Detailed DSSR results for the G-quadruplex: PDB entry 6t51
Created and maintained by Xiang-Jun Lu <xiangjun@x3dna.org>
Citation: Please cite the NAR'20 DSSR-PyMOL schematics paper and/or the NAR'15 DSSR method paper.
Summary information
- PDB id
- 6t51
- Class
- DNA
- Method
- NMR
- Summary
- NMR structure of kras22rt g-quadruplex forming within kras promoter region at physological temperature
- Reference
- Marquevielle J, Kumar MVV, Mergny JL, Salgado GF (2018): "1H,13C, and15N chemical shift assignments of a G-quadruplex forming sequence within the KRAS proto-oncogene promoter region." Biomol.Nmr Assign., 12, 123-127. doi: 10.1007/s12104-017-9793-0.
- Abstract
- Single stranded guanine rich DNA (or RNA) sequences adopt noncanonical secondary structures called G-quadruplexes (G4). Functionally, quadruplexes control gene transcription and regulate activities such as replication, gene recombination or alternative splicing. Hence they are potential targets for cancer, neuronal, and viral related diseases. KRAS is one of the most mutated oncogenes in the genome of cancer cells and contains a nuclease hypersensitive element (NHE) sequence capable of forming G-quadruplexes via its six runs of guanines. In our work, we are interested in the NMR structure of the major G4 scaffold formed in the KRAS NHE region with a mutated sequence of 22 residues. Here, we report 1H, 13C and 15N chemical shift assignments the G4 formed within KRAS22RT sequence.
- G4 notes
- 3 G-tetrads, 1 G4 helix, 1 G4 stem, 3(-P-P-P), parallel(4+0), UUUU
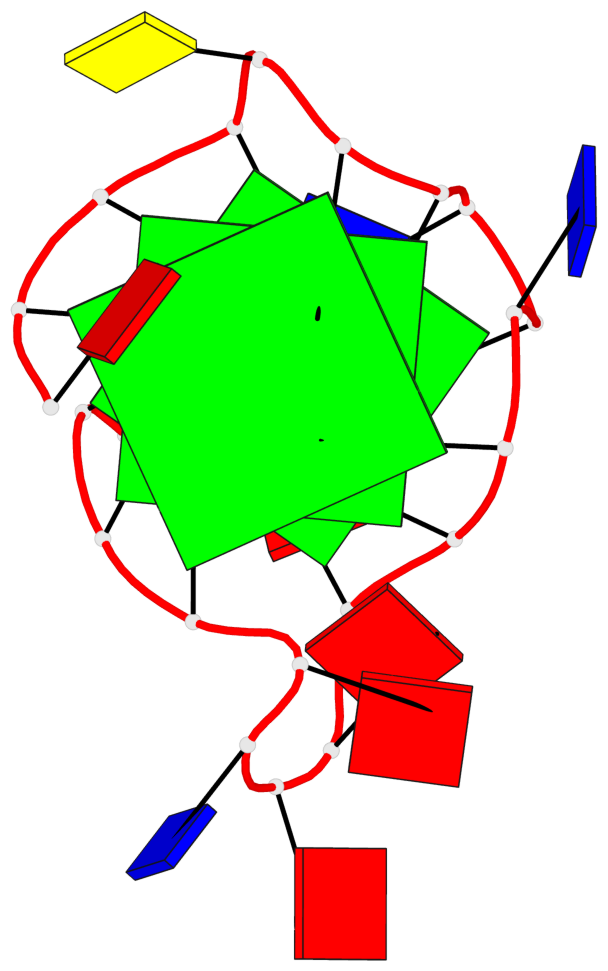
