Detailed DSSR results for the G-quadruplex: PDB entry 6q6r
Created and maintained by Xiang-Jun Lu <xiangjun@x3dna.org>
Citation: Please cite the NAR'20 DSSR-PyMOL schematics paper and/or the NAR'15 DSSR method paper.
Summary information
- PDB id
- 6q6r
- Class
- DNA binding protein
- Method
- X-ray (1.5 Å)
- Summary
- Recognition of different base tetrads by rhau: x-ray crystal structure of g4 recognition motif bound to the 3-end tetrad of a DNA g-quadruplex
- Reference
- Heddi B, Cheong VV, Schmitt E, Mechulam Y, Phan AT (2020): "Recognition of different base tetrads by RHAU (DHX36): X-ray crystal structure of the G4 recognition motif bound to the 3'-end tetrad of a DNA G-quadruplex." J.Struct.Biol., 209, 107399. doi: 10.1016/j.jsb.2019.10.001.
- Abstract
- G-quadruplexes (G4) are secondary structures of nucleic acids that can form in cells and have diverse biological functions. Several biologically important proteins interact with G-quadruplexes, of which RHAU (or DHX36) - a helicase from the DEAH-box superfamily, was shown to bind and unwind G-quadruplexes efficiently. We report a X-ray co-crystal structure at 1.5 Å resolution of an N-terminal fragment of RHAU bound to an exposed tetrad of a parallel-stranded G-quadruplex. The