Detailed DSSR results for the G-quadruplex: PDB entry 6jjf
Created and maintained by Xiang-Jun Lu <xiangjun@x3dna.org>
Citation: Please cite the NAR'20 DSSR-PyMOL schematics paper and/or the NAR'15 DSSR method paper.
Summary information
- PDB id
- 6jjf
- Class
- DNA
- Method
- X-ray (1.47 Å)
- Summary
- Crystal structure of a two-quartet DNA mixed-parallel-antiparallel g-quadruplex
- Reference
- Zhang Y, El Omari K, Duman R, Liu S, Haider S, Wagner A, Parkinson GN, Wei D (2020): "Native de novo structural determinations of non-canonical nucleic acid motifs by X-ray crystallography at long wavelengths." Nucleic Acids Res., 48, 9886-9898. doi: 10.1093/nar/gkaa439.
- Abstract
- Obtaining phase information remains a formidable challenge for nucleic acid structure determination. The introduction of an X-ray synchrotron beamline designed to be tunable to long wavelengths at Diamond Light Source has opened the possibility to native de novo structure determinations by the use of intrinsic scattering elements. This provides opportunities to overcome the limitations of introducing modifying nucleotides, often required to derive phasing information. In this paper, we build on established methods to generate new tools for nucleic acid structure determinations. We report on the use of (i) native intrinsic potassium single-wavelength anomalous dispersion methods (K-SAD), (ii) use of anomalous scattering elements integral