., 47, 6059-6072. doi: 10.1093/nar/gkz419.
- Abstract
- Comprehensive understanding of structure and recognition properties of regulatory nucleic acid elements in real time and atomic level is highly important to devise efficient therapeutic strategies. Here, we report the establishment of an innovative biophysical platform using a dual-app nucleoside analog, which serves as a common probe to detect and correlate different GQ structures and ligand binding under equilibrium conditions and in 3D by fluorescence and X-ray crystallography techniques. The probe (SedU) is composed of a microenvironment-sensitive fluorophore and an excellent anomalous X-ray scatterer (Se), which is assembled by attaching a selenophene ring at 5-position of 2'-deoxyuridine. SedU incorporated into the loop region of human telomeric DNA repeat fluorescently distinguished subtle differences in GQ topologies and enabled quantify ligand binding to different topologies. Importantly, anomalous X-ray dispersion signal from Se could be used to determine the structure of GQs. As the probe is minimally perturbing, a direct comparison of fluorescence data and crystal structures provided structural insights on how the probe senses different GQ conformations without affecting the native fold. Taken together, our dual-app probe represents a new class of tool that opens up new experimental strategies to concurrently investigate nucleic acid structure and recognition in real time and 3D.
- G4 notes
- 3 G-tetrads, 1 G4 helix, 1 G4 stem, 3(-P-P), parallel(4+0), UUUU
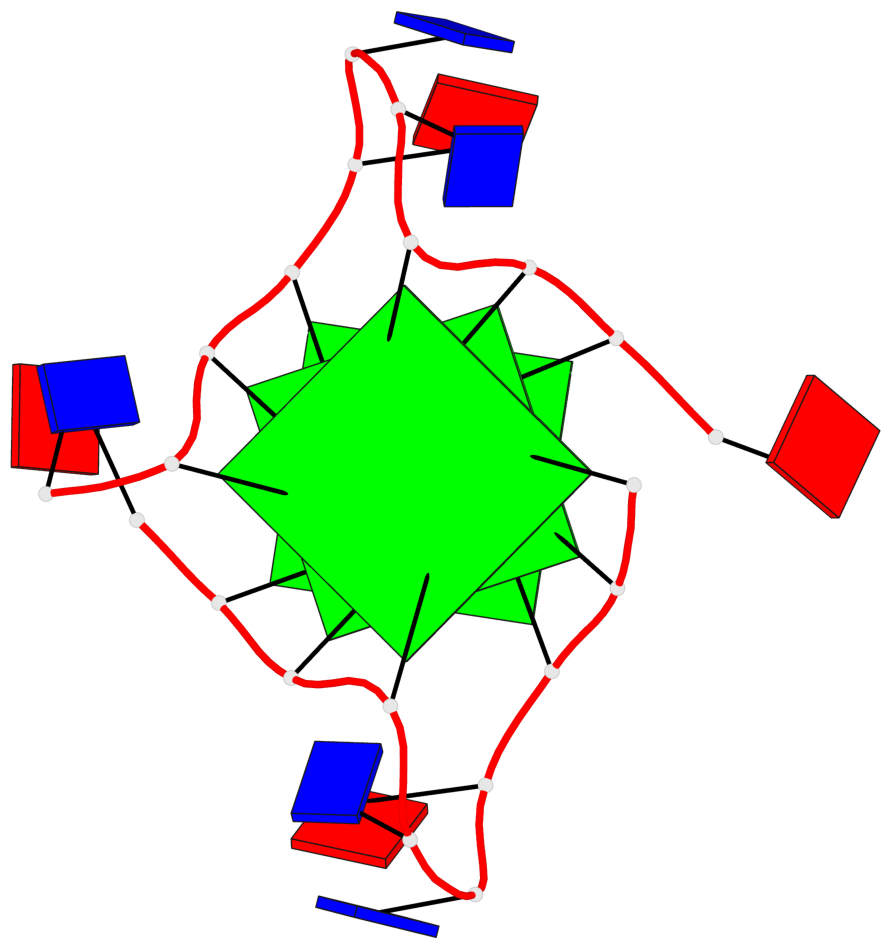