RHAU peptide folds into an L-shaped α-helix, and binds to a G-quadruplex through π-stacking and electrostatic interactions. X-ray crystal structure of our complex identified key amino acid residues important for G-quadruplex-peptide binding interaction at the 3'-end G•G•G•G tetrad. Together with previous solution and crystal structures of RHAU bound to the 5'-end G•G•G•G and G•G•A•T tetrads, our crystal structure highlights the occurrence of a robust G-quadruplex recognition motif within RHAU that can adapt to different accessible tetrads.
- G4 notes
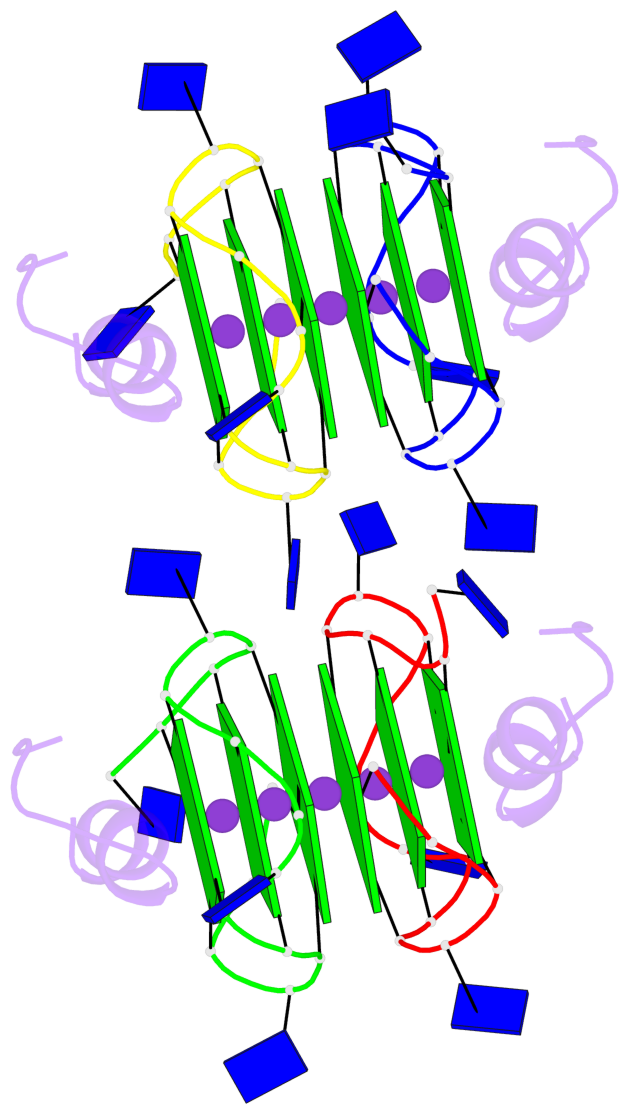