to the crystallization buffer (extrinsic cobalt and intrinsic potassium ions), (iii) extrinsic bromine and intrinsic phosphorus SAD to solve complex nucleic acid structures. Using the reported methods we solved the structures of (i) Pseudorabies virus (PRV) RNA G-quadruplex and ligand complex, (ii) PRV DNA G-quadruplex, and (iii) an i-motif of human telomeric sequence. Our results highlight the utility of using intrinsic scattering as a pathway to solve and determine non-canonical nucleic acid motifs and reveal the variability of topology, influence of ligand binding, and glycosidic angle rearrangements seen between RNA and DNA G-quadruplexes of the same sequence.
- G4 notes
- 4 G-tetrads, 1 G4 helix, 2 G4 stems, 1 G4 coaxial stack, (2+2), UDUD; parallel(4+0), UUUU, coaxial interfaces: mixed
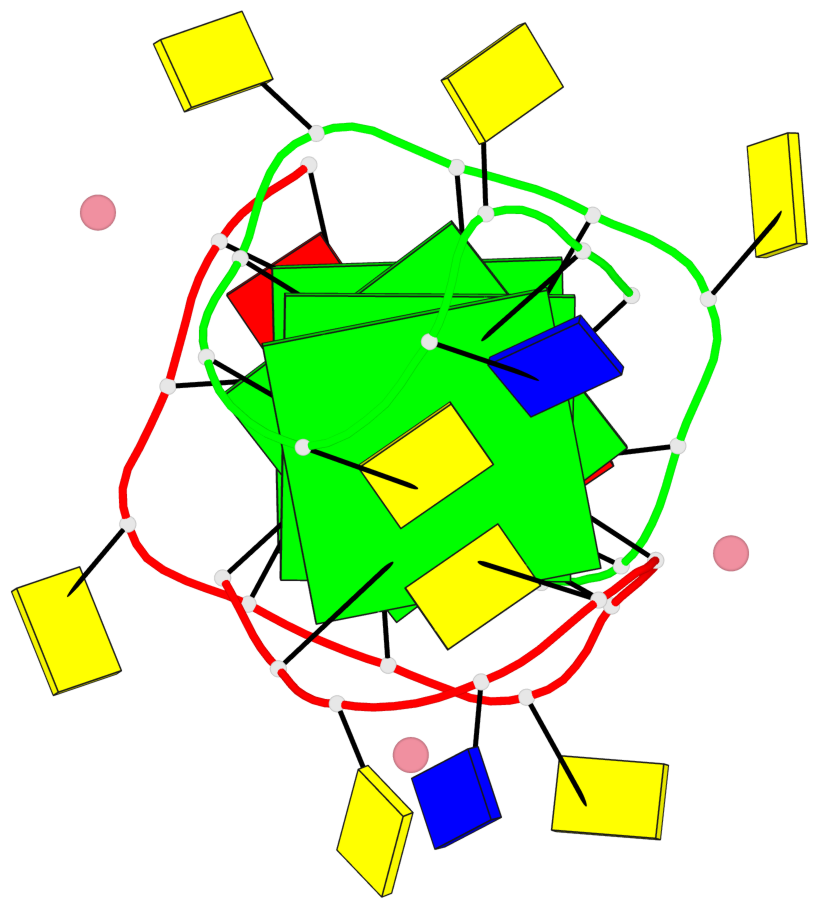
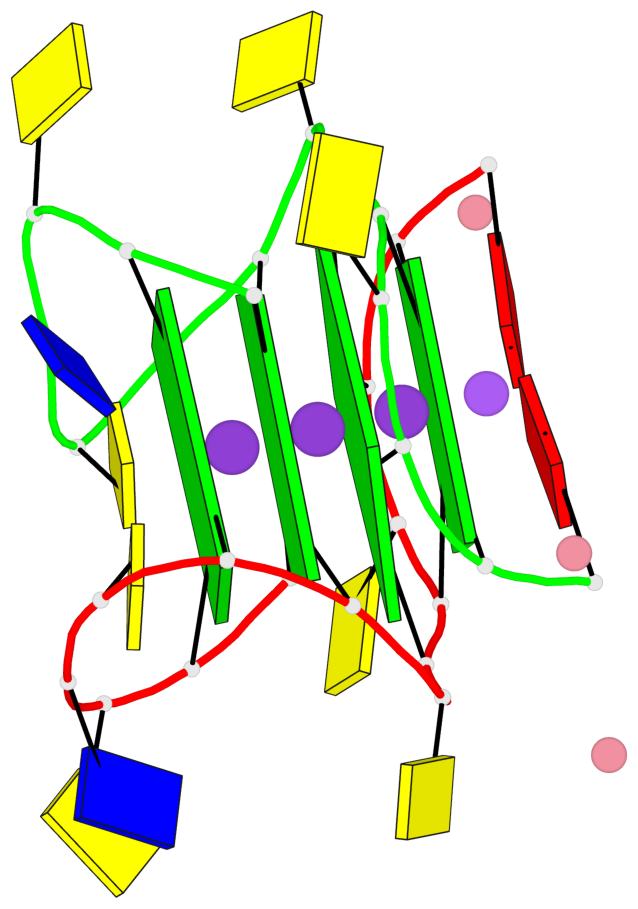
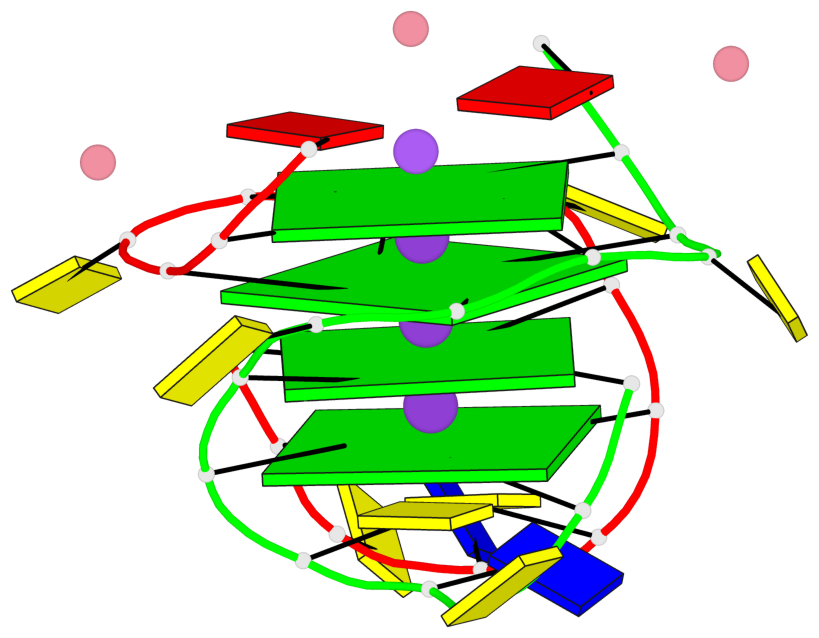
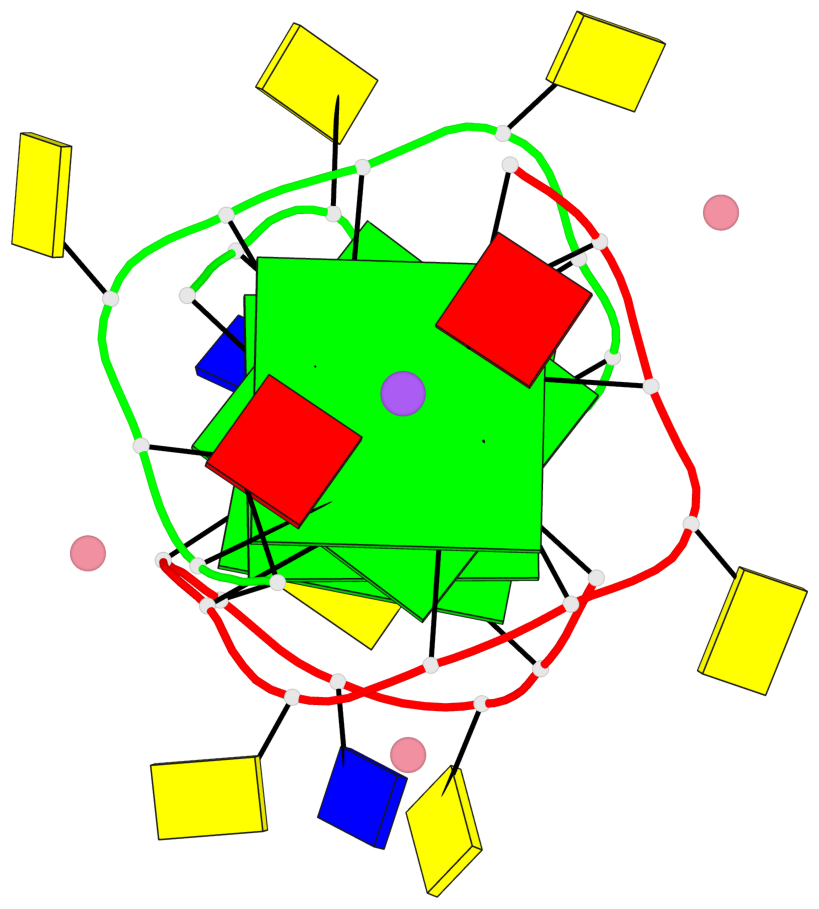