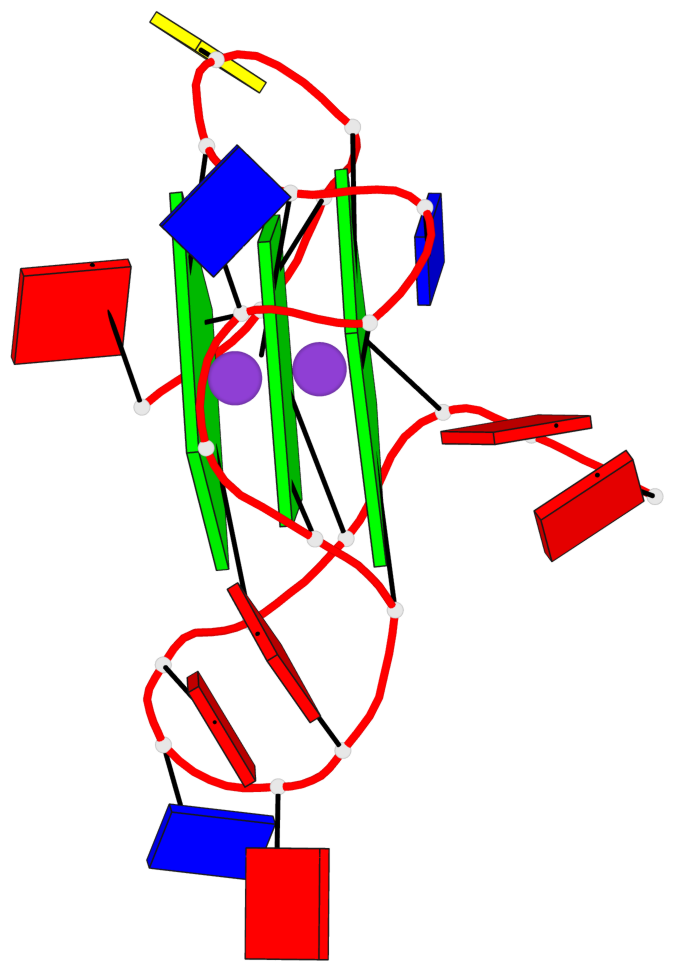
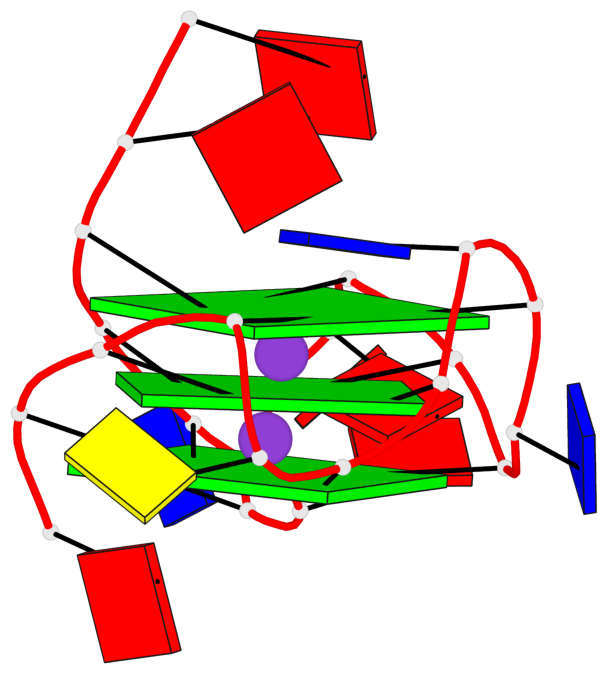
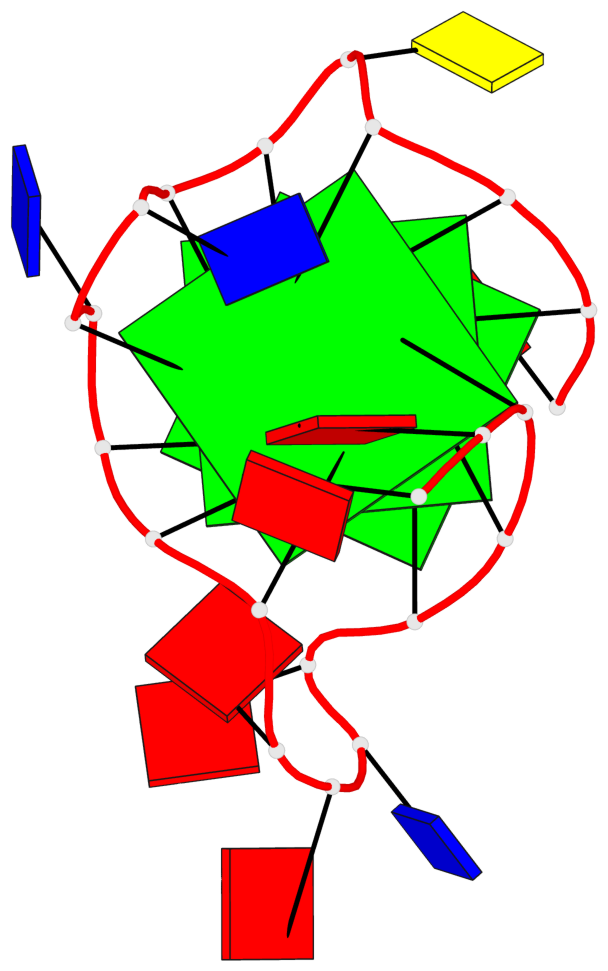
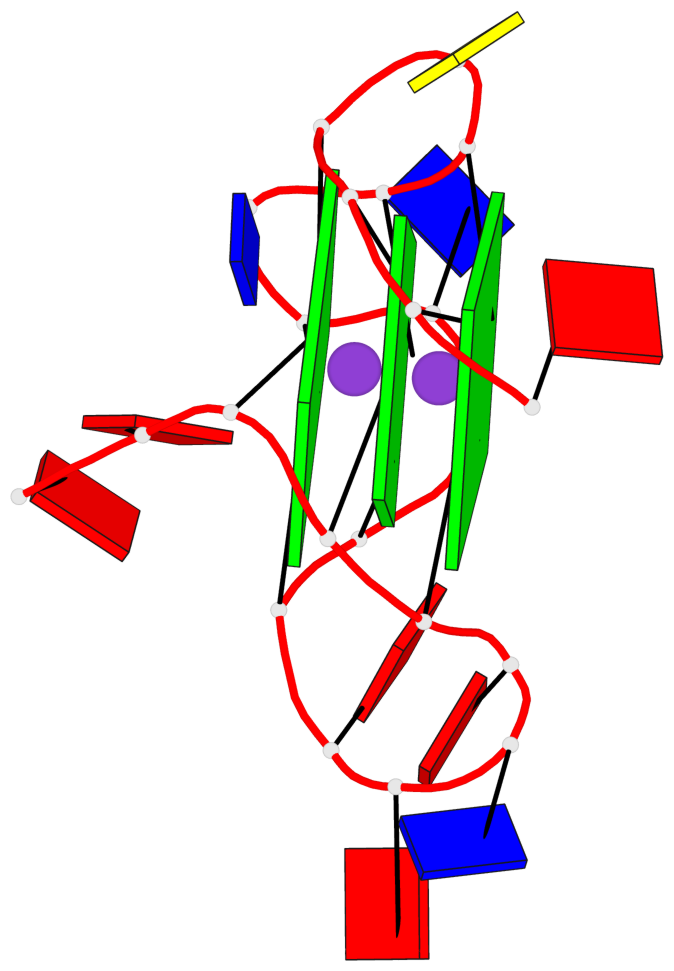
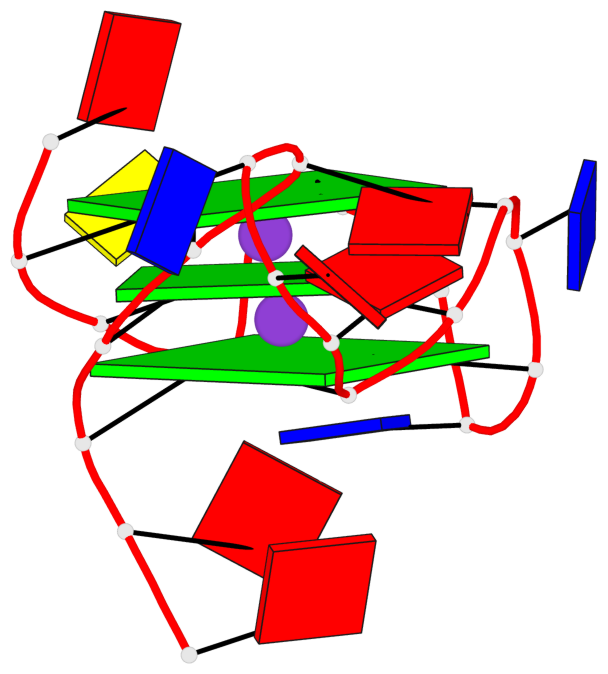
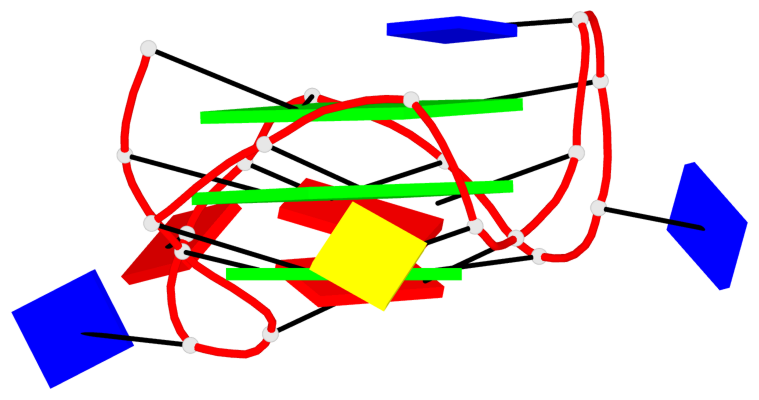
Base-block schematics in six views
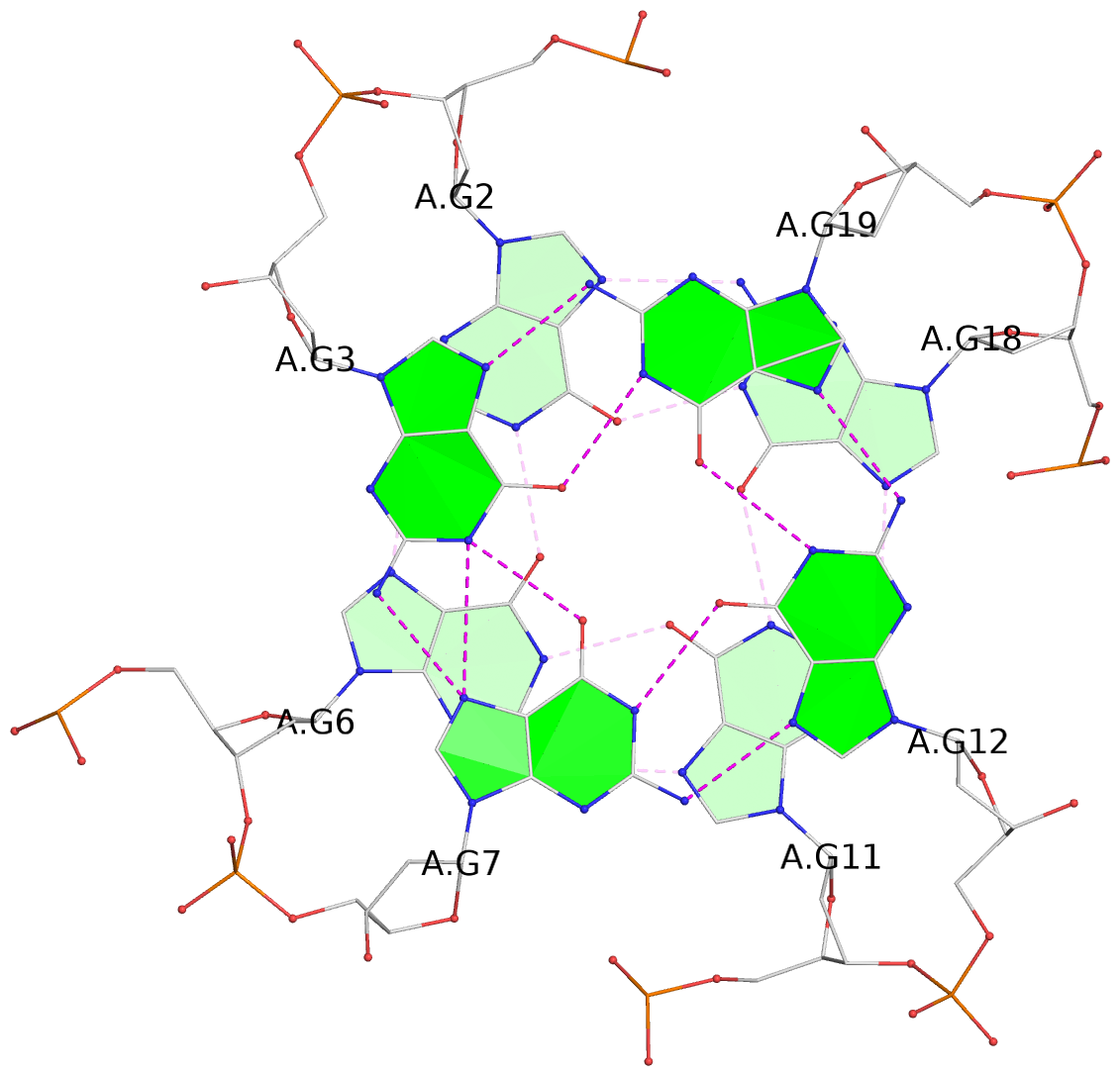
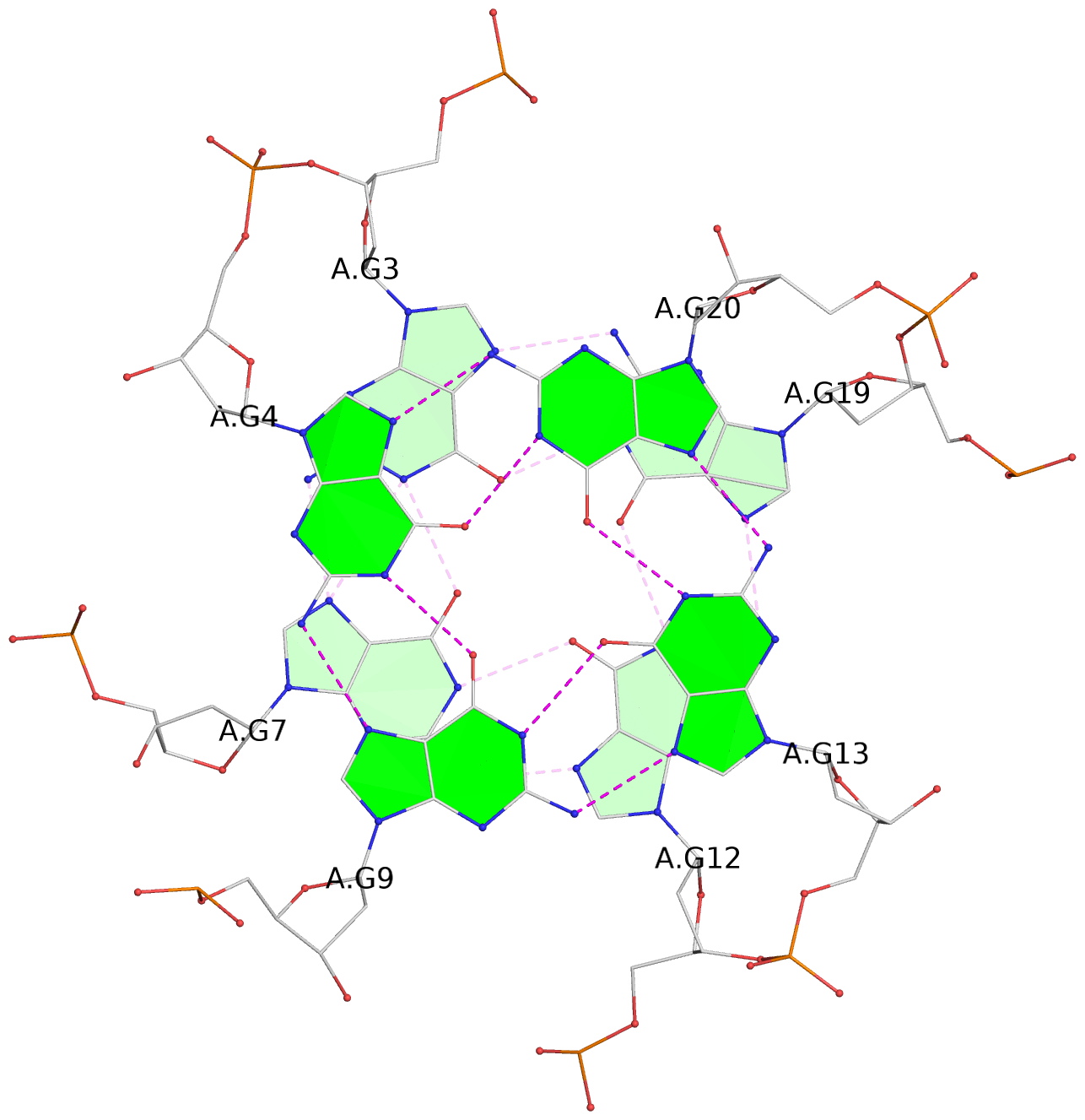
List of 3 G-tetrads
1 glyco-bond=---- sugar=---- groove=---- planarity=0.186 type=other nts=4 GGGG A.DG2,A.DG6,A.DG11,A.DG18 2 glyco-bond=---- sugar=..-- groove=---- planarity=0.252 type=other nts=4 GGGG A.DG3,A.DG7,A.DG12,A.DG19 3 glyco-bond=---- sugar=---- groove=---- planarity=0.163 type=other nts=4 GGGG A.DG4,A.DG9,A.DG13,A.DG20
List of 1 G4-helix
In DSSR, a G4-helix is defined by stacking interactions of G-tetrads, regardless of backbone connectivity, and may contain more than one G4-stem.
Helix#1, 3 G-tetrad layers, INTRA-molecular, with 1 stem
List of 1 G4-stem
In DSSR, a G4-stem is defined as a G4-helix with backbone connectivity. Bulges are also allowed along each of the four strands.