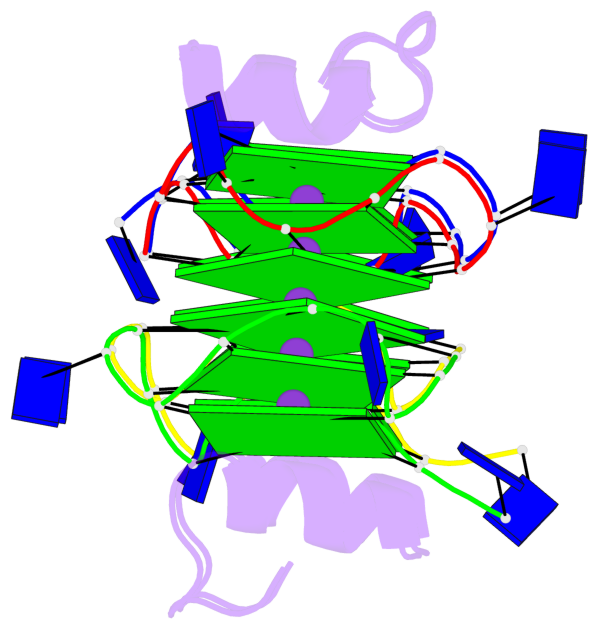
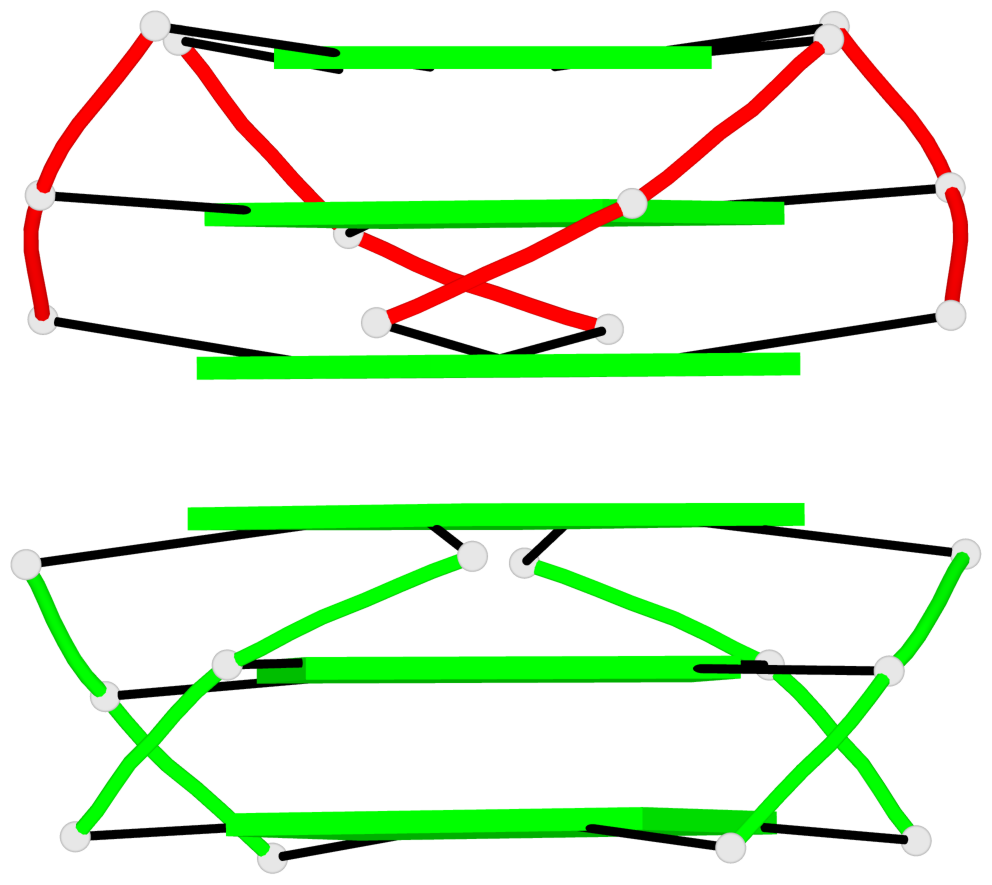
- 12 G-tetrads, 2 G4 helices, 4 G4 stems, 2 G4 coaxial stacks, 3(-P-P-P), parallel(4+0), UUUU, coaxial interfaces: 5'/5'
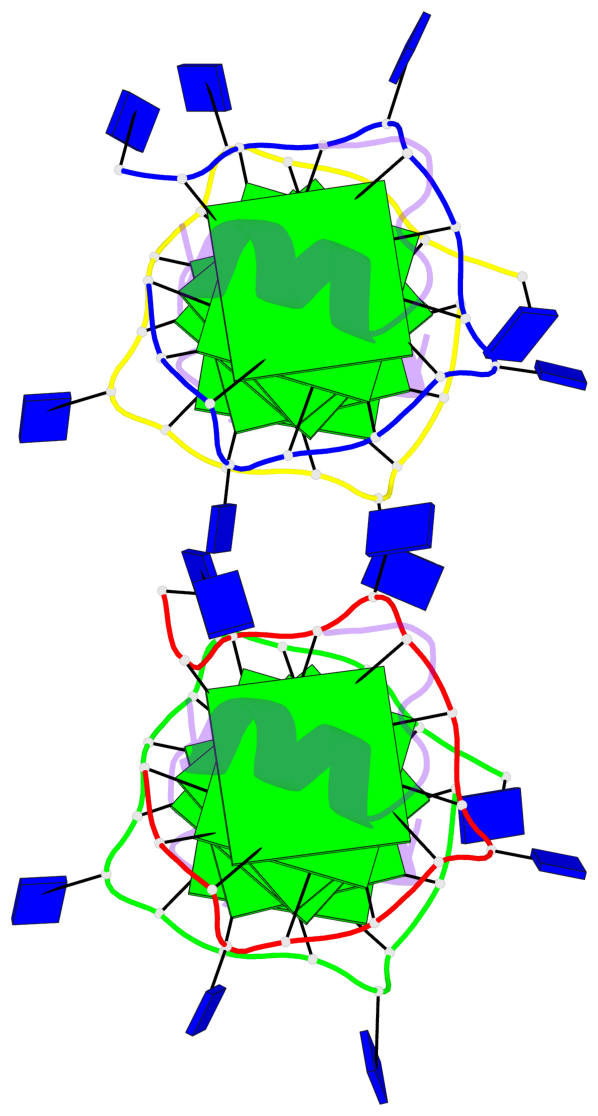
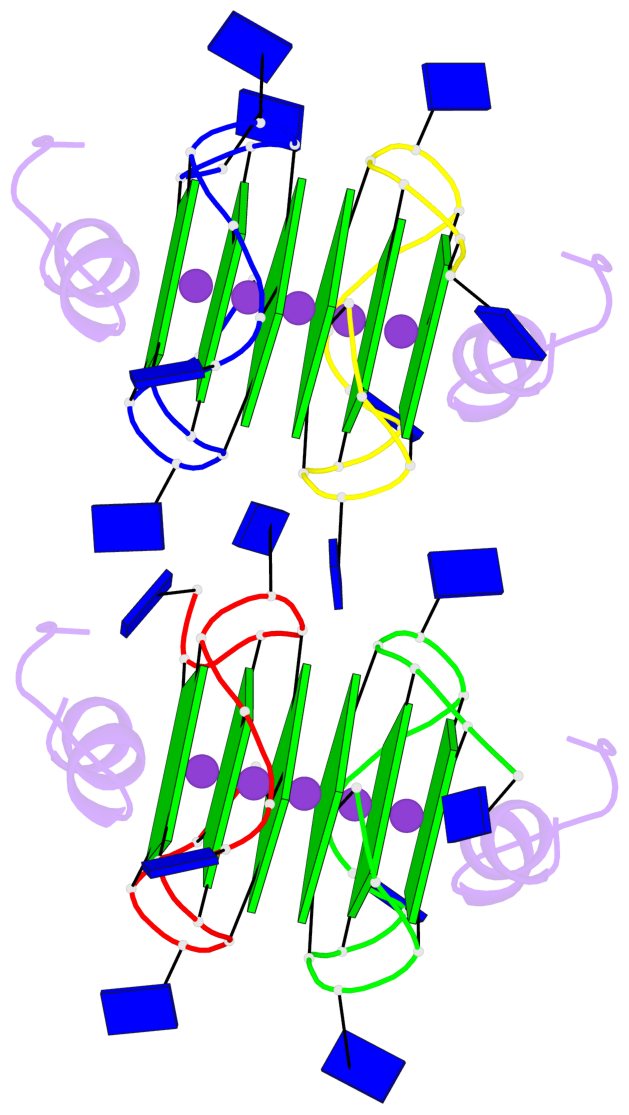
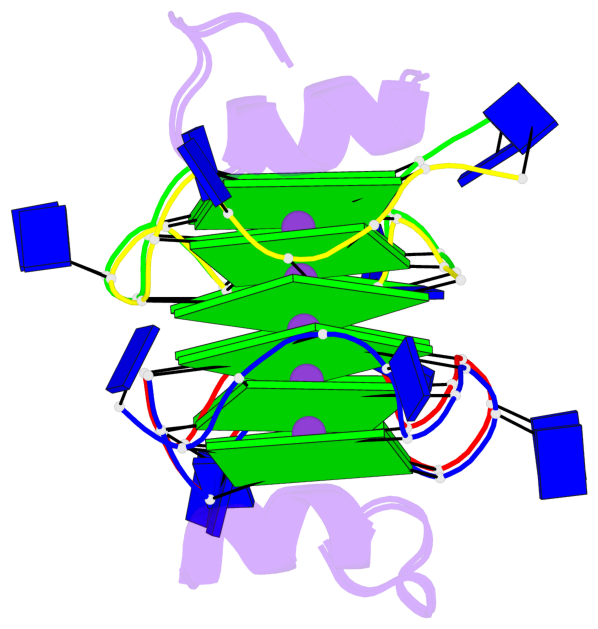
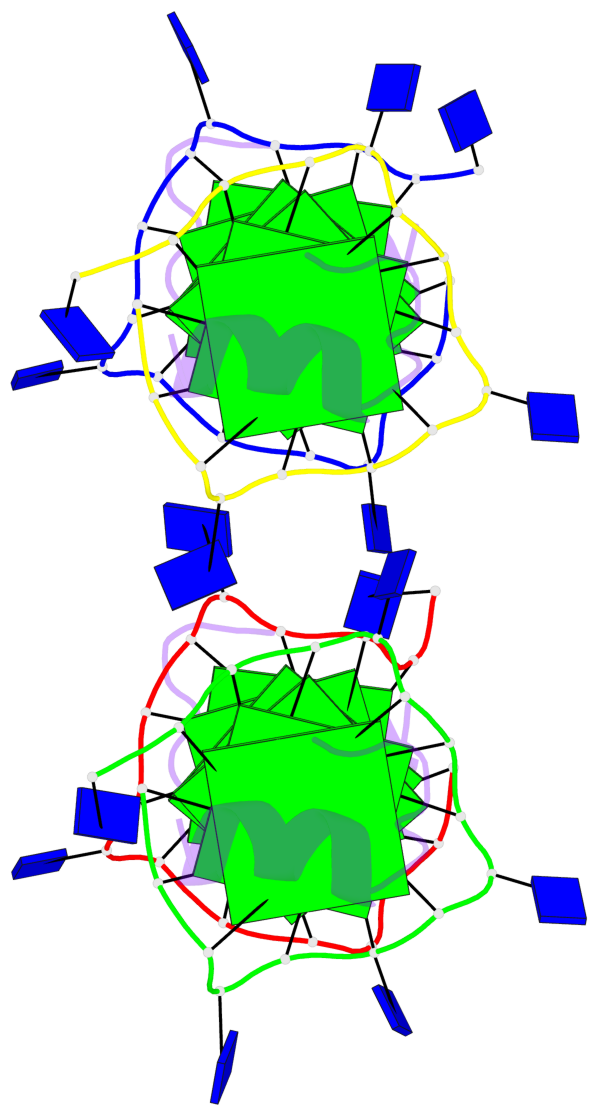
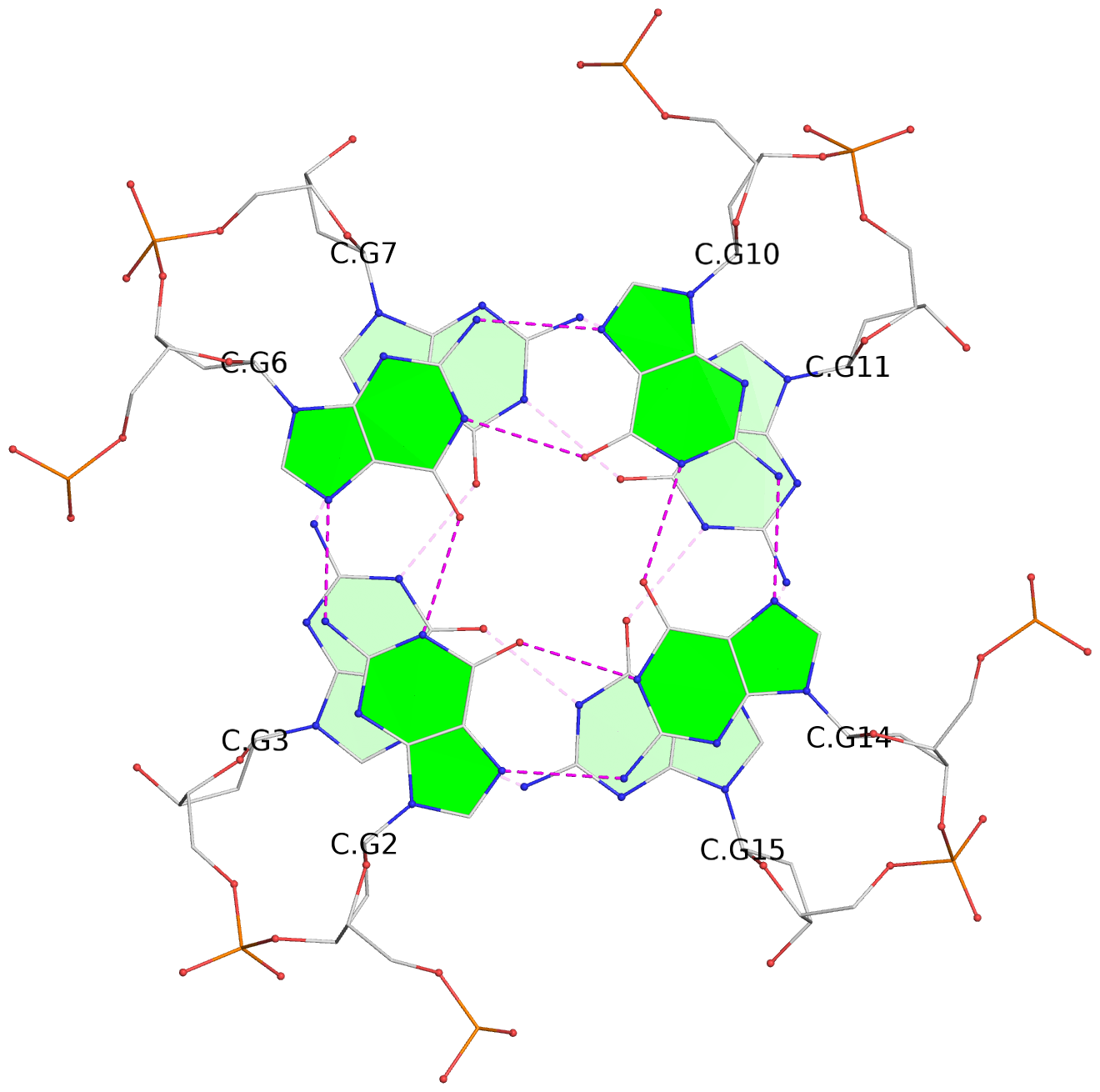
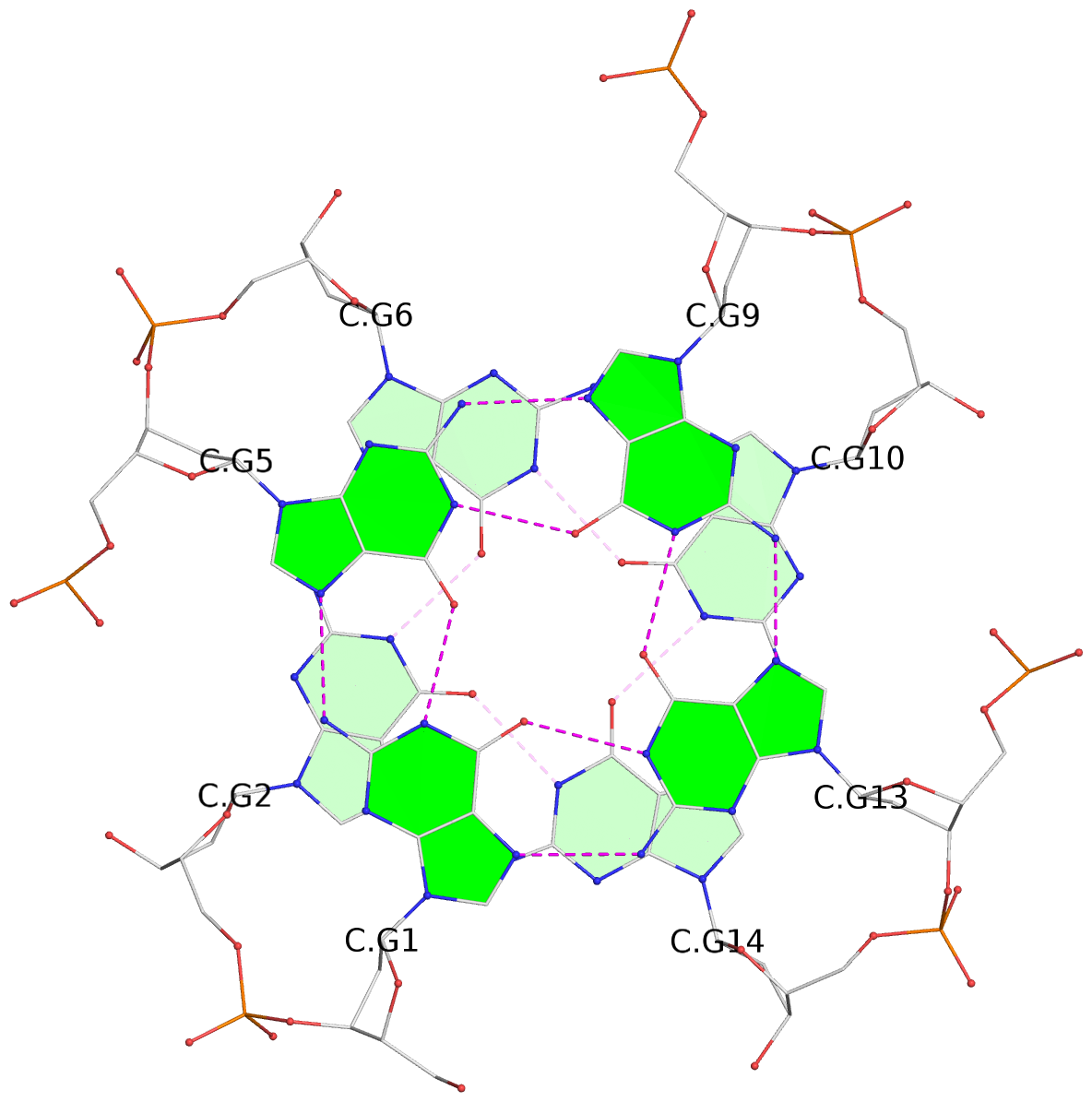
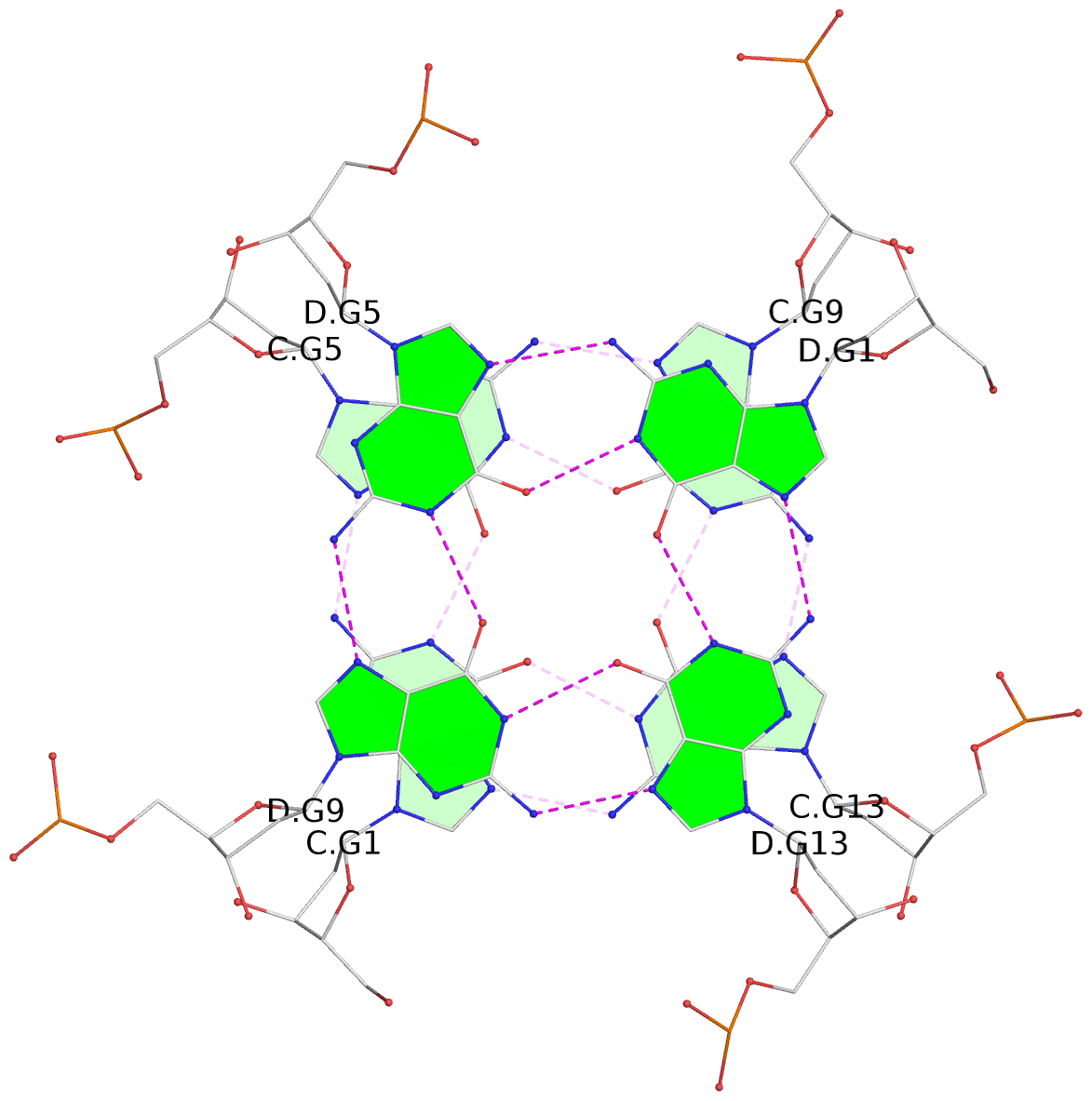
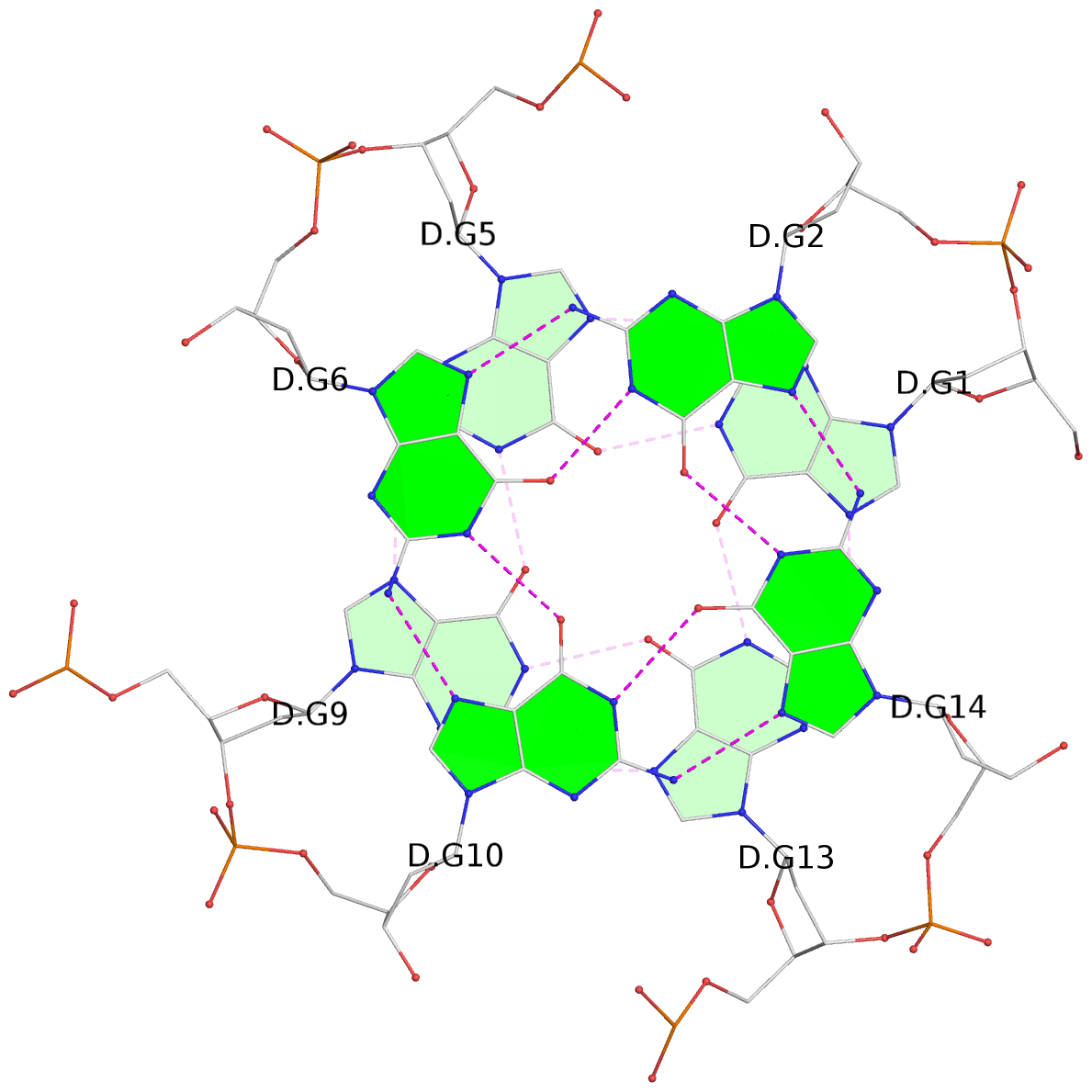
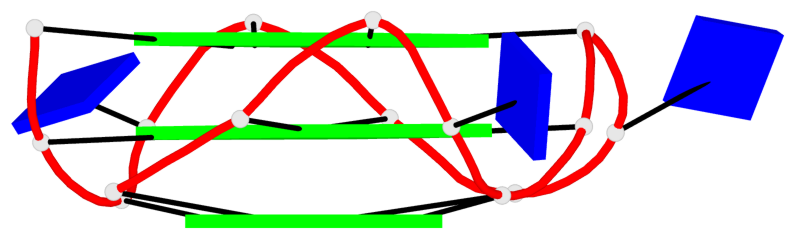
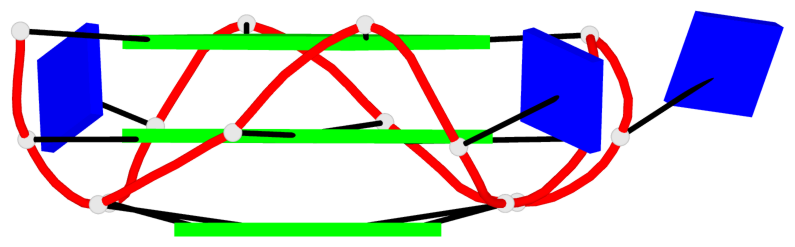
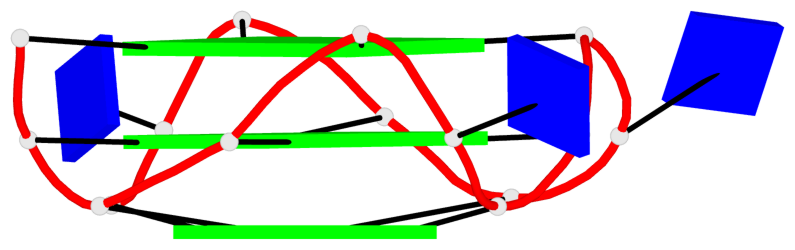
