Detailed DSSR results for the G-quadruplex: PDB entry 6r9k
Created and maintained by Xiang-Jun Lu <xiangjun@x3dna.org>
Citation: Please cite the NAR'20 DSSR-PyMOL schematics paper and/or the NAR'15 DSSR method paper.
Summary information
- PDB id
- 6r9k
- Class
- DNA
- Method
- NMR
- Summary
- A quadruplex hybrid structure with lpp loop orientation and 3 syn residues
- Reference
- Karg B, Mohr S, Weisz K (2019): "Duplex-Guided Refolding into Novel G-Quadruplex (3+1) Hybrid Conformations." Angew.Chem.Int.Ed.Engl., 58, 11068-11071. doi: 10.1002/anie.201905372.
- Abstract
- The oligomer d(GCGTG3 TCAG3 TG3 TG3 ACGC) with short complementary flanking sequences at the 5'- and 3'-ends was shown to fold into three different DNA G-quadruplex species. In contrast, a corresponding oligomer that lacks base complementarity between the two overhang sequences folds into a single parallel G-quadruplex. The three coexisting quadruplex structures were unambiguously identified and structurally characterized through detailed spectral comparisons with well-defined G-quadruplexes formed upon the deliberate incorporation of syn-favoring 8-bromoguanosine analogues into specific positions of the G-core. Two (3+1) hybrid structures coexist with the parallel fold and feature a novel lateral-propeller-propeller loop architecture that has not yet been