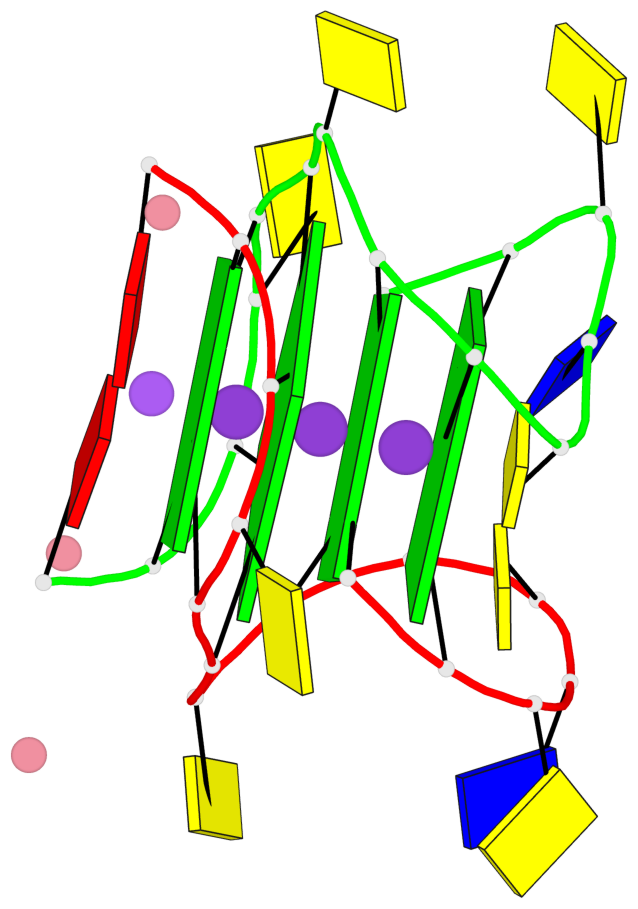
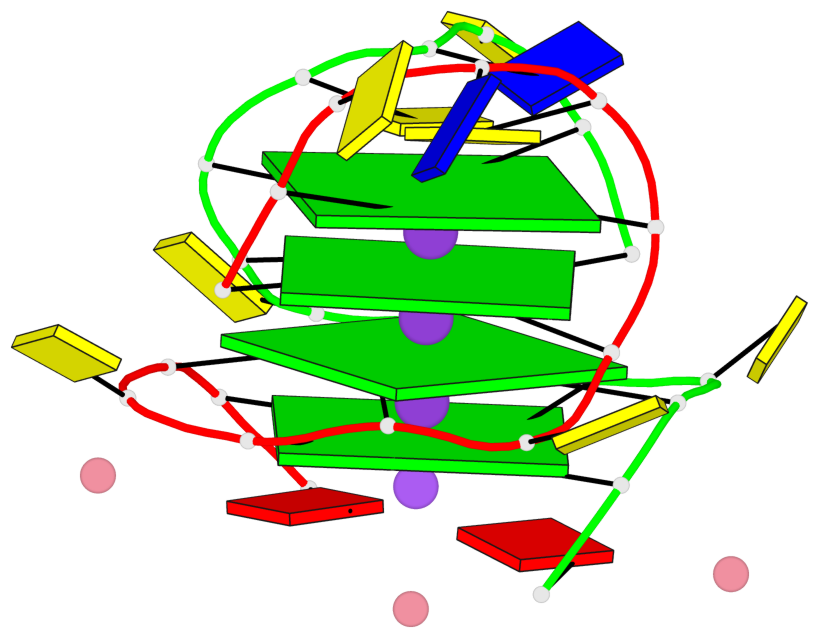
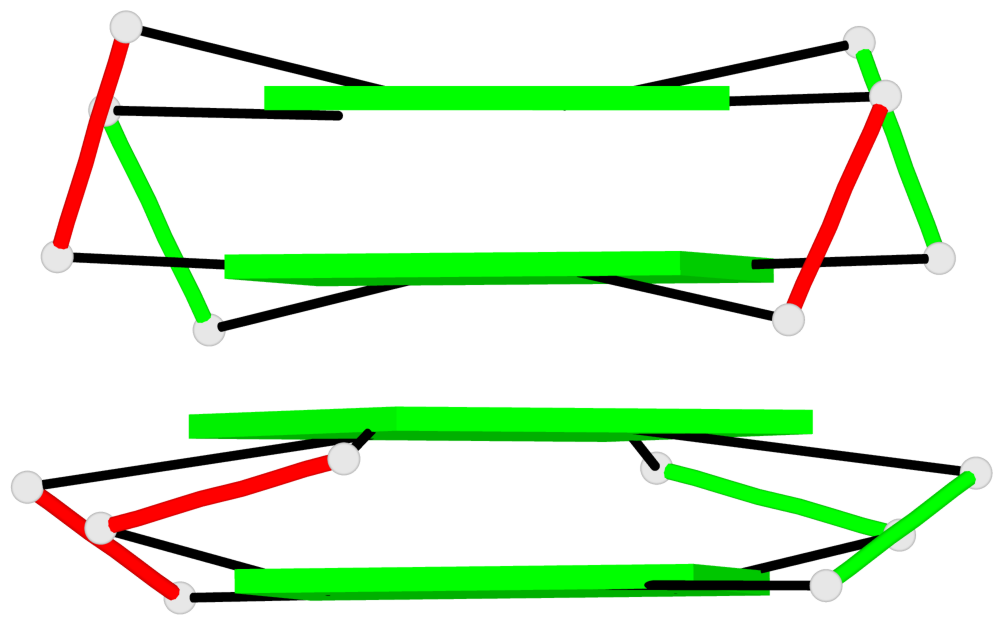
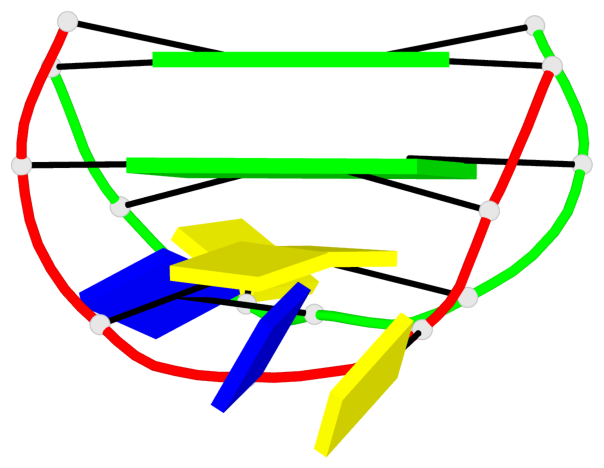
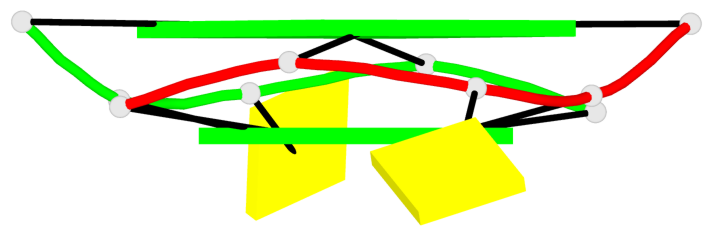
Base-block schematics in six views
List of 4 G-tetrads
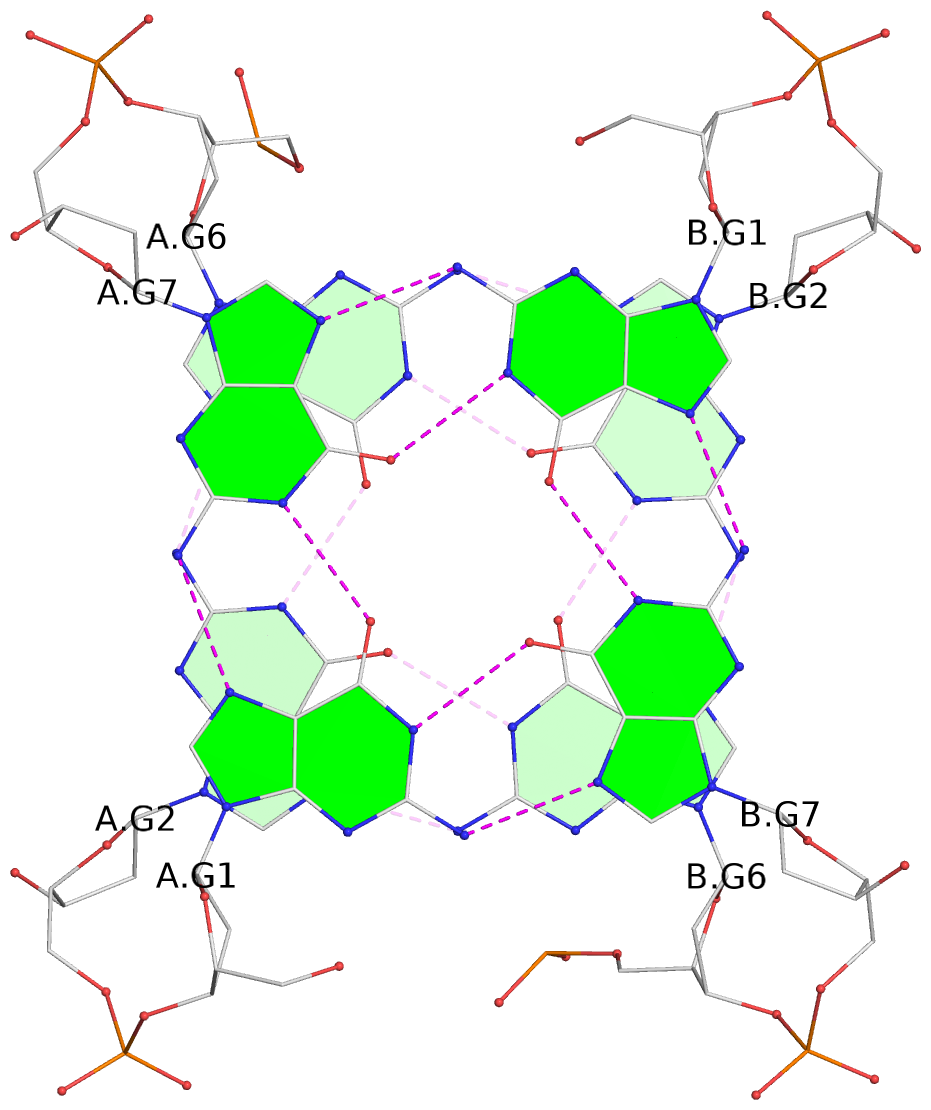
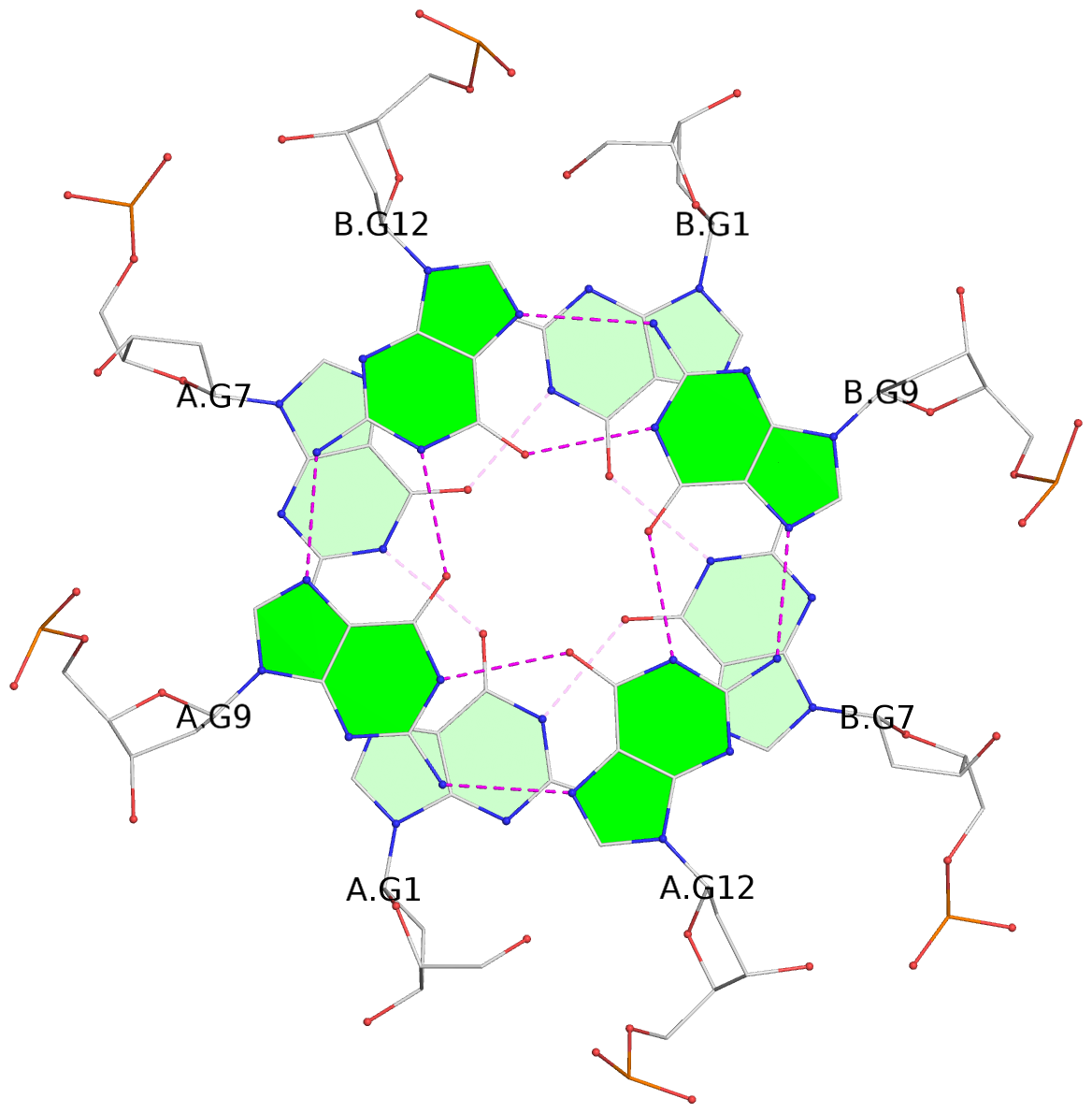
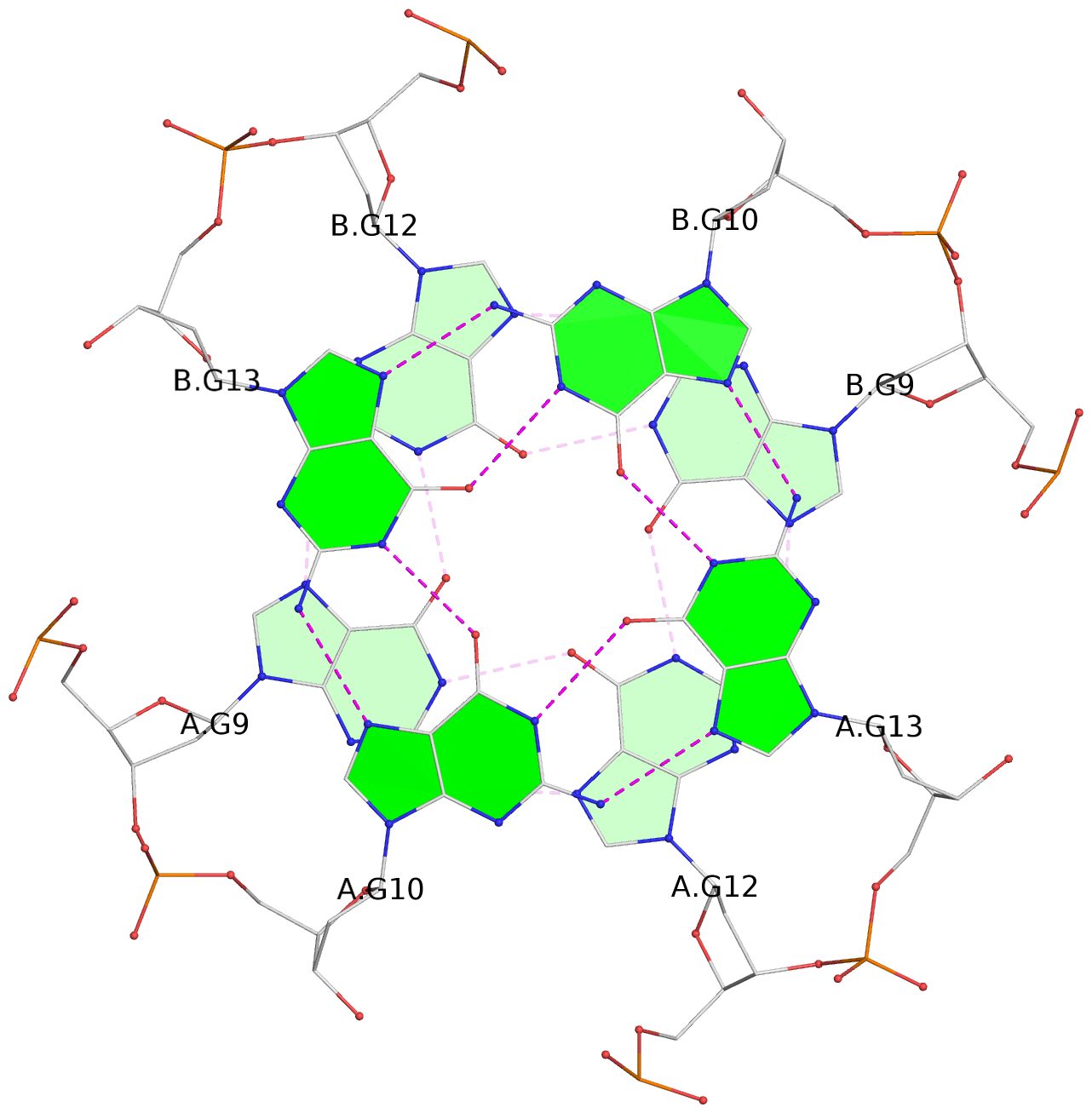
1 glyco-bond=s-s- sugar=---- groove=wnwn planarity=0.186 type=saddle nts=4 