Base-block schematics in six views
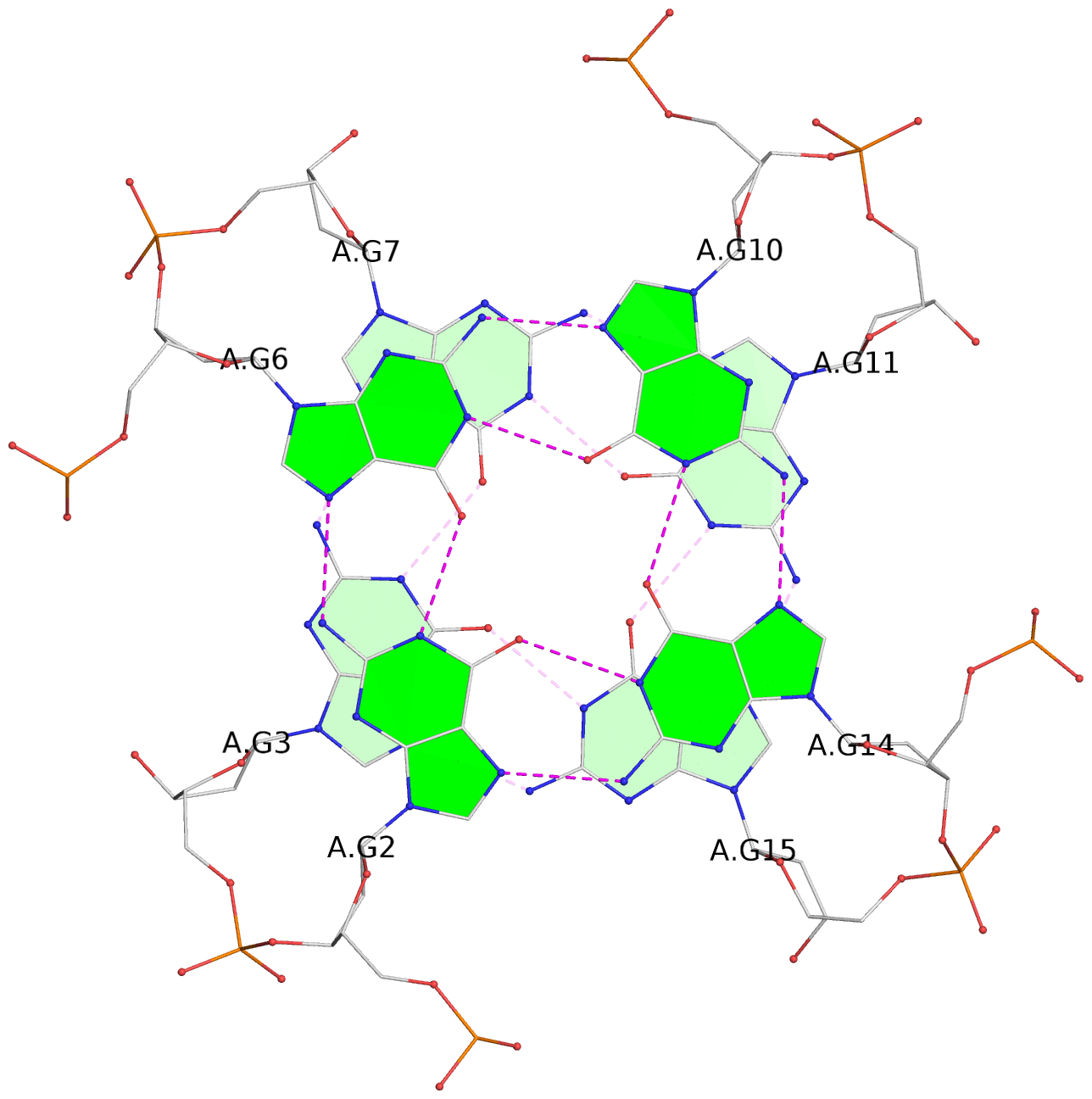
List of 12 G-tetrads
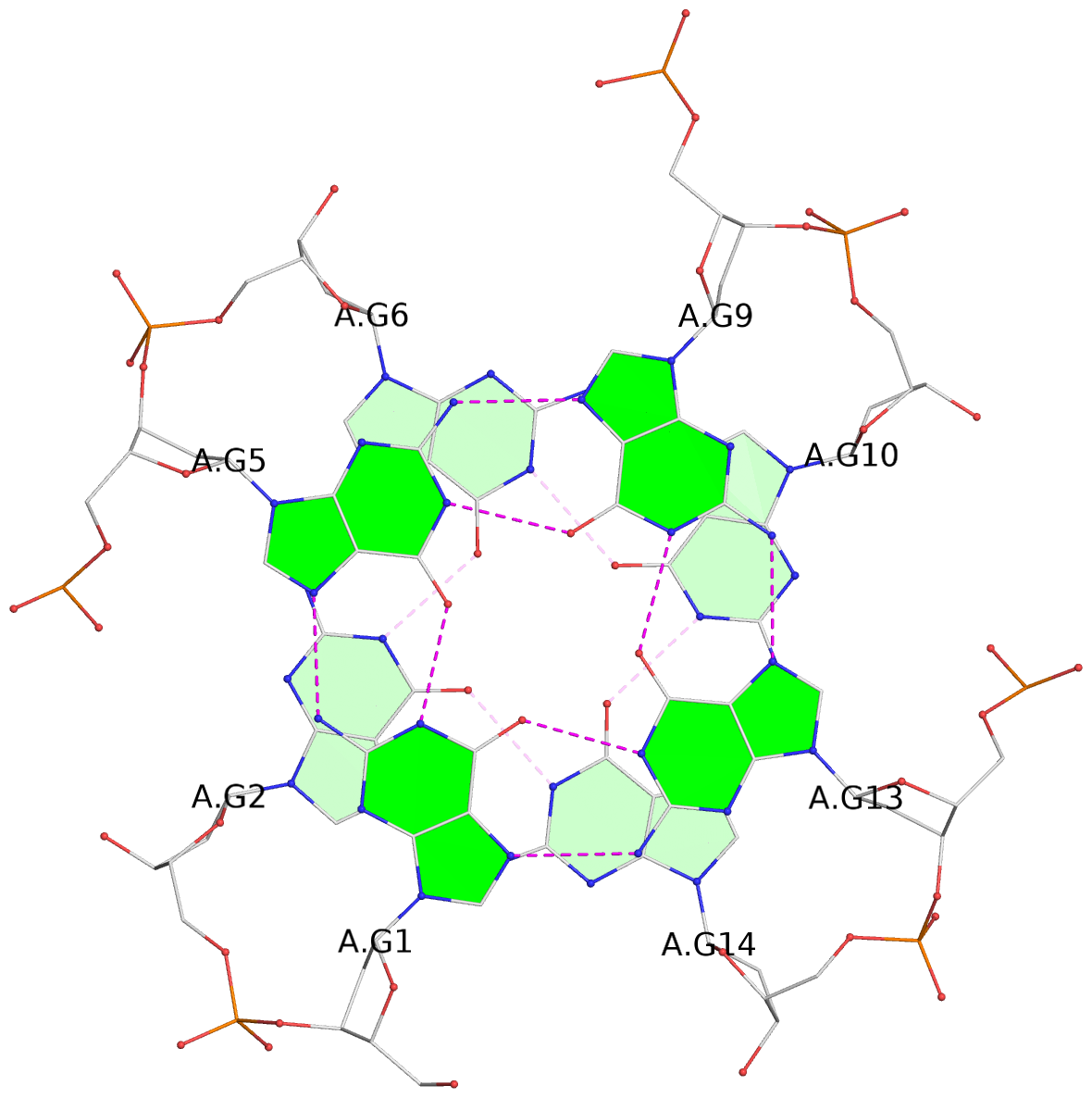
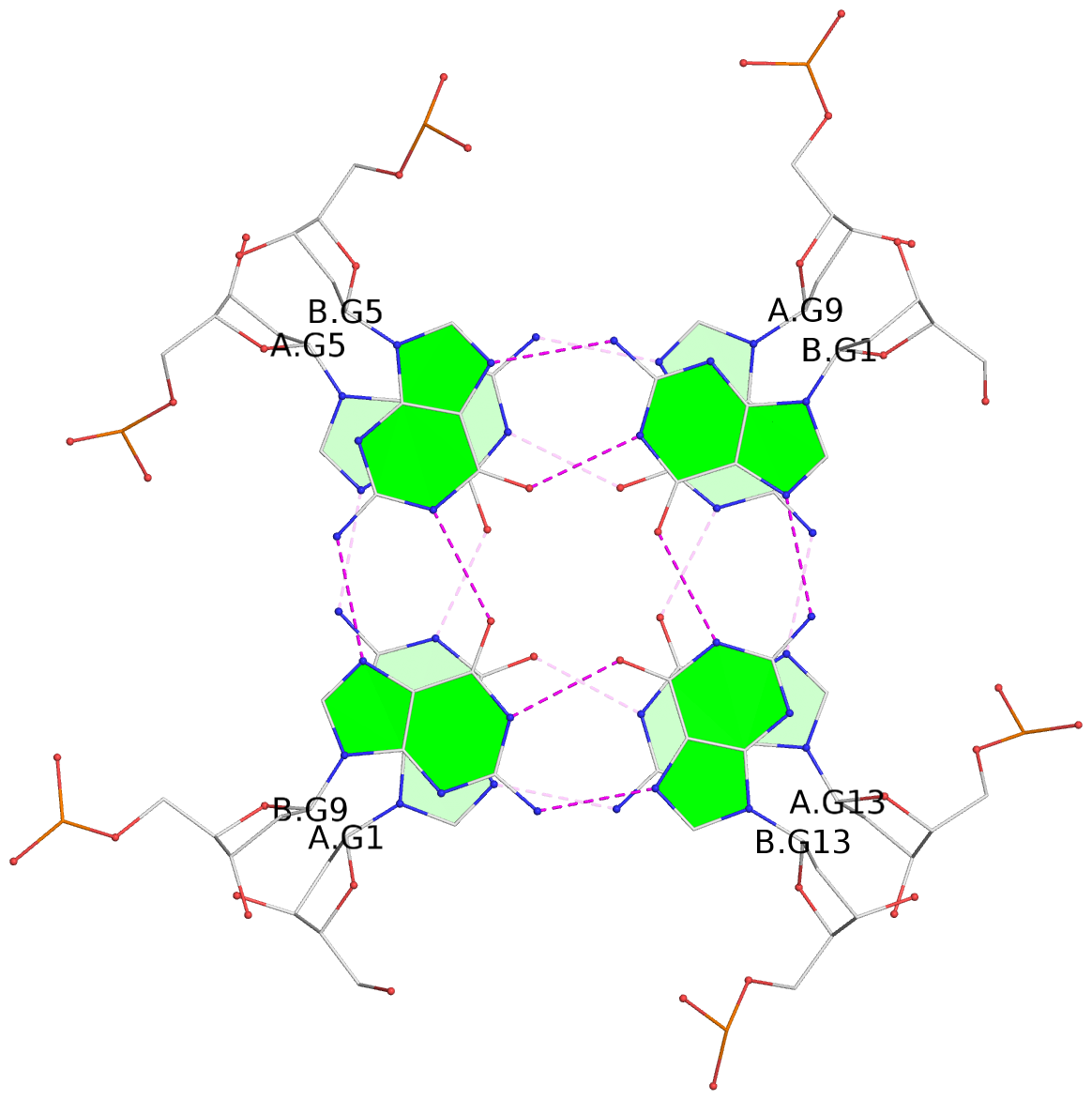
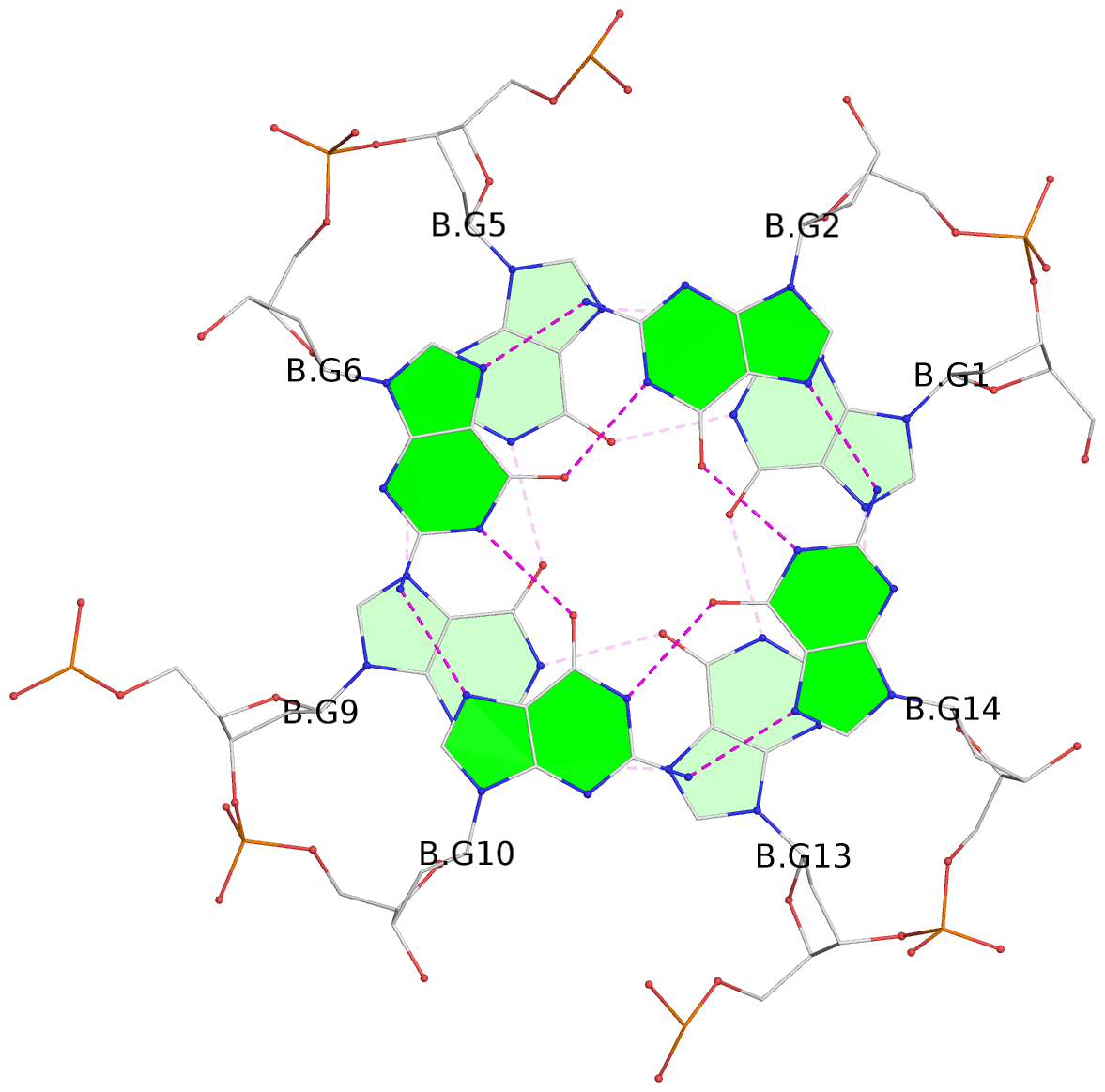
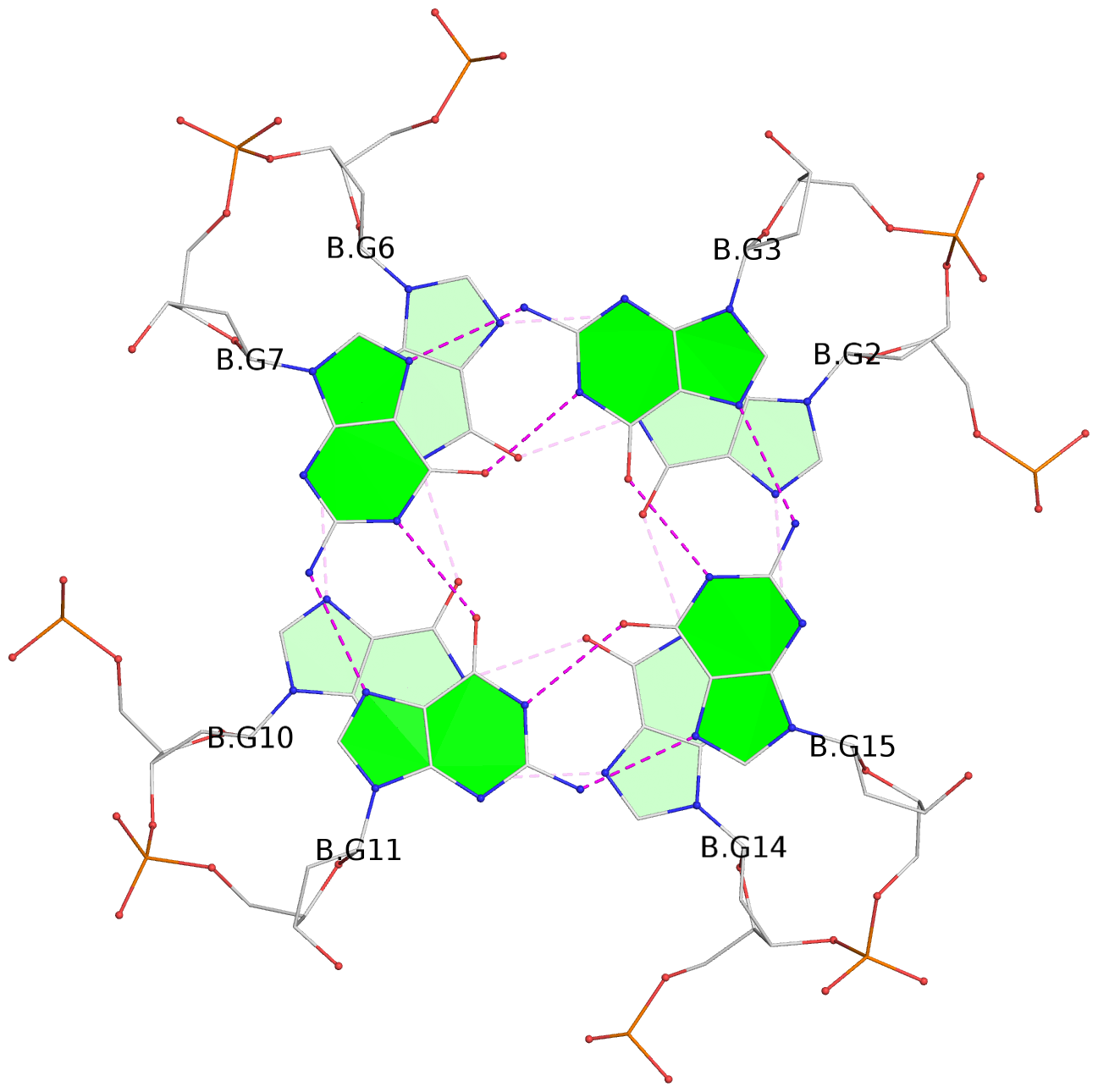
1 glyco-bond=---- sugar=---- groove=---- planarity=0.084 type=planar nts=4 GGGG A.DG1,A.DG5,A.DG9,A.DG13 2 glyco-bond=---- sugar=---- groove=---- planarity=0.129 type=planar nts=4 GGGG A.DG2,A.DG6,A.DG10,A.DG14 3 glyco-bond=---- sugar=---- groove=---- planarity=0.191 type=other nts=4 GGGG A.DG3,A.DG7,A.DG11,A.DG15 4 