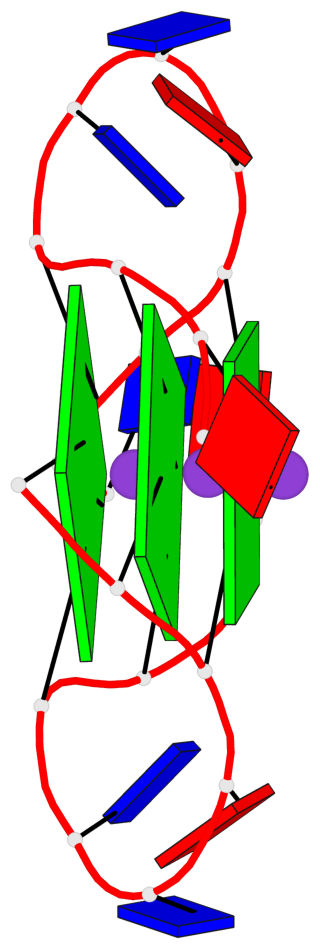
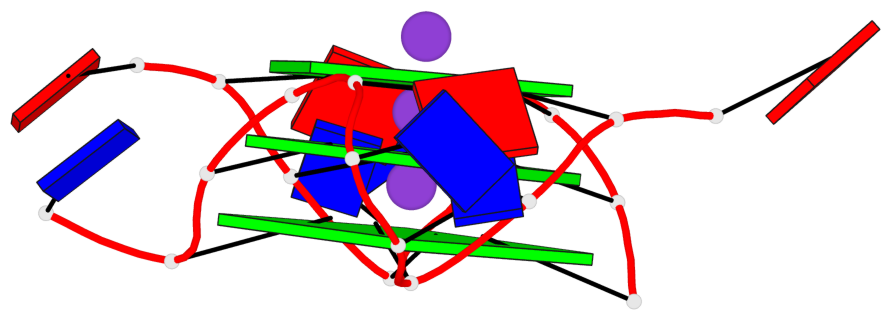
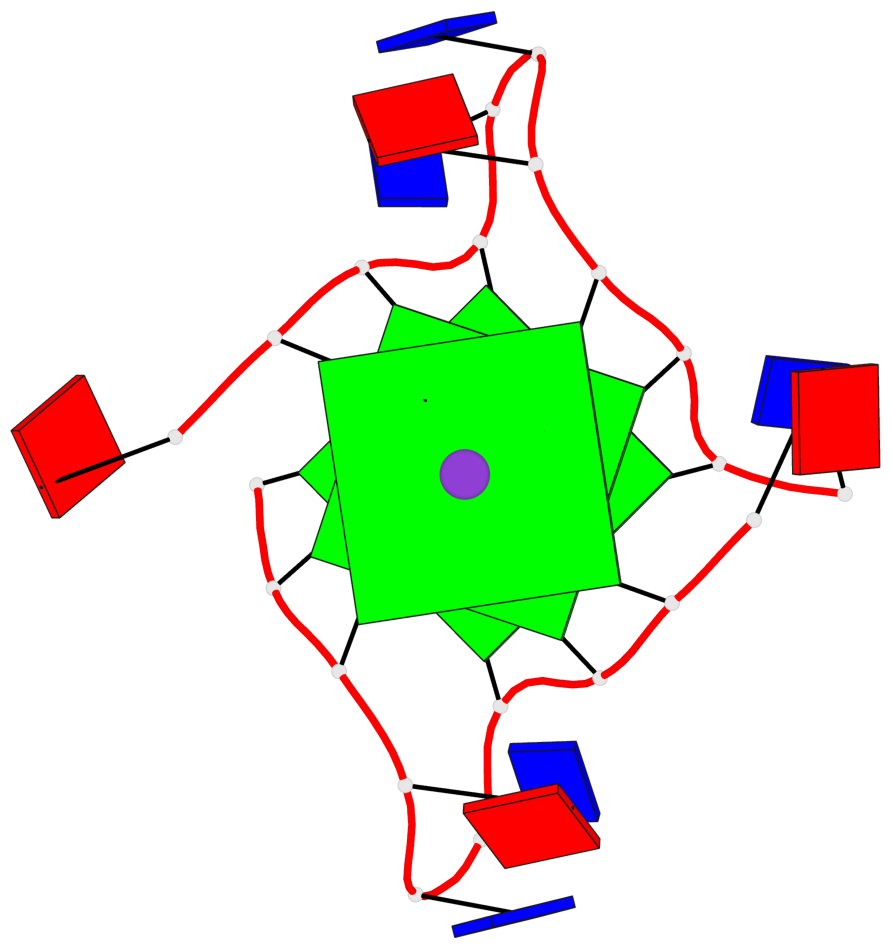
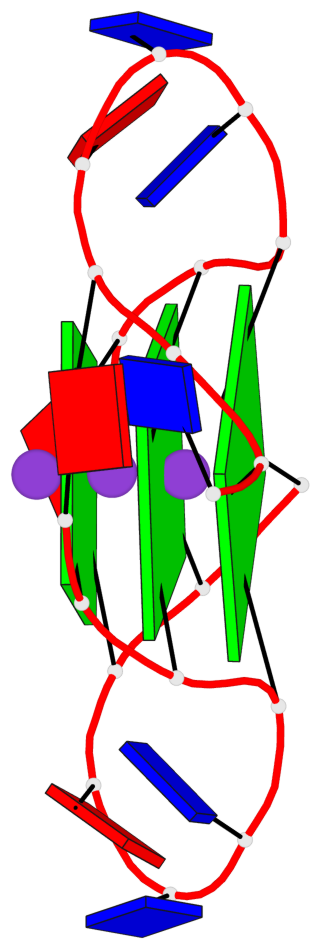
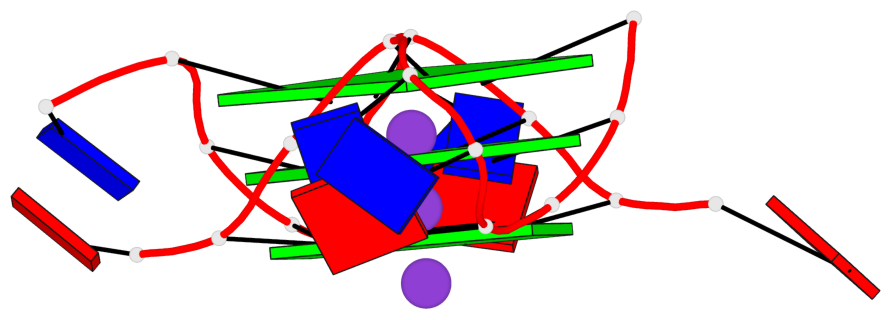
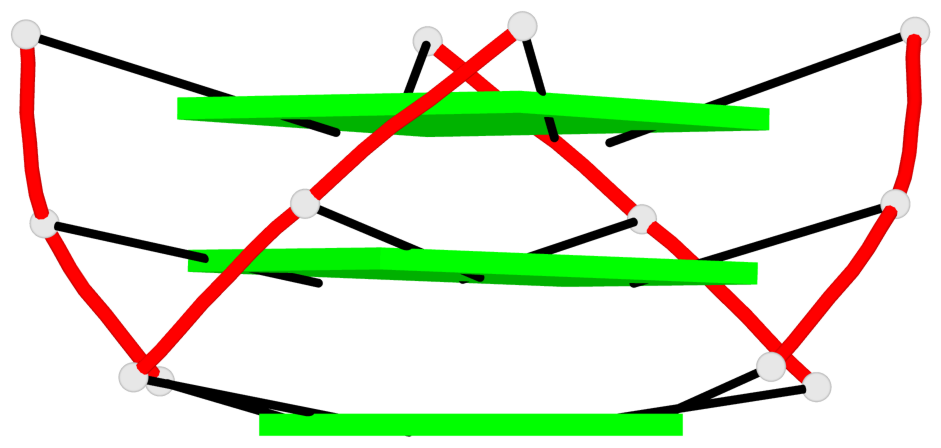
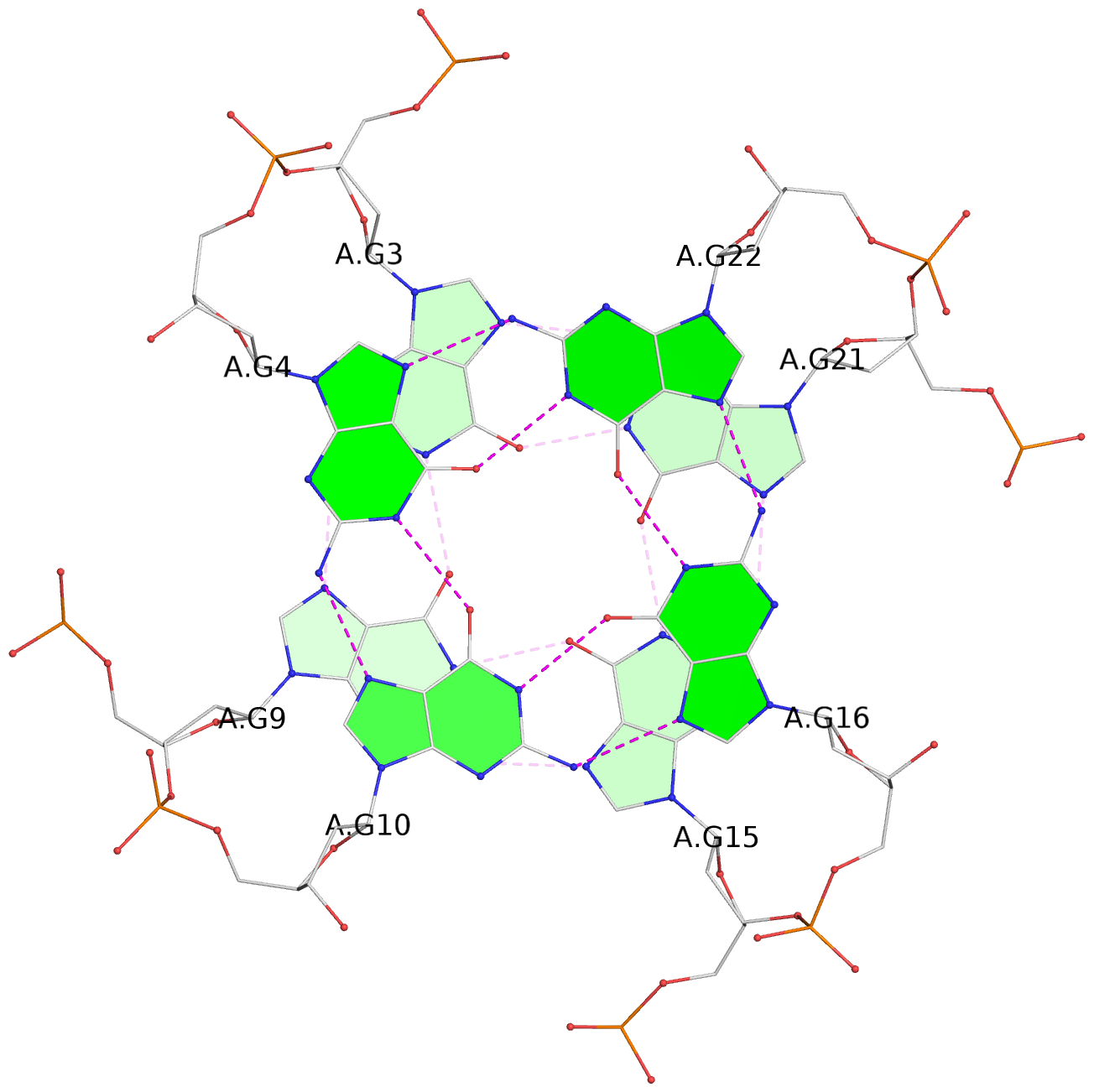
Base-block schematics in six views
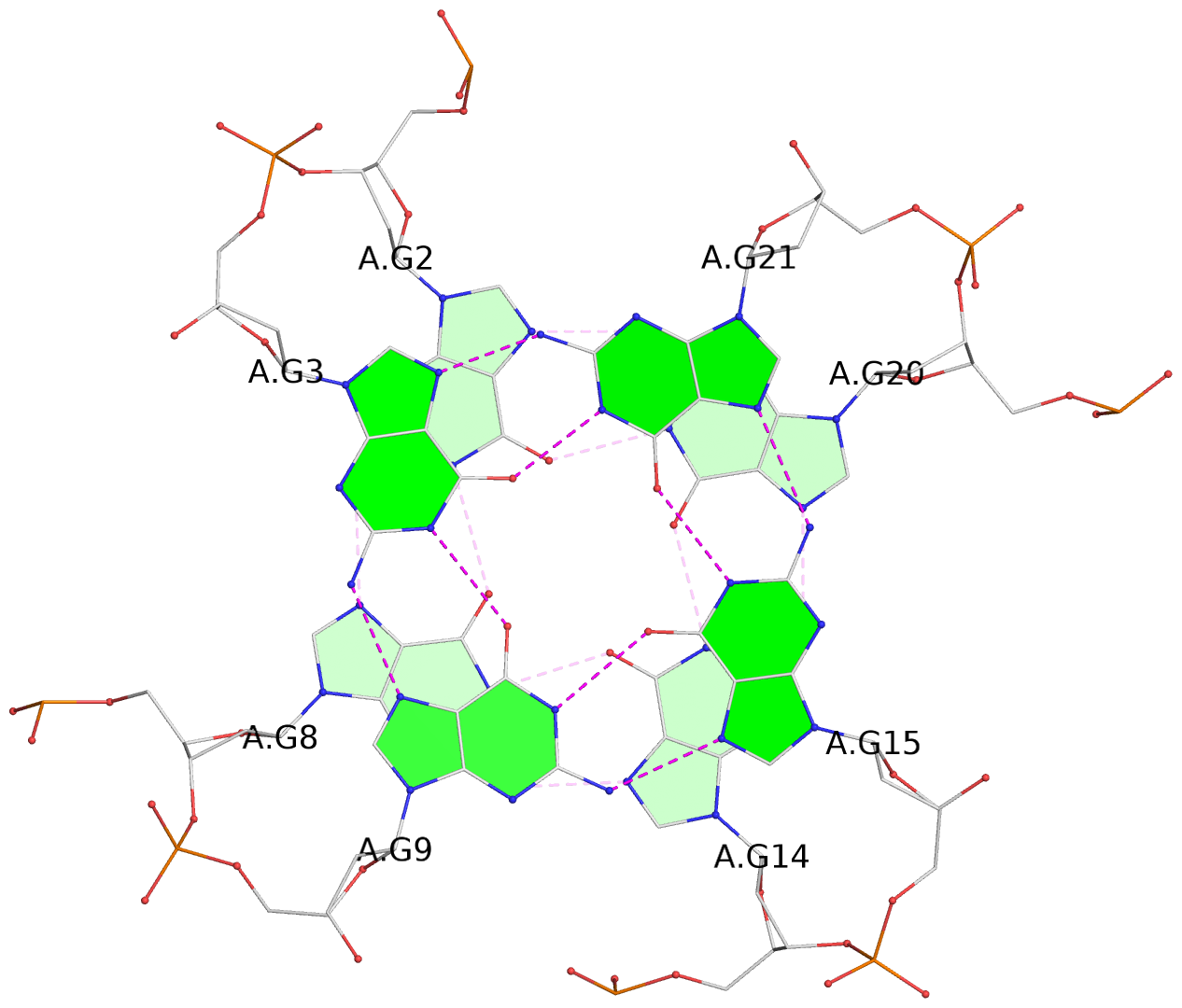
List of 3 G-tetrads
1 glyco-bond=---- sugar=---- groove=---- planarity=0.132 type=planar nts=4 GGGG A.DG2,A.DG8,A.DG14,A.DG20 2 glyco-bond=---- sugar=---- groove=---- planarity=0.279 type=bowl nts=4 GGGG A.DG3,A.DG9,A.DG15,A.DG21 3 glyco-bond=---- sugar=---- groove=---- planarity=0.410 type=bowl nts=4 GGGG A.DG4,A.DG10,A.DG16,A.DG22
List of 1 G4-helix
In DSSR, a G4-helix is defined by stacking interactions of G-tetrads, regardless of backbone connectivity, and may contain more than one G4-stem.
Helix#1, 3 G-tetrad layers, INTRA-molecular, with 1 stem
List of 1 G4-stem
In DSSR, a G4-stem is defined as a G4-helix with backbone connectivity. Bulges are also allowed along each of the four strands.