GGGG A.DG1,A.DG7,B.DG1,B.DG7 2 glyco-bond=-s-s sugar=---- groove=wnwn planarity=0.254 type=bowl-2 nts=4 GGGG A.DG2,A.DG6,B.DG2,B.DG6 3 glyco-bond=---- sugar=---- groove=---- planarity=0.182 type=other nts=4 GGGG A.DG9,A.DG12,B.DG9,B.DG12 4 glyco-bond=---- sugar=---- groove=---- planarity=0.253 type=other nts=4 GGGG A.DG10,A.DG13,B.DG10,B.DG13
List of 1 G4-helix
In DSSR, a G4-helix is defined by stacking interactions of G-tetrads, regardless of backbone connectivity, and may contain more than one G4-stem.
Helix#1, 4 G-tetrad layers, inter-molecular, with 2 stems
List of 2 G4-stems
In DSSR, a G4-stem is defined as a G4-helix with backbone connectivity. Bulges are also allowed along each of the four strands.
Stem#1, 2 G-tetrad layers, 2 loops, inter-molecular, UDUD, anti-parallel, (2+2)
Stem#2, 2 G-tetrad layers, 2 loops, inter-molecular, UUUU, parallel, parallel(4+0)
List of 1 G4 coaxial stack
1 G4 helix#1 contains 2 G4 stems: [#1,#2] [mixed]