glyco-bond=---- sugar=---- groove=---- planarity=0.084 type=planar nts=4 GGGG B.DG1,B.DG5,B.DG9,B.DG13 5 glyco-bond=---- sugar=---- groove=---- planarity=0.115 type=planar nts=4 GGGG B.DG2,B.DG6,B.DG10,B.DG14 6 glyco-bond=---- sugar=---- groove=---- planarity=0.149 type=planar nts=4 GGGG B.DG3,B.DG7,B.DG11,B.DG15 7 glyco-bond=---- sugar=---- groove=---- planarity=0.056 type=planar nts=4 GGGG C.DG1,C.DG5,C.DG9,C.DG13 8 glyco-bond=---- sugar=---- groove=---- planarity=0.108 type=planar nts=4 GGGG C.DG2,C.DG6,C.DG10,C.DG14 9 glyco-bond=---- sugar=---- groove=---- planarity=0.164 type=other nts=4 GGGG C.DG3,C.DG7,C.DG11,C.DG15 10 glyco-bond=---- sugar=---- groove=---- planarity=0.058 type=planar nts=4 GGGG D.DG1,D.DG5,D.DG9,D.DG13 11 glyco-bond=---- sugar=---- groove=---- planarity=0.089 type=planar nts=4 GGGG D.DG2,D.DG6,D.DG10,D.DG14 12 glyco-bond=---- sugar=---- groove=---- planarity=0.154 type=planar nts=4 GGGG D.DG3,D.DG7,D.DG11,D.DG15
List of 2 G4-helices
In DSSR, a G4-helix is defined by stacking interactions of G-tetrads, regardless of backbone connectivity, and may contain more than one G4-stem.
Helix#1, 6 G-tetrad layers, inter-molecular, with 2 stems
Helix#2, 6 G-tetrad layers, inter-molecular, with 2 stems
List of 4 G4-stems
In DSSR, a G4-stem is defined as a G4-helix with backbone connectivity. Bulges are also allowed along each of the four strands.
Stem#1, 3 G-tetrad layers, 3 loops, INTRA-molecular, UUUU, parallel, 3(-P-P-P), parallel(4+0)
Stem#2, 3 G-tetrad layers, 3 loops, INTRA-molecular, UUUU, parallel, 3(-P-P-P), parallel(4+0)
Stem#3, 3 G-tetrad layers, 3 loops, INTRA-molecular, UUUU, parallel, 3(-P-P-P), parallel(4+0)
Stem#4, 3 G-tetrad layers, 3 loops, INTRA-molecular, UUUU, parallel, 3(-P-P-P), parallel(4+0)
List of 2 G4 coaxial stacks
1 G4 helix#1 contains 2 G4 stems: [#1,#2] [5'/5'] 2 G4 helix#2 contains 2 G4 stems: [#3,#4] [5'/5']