confirmed experimentally. Both hybrid quadruplexes adopt the same topology and only differ in their pattern of anti→syn transitions and tetrad stackings.
- G4 notes
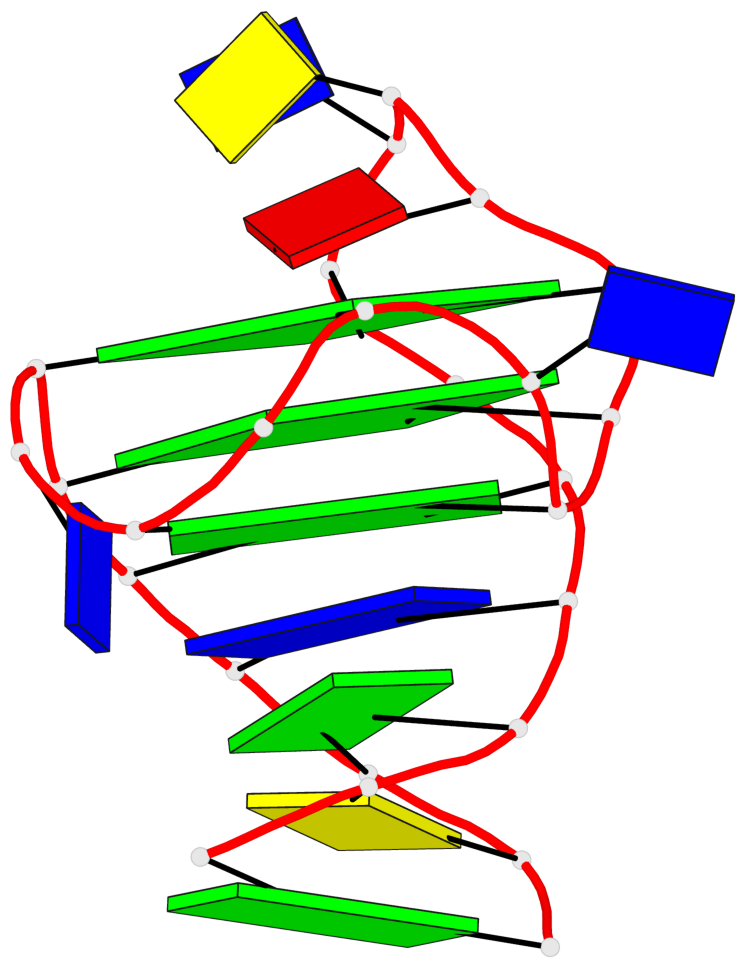
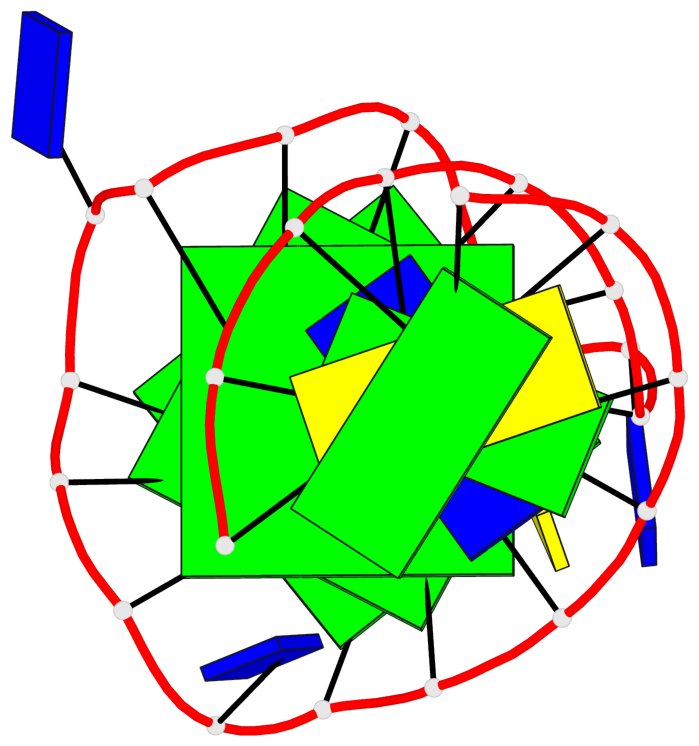
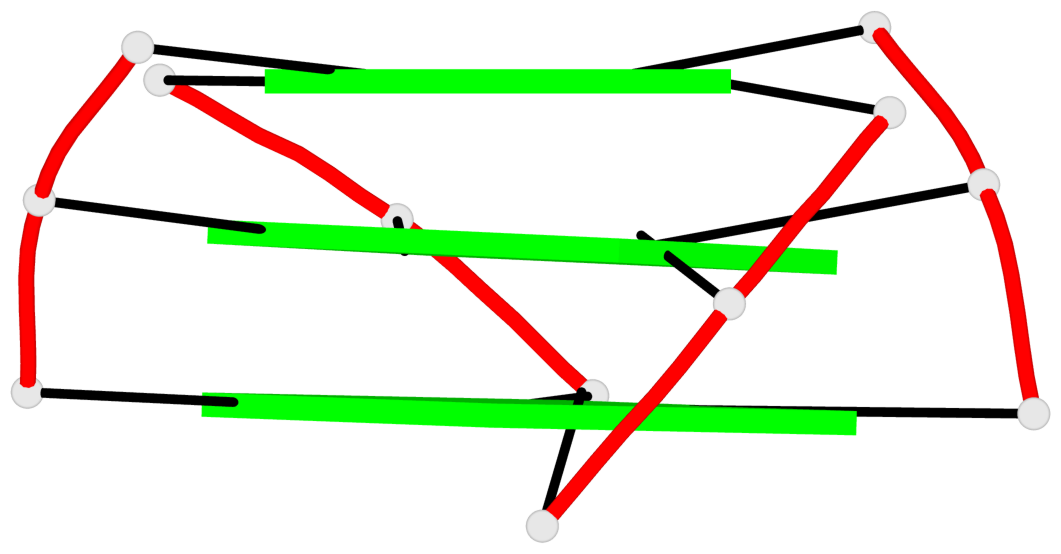
- 3 G-tetrads, 1 G4 helix, 1 G4 stem, 3(+Ln+P+P), hybrid-1R(1+3), UDDD
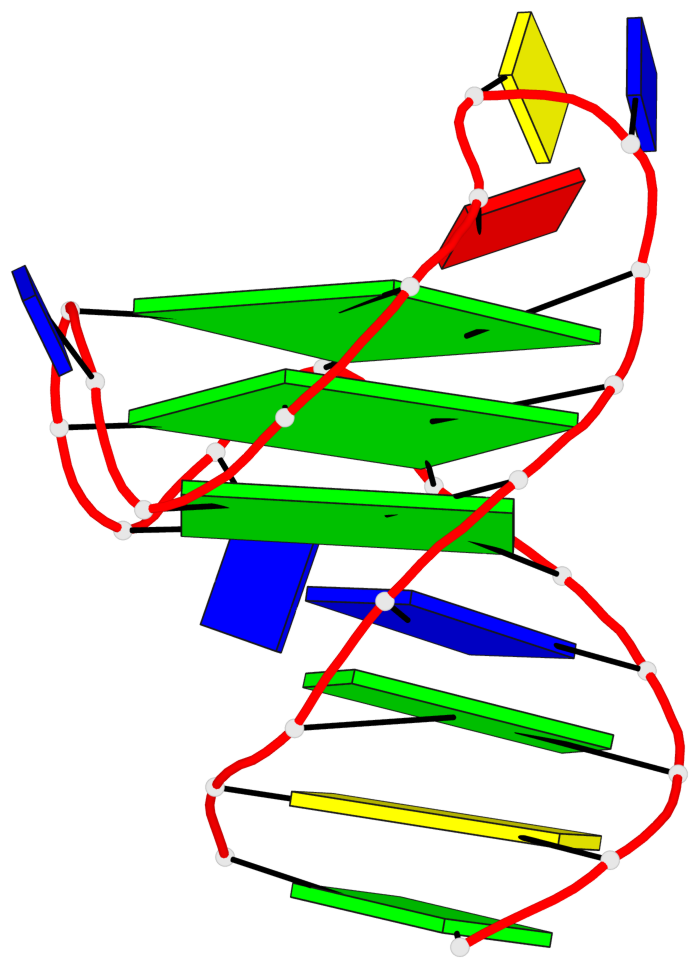
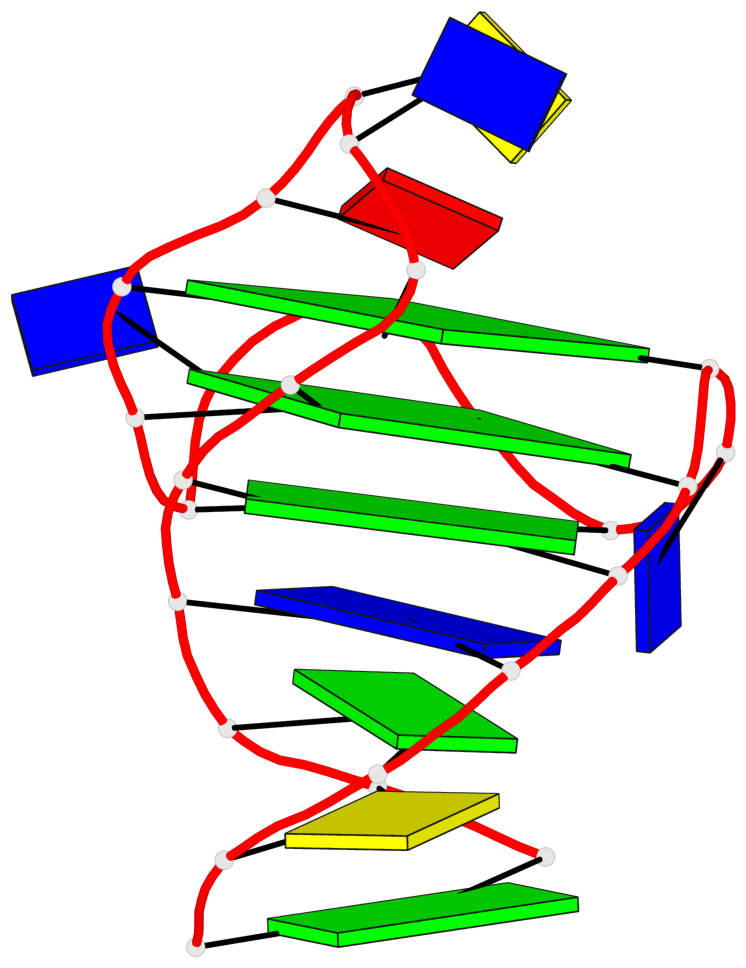
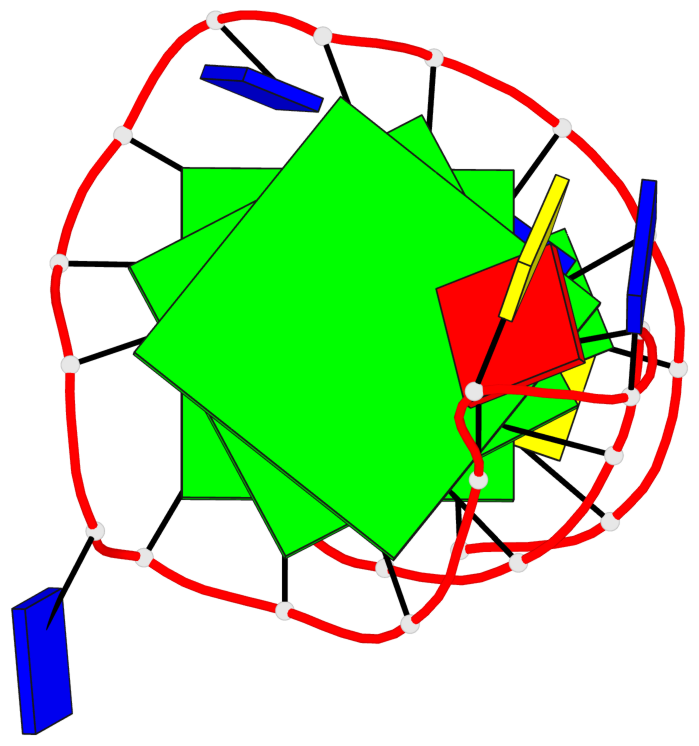
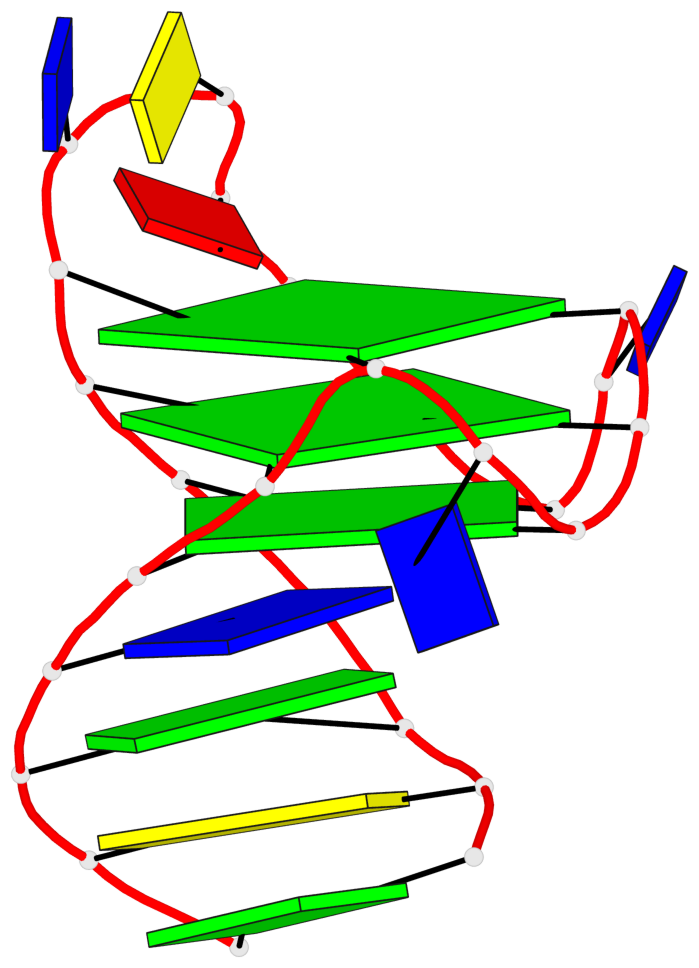
Base-block schematics in six views
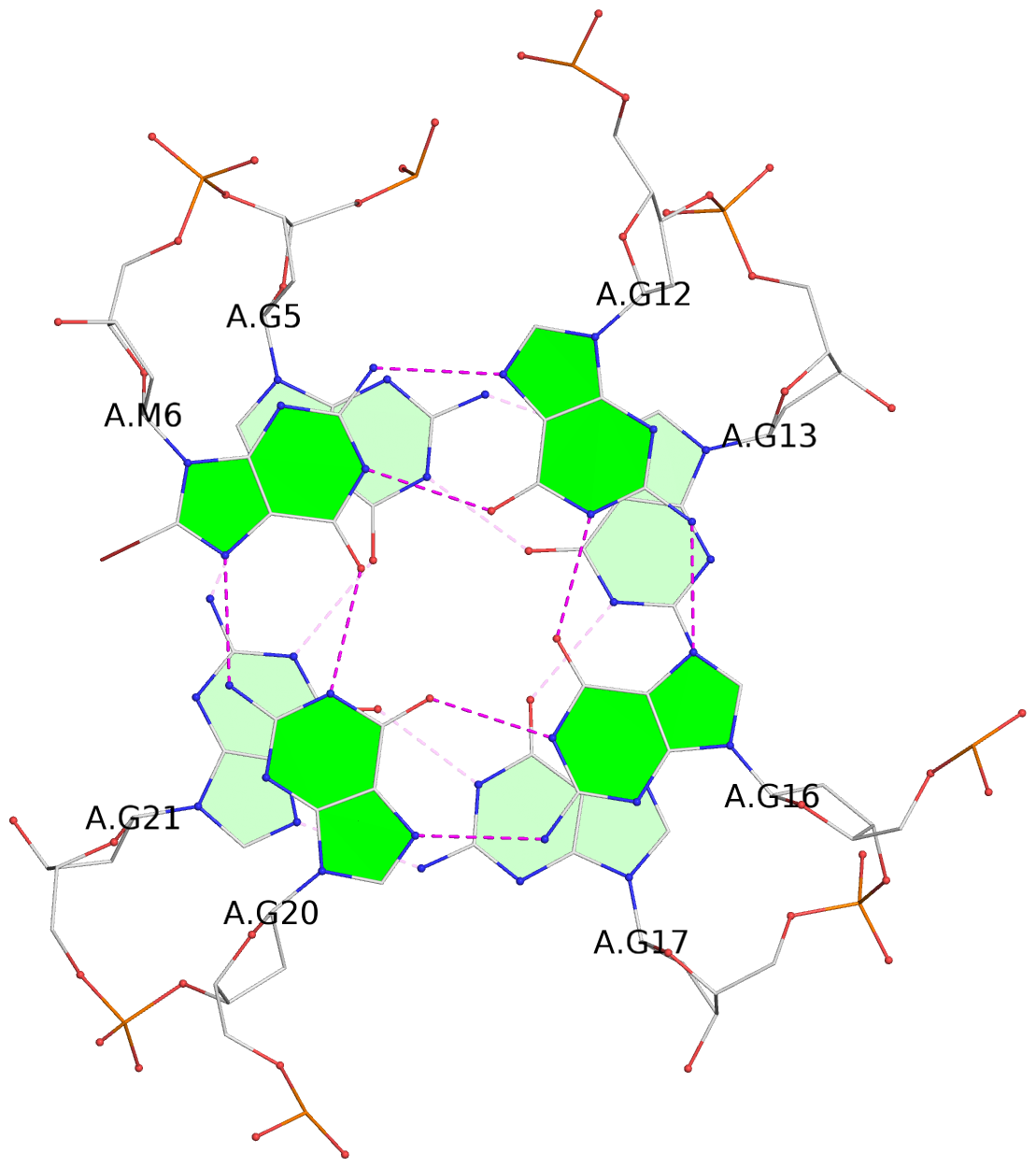
List of 3 G-tetrads
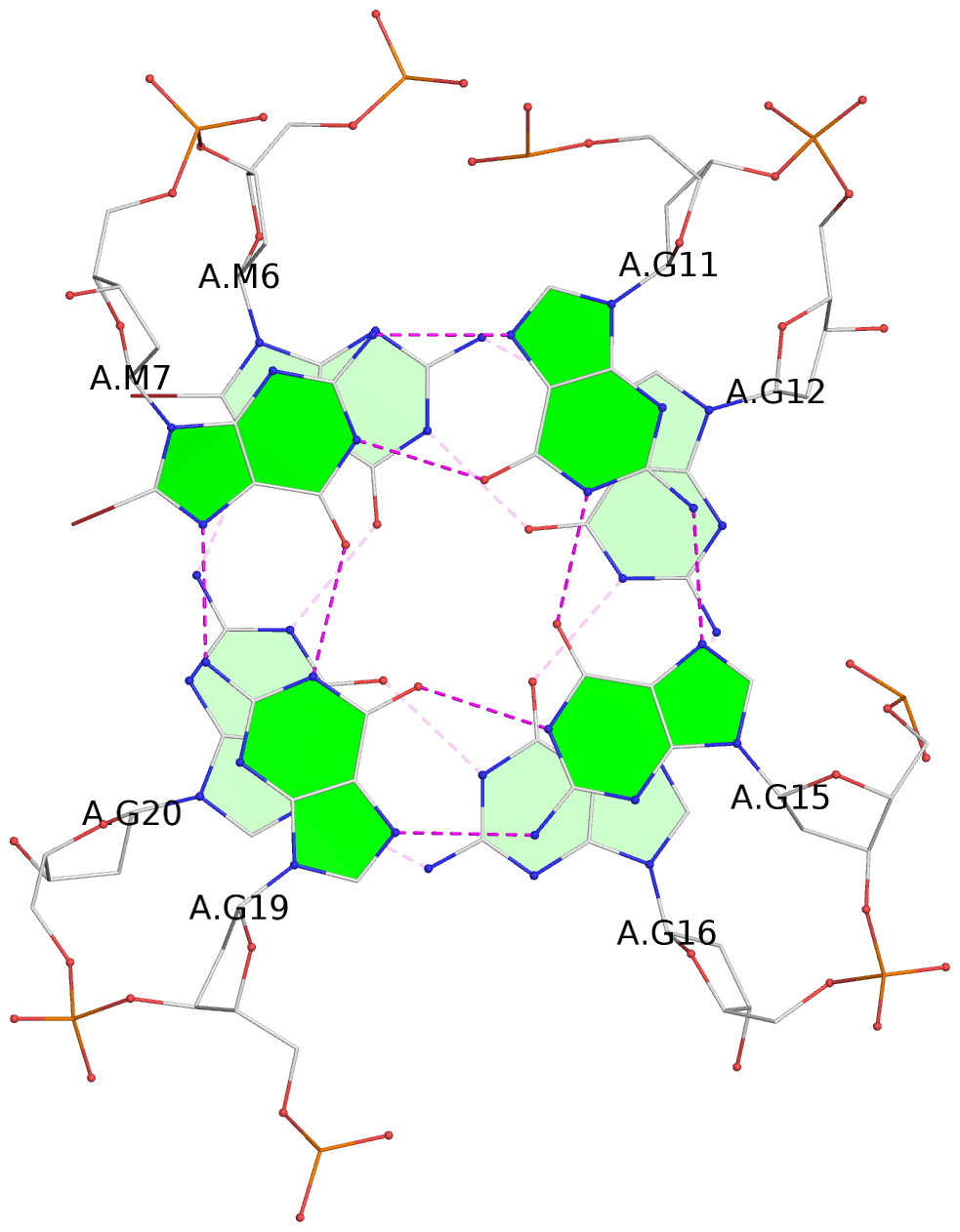
1 glyco-bond=s--- sugar=---- groove=w--n planarity=0.108 type=planar nts=4 GGGG A.DG5,A.DG21,A.DG17,A.DG13 2 glyco-bond=s--- sugar=---3 groove=w--n planarity=0.139 type=planar nts=4 gGGG A.BGM6,A.DG20,A.DG16,A.DG12 3 glyco-bond=s--- sugar=---- groove=w--n planarity=0.284 type=other nts=4 gGGG A.BGM7,A.DG19,A.DG15,A.DG11
List of 1 G4-helix
In DSSR, a G4-helix is defined by stacking interactions of G-tetrads, regardless of backbone connectivity, and may contain more than one G4-stem.
Helix#1, 3 G-tetrad layers, INTRA-molecular, with 1 stem
List of 1 G4-stem
In DSSR, a G4-stem is defined as a G4-helix with backbone connectivity. Bulges are also allowed along each of the four strands.